diff --git a/.gitignore b/.gitignore
index 9732e6cf..1b663cc0 100644
--- a/.gitignore
+++ b/.gitignore
@@ -1,4 +1,3 @@
-Secrets/
*.lock*
*.env
*.bak
@@ -10,4 +9,14 @@ Secrets/
*.pytest_cache
*.cache
*pyc
-docker-compose.override.yml
+#docker-compose.override.yml
+# for local dev
+credential*.json
+venv/
+.idea/
+
+/fence/
+
+/output/
+
+
diff --git a/README.md b/README.md
index 7c110379..34710eee 100644
--- a/README.md
+++ b/README.md
@@ -1,6 +1,17 @@
+
+ACED - Training
+===
+> This repo was established as a sandbox to introduce Gen3 in a local, docker-compose laptop friendly environment.
+
+Follow the instructions below to introduce Gen3. Then, see ACED specific steps [here](docs/aced.md)
+
+
+
Compose-Services
===
+
+
Docker-compose setup for experimental commons, small commons, or local development of the Gen3 stack. Production use should use [cloud-automation](https://github.com/uc-cdis/cloud-automation).
This setup uses Docker containers for the [Gen3 microservices](https://github.com/uc-cdis/) and nginx. The microservices and nginx images are pulled from quay.io (master), while Postgres (9.5) images are pulled from Docker Hub. Nginx is used as a reverse proxy to each of the services.
diff --git a/datadictionary/README.md b/datadictionary/README.md
index 60d1792b..5ec4b641 100644
--- a/datadictionary/README.md
+++ b/datadictionary/README.md
@@ -1,3 +1,15 @@
+
+
+ACED - Training
+===
+> This repo was established as a sandbox to introduce Gen3 in a local, docker-compose laptop friendly environment.
+
+```commandline
+cp ../data_model/DATA/gen3/*.yaml datadictionary/gdcdictionary/schemas/
+
+
+```
+
# Data Dictionary
The data dictionary provides the first level of validation for all data
diff --git a/datadictionary/aced.json b/datadictionary/aced.json
new file mode 100644
index 00000000..c8ce19ce
--- /dev/null
+++ b/datadictionary/aced.json
@@ -0,0 +1 @@
+{"DocumentReference.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "additionalProperties": false, "category": "data_file", "description": " A reference to a document.\n \n A reference to a document of any kind for any purpose. Provides metadata\n about the document so that the document can be discovered and managed. The\n scope of a document is any seralized object with a mime-type, so includes\n formal patient centric documents (CDA), cliical notes, scanned paper, and\n non-patient specific documents like policy text.\n ", "id": "DocumentReference", "links": [{"backref": "DocumentReferences", "label": "Patients", "multiplicity": "many_to_many", "name": "Patients", "required": true, "target_type": "Patient"}, {"backref": "DocumentReferences", "label": "Organizations", "multiplicity": "many_to_many", "name": "Organizations", "required": false, "target_type": "Organization"}], "namespace": "http://hl7.org/fhir", "program": "*", "project": "*", "properties": {"$ref": "_definitions.yaml#/data_file_properties", "data_category": {"term": {"$ref": "_terms.yaml#/data_category"}, "type": "string"}, "data_type": {"term": {"$ref": "_terms.yaml#/data_type"}, "type": "string"}, "data_format": {"term": {"$ref": "_terms.yaml#/data_format"}, "type": "string"}, "author_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "author_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "category_coding_us_core_documentreference_category": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "content_attachment_contentType": {"description": "Mime type of the content, with charset etc..", "type": ["string", "null"]}, "content_attachment_data": {"description": "Data inline, base64ed.", "type": ["string", "null"]}, "content_attachment_md5": {"description": "", "type": ["string", "null"]}, "content_attachment_size": {"description": "Number of bytes of content (if url provided).", "type": ["number", "null"]}, "content_attachment_url": {"description": "Uri where the data can be found.", "type": ["string", "null"]}, "content_format_IHE_FormatCode_codesystem": {"description": "Format/content rules for the document.", "type": ["string", "null"]}, "context_encounter_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "context_period_end": {"description": "End time with inclusive boundary, if not ongoing.", "type": ["string", "null"]}, "context_period_start": {"description": "Starting time with inclusive boundary.", "type": ["string", "null"]}, "created_datetime": {"$ref": "_definitions.yaml#/datetime"}, "custodian_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "custodian_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "date": {"description": "When this document reference was created.", "type": ["string", "null"]}, "id": {"description": "", "type": ["string", "null"]}, "identifier_urn_ietf_rfc_3986": {"description": "Other identifiers for the document.", "type": ["string", "null"]}, "meta_profile": {"description": "Profiles this resource claims to conform to.", "type": ["string", "null"]}, "project_id": {"$ref": "_definitions.yaml#/project_id"}, "resourceType": {"description": "One of the resource types defined as part of this version of FHIR", "type": ["string", "null"]}, "state": {"$ref": "_definitions.yaml#/state"}, "status": {"description": "The status of this document reference.. http://hl7.org/fhir/document-reference-status", "enum": ["current", "superseded", "entered-in-error"], "term": {"description": "The status of this document reference.. http://hl7.org/fhir/document-reference-status", "termDef": {"cde_id": "http://hl7.org/fhir/document-reference-status", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/document-reference-status", "term_url": "http://hl7.org/fhir/document-reference-status"}}}, "subject_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "submitter_id": {"type": ["string", "null"]}, "type": {"type": "string"}, "type_coding_0_code": {"description": "Symbol in syntax defined by the system.", "type": ["string", "null"]}, "type_coding_0_display": {"description": "Representation defined by the system.", "type": ["string", "null"]}, "type_coding_0_system": {"description": "Identity of the terminology system.", "type": ["string", "null"]}, "type_coding_1_code": {"description": "Symbol in syntax defined by the system.", "type": ["string", "null"]}, "type_coding_1_display": {"description": "Representation defined by the system.", "type": ["string", "null"]}, "type_coding_1_system": {"description": "Identity of the terminology system.", "type": ["string", "null"]}, "updated_datetime": {"$ref": "_definitions.yaml#/datetime"}}, "required": ["submitter_id", "type", "status"], "submittable": true, "systemProperties": ["id", "project_id", "created_datetime", "updated_datetime", "state"], "title": "DocumentReference", "type": "object", "uniqueKeys": [["id"], ["project_id", "submitter_id"]], "validators": null}, "Observation.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "additionalProperties": false, "category": "Clinical", "description": " Measurements and simple assertions.\n \n Measurements and simple assertions made about a patient, device or other\n subject.\n ", "id": "Observation", "links": [{"backref": "Observations", "label": "ResearchStudies", "multiplicity": "many_to_many", "name": "ResearchStudies", "required": false, "target_type": "ResearchStudy"}, {"backref": "Observations", "label": "Specimen", "multiplicity": "many_to_many", "name": "Specimen", "required": false, "target_type": "Specimen"}, {"backref": "Observations", "label": "Patients", "multiplicity": "many_to_many", "name": "Patients", "required": true, "target_type": "Patient"}, {"backref": "Observations", "label": "Encounters", "multiplicity": "many_to_many", "name": "Encounters", "required": false, "target_type": "Encounter"}], "namespace": "http://hl7.org/fhir", "program": "*", "project": "*", "properties": {"category_coding_observation_category": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "code_coding_0_code": {"description": "Symbol in syntax defined by the system.", "type": ["string", "null"]}, "code_coding_0_display": {"description": "Representation defined by the system.", "type": ["string", "null"]}, "code_coding_0_system": {"description": "Identity of the terminology system.", "type": ["string", "null"]}, "code_coding_1_code": {"description": "Symbol in syntax defined by the system.", "type": ["string", "null"]}, "code_coding_1_display": {"description": "Representation defined by the system.", "type": ["string", "null"]}, "code_coding_1_system": {"description": "Identity of the terminology system.", "type": ["string", "null"]}, "code_coding_loinc_org": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "code_coding_sct": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "code_text": {"description": "Plain text representation of the concept.", "type": ["string", "null"]}, "component_0_code_coding_loinc_org": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "component_0_code_text": {"description": "Plain text representation of the concept.", "type": ["string", "null"]}, "component_0_valueQuantity_code": {"description": "Coded form of the unit.", "type": ["string", "null"]}, "component_0_valueQuantity_system": {"description": "System that defines coded unit form.", "type": ["string", "null"]}, "component_0_valueQuantity_unit": {"description": "Unit representation.", "type": ["string", "null"]}, "component_0_valueQuantity_value": {"description": "Numerical value (with implicit precision).", "type": ["number", "null"]}, "component_1_code_coding_loinc_org": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "component_1_code_text": {"description": "Plain text representation of the concept.", "type": ["string", "null"]}, "component_1_valueQuantity_code": {"description": "Coded form of the unit.", "type": ["string", "null"]}, "component_1_valueQuantity_system": {"description": "System that defines coded unit form.", "type": ["string", "null"]}, "component_1_valueQuantity_unit": {"description": "Unit representation.", "type": ["string", "null"]}, "component_1_valueQuantity_value": {"description": "Numerical value (with implicit precision).", "type": ["number", "null"]}, "created_datetime": {"$ref": "_definitions.yaml#/datetime"}, "effectiveDateTime": {"description": "Clinically relevant time/time-period for observation.", "type": ["string", "null"]}, "encounter_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "id": {"description": "", "type": ["string", "null"]}, "issued": {"description": "Date/Time this version was made available.", "type": ["string", "null"]}, "meta_profile": {"description": "Profiles this resource claims to conform to.", "type": ["string", "null"]}, "meta_profile_0": {"description": "Profiles this resource claims to conform to.", "type": ["string", "null"]}, "meta_profile_1": {"description": "Profiles this resource claims to conform to.", "type": ["string", "null"]}, "project_id": {"$ref": "_definitions.yaml#/project_id"}, "resourceType": {"description": "One of the resource types defined as part of this version of FHIR", "type": ["string", "null"]}, "state": {"$ref": "_definitions.yaml#/state"}, "status": {"description": "The status of the result value.. http://hl7.org/fhir/observation-status", "enum": ["registered", "preliminary", "final", "amended", "corrected", "cancelled", "entered-in-error", "unknown"], "term": {"description": "The status of the result value.. http://hl7.org/fhir/observation-status", "termDef": {"cde_id": "http://hl7.org/fhir/observation-status", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/observation-status", "term_url": "http://hl7.org/fhir/observation-status"}}}, "subject_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "submitter_id": {"type": ["string", "null"]}, "type": {"type": "string"}, "updated_datetime": {"$ref": "_definitions.yaml#/datetime"}, "valueCodeableConcept_coding_loinc_org": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "valueCodeableConcept_coding_sct": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "valueCodeableConcept_text": {"description": "Plain text representation of the concept.", "type": ["string", "null"]}, "valueQuantity_code": {"description": "Coded form of the unit.", "type": ["string", "null"]}, "valueQuantity_system": {"description": "System that defines coded unit form.", "type": ["string", "null"]}, "valueQuantity_unit": {"description": "Unit representation.", "type": ["string", "null"]}, "valueQuantity_value": {"description": "Numerical value (with implicit precision).", "type": ["number", "null"]}, "valueSampledData_data": {"description": "Decimal values with spaces, or \"E\" | \"U\" | \"L\".", "type": ["string", "null"]}, "valueSampledData_dimensions": {"description": "Number of sample points at each time point.", "type": ["number"]}, "valueSampledData_origin_system": {"description": "System that defines coded unit form.", "type": ["string", "null"]}, "valueSampledData_origin_value": {"description": "Numerical value (with implicit precision).", "type": ["number", "null"]}, "valueSampledData_period": {"description": "Number of milliseconds between samples.", "type": ["number"]}, "valueString": {"description": "Actual result.", "type": ["string", "null"]}}, "required": ["submitter_id", "type", "status"], "submittable": true, "systemProperties": ["id", "project_id", "created_datetime", "updated_datetime", "state"], "title": "Observation", "type": "object", "uniqueKeys": [["id"], ["project_id", "submitter_id"]], "validators": null}, "Patient.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "additionalProperties": false, "category": "Administrative", "description": " Information about an individual or animal receiving health care services.\n \n Demographics and other administrative information about an individual or\n animal receiving care or other health-related services.\n ", "id": "Patient", "links": [{"backref": "Patients", "label": "Organizations", "multiplicity": "many_to_many", "name": "Organizations", "required": true, "target_type": "Organization"}], "namespace": "http://hl7.org/fhir", "program": "*", "project": "*", "properties": {"address_city": {"description": "Name of city, town etc..", "type": ["string", "null"]}, "address_country": {"description": "Country (e.g. can be ISO 3166 2 or 3 letter code).", "type": ["string", "null"]}, "address_geolocation_latitude": {"description": "The absolute geographic location of the Location, expressed using the WGS84 datum (This is the same co-ordinate system used in KML).\nLatitude. The value domain and the interpretation are the same as for the text of the latitude element in KML (see notes below).", "type": ["number", "null"]}, "address_geolocation_longitude": {"description": "The absolute geographic location of the Location, expressed using the WGS84 datum (This is the same co-ordinate system used in KML).\nLongitude. The value domain and the interpretation are the same as for the text of the longitude element in KML (see notes below).", "type": ["number", "null"]}, "address_line": {"description": "Street name, number, direction & P.O. Box etc..", "type": ["string", "null"]}, "address_postalCode": {"description": "Postal code for area.", "type": ["string", "null"]}, "address_state": {"description": "Sub-unit of country (abbreviations ok).", "type": ["string", "null"]}, "birthDate": {"description": "The date of birth for the individual.", "type": ["string", "null"]}, "communication_language_coding_iso639_1": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "communication_language_coding_urn_ietf_bcp_47": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "communication_language_text": {"description": "Plain text representation of the concept.", "type": ["string", "null"]}, "created_datetime": {"$ref": "_definitions.yaml#/datetime"}, "deceasedDateTime": {"description": "Indicates if the individual is deceased or not.", "type": ["string", "null"]}, "disability_adjusted_life_years": {"description": "Disability Adjusted Life Years as defined in the literature and summarized at .", "type": ["number", "null"]}, "gender": {"description": "Administrative Gender - the gender that the patient is considered to have for administration and record keeping purposes.. http://hl7.org/fhir/administrative-gender", "enum": ["male", "female", "other", "unknown"], "term": {"description": "Administrative Gender - the gender that the patient is considered to have for administration and record keeping purposes.. http://hl7.org/fhir/administrative-gender", "termDef": {"cde_id": "http://hl7.org/fhir/administrative-gender", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/administrative-gender", "term_url": "http://hl7.org/fhir/administrative-gender"}}}, "id": {"description": "", "type": ["string", "null"]}, "identifier_DL": {"description": "An identifier for this patient.", "type": ["string", "null"]}, "identifier_MR": {"description": "An identifier for this patient.", "type": ["string", "null"]}, "identifier_PPN": {"description": "An identifier for this patient.", "type": ["string", "null"]}, "identifier_SS": {"description": "An identifier for this patient.", "type": ["string", "null"]}, "identifier_synthea": {"description": "An identifier for this patient.", "type": ["string", "null"]}, "maritalStatus_coding_v3_MaritalStatus": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "maritalStatus_text": {"description": "Plain text representation of the concept.", "type": ["string", "null"]}, "meta_profile": {"description": "Profiles this resource claims to conform to.", "type": ["string", "null"]}, "multipleBirthBoolean": {"description": "Whether patient is part of a multiple birth.", "type": ["boolean", "null"]}, "multipleBirthInteger": {"description": "Whether patient is part of a multiple birth.", "type": ["number", "null"]}, "name_0_family": {"description": "Family name (often called 'Surname').", "type": ["string", "null"]}, "name_0_given": {"description": "Given names (not always 'first'). Includes middle names.", "type": ["string", "null"]}, "name_0_prefix": {"description": "Parts that come before the name.", "type": ["string", "null"]}, "name_0_suffix": {"description": "Parts that come after the name.", "type": ["string", "null"]}, "name_0_use": {"description": "Identifies the purpose for this name.. http://hl7.org/fhir/name-use", "enum": ["usual", "official", "temp", "nickname", "anonymous", "old", "maiden"], "term": {"description": "Identifies the purpose for this name.. http://hl7.org/fhir/name-use", "termDef": {"cde_id": "http://hl7.org/fhir/name-use", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/name-use", "term_url": "http://hl7.org/fhir/name-use"}}}, "name_1_family": {"description": "Family name (often called 'Surname').", "type": ["string", "null"]}, "name_1_given": {"description": "Given names (not always 'first'). Includes middle names.", "type": ["string", "null"]}, "name_1_prefix": {"description": "Parts that come before the name.", "type": ["string", "null"]}, "name_1_suffix": {"description": "Parts that come after the name.", "type": ["string", "null"]}, "name_1_use": {"description": "Identifies the purpose for this name.. http://hl7.org/fhir/name-use", "enum": ["usual", "official", "temp", "nickname", "anonymous", "old", "maiden"], "term": {"description": "Identifies the purpose for this name.. http://hl7.org/fhir/name-use", "termDef": {"cde_id": "http://hl7.org/fhir/name-use", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/name-use", "term_url": "http://hl7.org/fhir/name-use"}}}, "name_family": {"description": "Family name (often called 'Surname').", "type": ["string", "null"]}, "name_given": {"description": "Given names (not always 'first'). Includes middle names.", "type": ["string", "null"]}, "name_prefix": {"description": "Parts that come before the name.", "type": ["string", "null"]}, "name_suffix": {"description": "Parts that come after the name.", "type": ["string", "null"]}, "name_use": {"description": "Identifies the purpose for this name.. http://hl7.org/fhir/name-use", "enum": ["usual", "official", "temp", "nickname", "anonymous", "old", "maiden"], "term": {"description": "Identifies the purpose for this name.. http://hl7.org/fhir/name-use", "termDef": {"cde_id": "http://hl7.org/fhir/name-use", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/name-use", "term_url": "http://hl7.org/fhir/name-use"}}}, "patient_birthPlace_city": {"description": "The registered place of birth of the patient. A sytem may use the address.text if they don't store the birthPlace address in discrete elements.", "type": ["string", "null"]}, "patient_birthPlace_country": {"description": "The registered place of birth of the patient. A sytem may use the address.text if they don't store the birthPlace address in discrete elements.", "type": ["string", "null"]}, "patient_birthPlace_state": {"description": "The registered place of birth of the patient. A sytem may use the address.text if they don't store the birthPlace address in discrete elements.", "type": ["string", "null"]}, "patient_mothersMaidenName": {"description": "Mother's maiden (unmarried) name, commonly collected to help verify patient identity.", "type": ["string", "null"]}, "project_id": {"$ref": "_definitions.yaml#/project_id"}, "quality_adjusted_life_years": {"description": "Quality Adjusted Life Years as defined in the literature and summarized at .", "type": ["number", "null"]}, "resourceType": {"description": "One of the resource types defined as part of this version of FHIR", "type": ["string", "null"]}, "state": {"$ref": "_definitions.yaml#/state"}, "submitter_id": {"type": ["string", "null"]}, "telecom_system": {"description": "Telecommunications form for contact point - what communications system is required to make use of the contact.. http://hl7.org/fhir/contact-point-system", "enum": ["phone", "fax", "email", "pager", "url", "sms", "other"], "term": {"description": "Telecommunications form for contact point - what communications system is required to make use of the contact.. http://hl7.org/fhir/contact-point-system", "termDef": {"cde_id": "http://hl7.org/fhir/contact-point-system", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/contact-point-system", "term_url": "http://hl7.org/fhir/contact-point-system"}}}, "telecom_use": {"description": "Identifies the purpose for the contact point.. http://hl7.org/fhir/contact-point-use", "enum": ["home", "work", "temp", "old", "mobile"], "term": {"description": "Identifies the purpose for the contact point.. http://hl7.org/fhir/contact-point-use", "termDef": {"cde_id": "http://hl7.org/fhir/contact-point-use", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/contact-point-use", "term_url": "http://hl7.org/fhir/contact-point-use"}}}, "telecom_value": {"description": "The actual contact point details.", "type": ["string", "null"]}, "text_div": {"description": "Limited xhtml content.", "type": ["string"]}, "text_status": {"description": "The status of the narrative - whether it's entirely generated (from just the defined data or the extensions too), or whether a human authored it and it may contain additional data.. http://hl7.org/fhir/narrative-status", "enum": ["generated", "extensions", "additional", "empty"], "term": {"description": "The status of the narrative - whether it's entirely generated (from just the defined data or the extensions too), or whether a human authored it and it may contain additional data.. http://hl7.org/fhir/narrative-status", "termDef": {"cde_id": "http://hl7.org/fhir/narrative-status", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/narrative-status", "term_url": "http://hl7.org/fhir/narrative-status"}}}, "type": {"type": "string"}, "updated_datetime": {"$ref": "_definitions.yaml#/datetime"}, "us_core_birthsex": {"description": "A code classifying the person's sex assigned at birth as specified by the [Office of the National Coordinator for Health IT (ONC)](https://www.healthit.gov/newsroom/about-onc). This extension aligns with the C-CDA Birth Sex Observation (LOINC 76689-9).", "type": ["string", "null"]}, "us_core_ethnicity_ombCategory": {"description": "Concepts classifying the person into a named category of humans sharing common history, traits, geographical origin or nationality. The ethnicity codes used to represent these concepts are based upon the [CDC ethnicity and Ethnicity Code Set Version 1.0](http://www.cdc.gov/phin/resources/vocabulary/index.html) which includes over 900 concepts for representing race and ethnicity of which 43 reference ethnicity. The ethnicity concepts are grouped by and pre-mapped to the 2 OMB ethnicity categories: - Hispanic or Latino - Not Hispanic or Latino.\nThe 2 ethnicity category codes according to the [OMB Standards for Maintaining, Collecting, and Presenting Federal Data on Race and Ethnicity, Statistical Policy Directive No. 15, as revised, October 30, 1997](https://www.govinfo.gov/content/pkg/FR-1997-10-30/pdf/97-28653.pdf).. http://hl7.org/fhir/us/core/ValueSet/omb-ethnicity-category", "enum": ["2135-2", "2186-5", "ASKU", "UNK"], "term": {"description": "Concepts classifying the person into a named category of humans sharing common history, traits, geographical origin or nationality. The ethnicity codes used to represent these concepts are based upon the [CDC ethnicity and Ethnicity Code Set Version 1.0](http://www.cdc.gov/phin/resources/vocabulary/index.html) which includes over 900 concepts for representing race and ethnicity of which 43 reference ethnicity. The ethnicity concepts are grouped by and pre-mapped to the 2 OMB ethnicity categories: - Hispanic or Latino - Not Hispanic or Latino.\nThe 2 ethnicity category codes according to the [OMB Standards for Maintaining, Collecting, and Presenting Federal Data on Race and Ethnicity, Statistical Policy Directive No. 15, as revised, October 30, 1997](https://www.govinfo.gov/content/pkg/FR-1997-10-30/pdf/97-28653.pdf).. http://hl7.org/fhir/us/core/ValueSet/omb-ethnicity-category", "termDef": {"cde_id": "http://hl7.org/fhir/us/core/ValueSet/omb-ethnicity-category", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/us/core/ValueSet/omb-ethnicity-category", "term_url": "http://hl7.org/fhir/us/core/ValueSet/omb-ethnicity-category"}}}, "us_core_ethnicity_text": {"description": "Concepts classifying the person into a named category of humans sharing common history, traits, geographical origin or nationality. The ethnicity codes used to represent these concepts are based upon the [CDC ethnicity and Ethnicity Code Set Version 1.0](http://www.cdc.gov/phin/resources/vocabulary/index.html) which includes over 900 concepts for representing race and ethnicity of which 43 reference ethnicity. The ethnicity concepts are grouped by and pre-mapped to the 2 OMB ethnicity categories: - Hispanic or Latino - Not Hispanic or Latino.\nPlain text representation of the ethnicity concept(s).", "type": ["string", "null"]}, "us_core_race_ombCategory": {"description": "Concepts classifying the person into a named category of humans sharing common history, traits, geographical origin or nationality. The race codes used to represent these concepts are based upon the [CDC Race and Ethnicity Code Set Version 1.0](http://www.cdc.gov/phin/resources/vocabulary/index.html) which includes over 900 concepts for representing race and ethnicity of which 921 reference race. The race concepts are grouped by and pre-mapped to the 5 OMB race categories:\n\n - American Indian or Alaska Native\n - Asian\n - Black or African American\n - Native Hawaiian or Other Pacific Islander\n - White.\nThe 5 race category codes according to the [OMB Standards for Maintaining, Collecting, and Presenting Federal Data on Race and Ethnicity, Statistical Policy Directive No. 15, as revised, October 30, 1997](https://www.govinfo.gov/content/pkg/FR-1997-10-30/pdf/97-28653.pdf).. http://hl7.org/fhir/us/core/ValueSet/omb-race-category", "enum": ["1002-5", "2028-9", "2054-5", "2076-8", "2106-3", "2131-1", "ASKU", "UNK"], "term": {"description": "Concepts classifying the person into a named category of humans sharing common history, traits, geographical origin or nationality. The race codes used to represent these concepts are based upon the [CDC Race and Ethnicity Code Set Version 1.0](http://www.cdc.gov/phin/resources/vocabulary/index.html) which includes over 900 concepts for representing race and ethnicity of which 921 reference race. The race concepts are grouped by and pre-mapped to the 5 OMB race categories:\n\n - American Indian or Alaska Native\n - Asian\n - Black or African American\n - Native Hawaiian or Other Pacific Islander\n - White.\nThe 5 race category codes according to the [OMB Standards for Maintaining, Collecting, and Presenting Federal Data on Race and Ethnicity, Statistical Policy Directive No. 15, as revised, October 30, 1997](https://www.govinfo.gov/content/pkg/FR-1997-10-30/pdf/97-28653.pdf).. http://hl7.org/fhir/us/core/ValueSet/omb-race-category", "termDef": {"cde_id": "http://hl7.org/fhir/us/core/ValueSet/omb-race-category", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/us/core/ValueSet/omb-race-category", "term_url": "http://hl7.org/fhir/us/core/ValueSet/omb-race-category"}}}, "us_core_race_text": {"description": "Concepts classifying the person into a named category of humans sharing common history, traits, geographical origin or nationality. The race codes used to represent these concepts are based upon the [CDC Race and Ethnicity Code Set Version 1.0](http://www.cdc.gov/phin/resources/vocabulary/index.html) which includes over 900 concepts for representing race and ethnicity of which 921 reference race. The race concepts are grouped by and pre-mapped to the 5 OMB race categories:\n\n - American Indian or Alaska Native\n - Asian\n - Black or African American\n - Native Hawaiian or Other Pacific Islander\n - White.\nPlain text representation of the race concept(s).", "type": ["string", "null"]}}, "required": ["submitter_id", "type", "text_status", "text_div"], "submittable": true, "systemProperties": ["id", "project_id", "created_datetime", "updated_datetime", "state"], "title": "Patient", "type": "object", "uniqueKeys": [["id"], ["project_id", "submitter_id"]], "validators": null}, "Practitioner.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "additionalProperties": false, "category": "Administrative", "description": " A person with a formal responsibility in the provisioning of healthcare or\n related services.\n \n A person who is directly or indirectly involved in the provisioning of\n healthcare.\n ", "id": "Practitioner", "links": [], "namespace": "http://hl7.org/fhir", "program": "*", "project": "*", "properties": {"created_datetime": {"$ref": "_definitions.yaml#/datetime"}, "gender": {"description": "Administrative Gender - the gender that the person is considered to have for administration and record keeping purposes.. http://hl7.org/fhir/administrative-gender", "enum": ["male", "female", "other", "unknown"], "term": {"description": "Administrative Gender - the gender that the person is considered to have for administration and record keeping purposes.. http://hl7.org/fhir/administrative-gender", "termDef": {"cde_id": "http://hl7.org/fhir/administrative-gender", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/administrative-gender", "term_url": "http://hl7.org/fhir/administrative-gender"}}}, "id": {"description": "", "type": ["string", "null"]}, "identifier_name": {"description": "An identifier for the person as this agent.", "type": ["string", "null"]}, "identifier_synthea": {"description": "An identifier for the person as this agent.", "type": ["string", "null"]}, "name_text": {"description": "Text representation of the full name.", "type": ["string", "null"]}, "project_id": {"$ref": "_definitions.yaml#/project_id"}, "qualification_code_coding_synthea": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "qualification_code_text": {"description": "Plain text representation of the concept.", "type": ["string", "null"]}, "qualification_issuer_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "resourceType": {"description": "One of the resource types defined as part of this version of FHIR", "type": ["string", "null"]}, "state": {"$ref": "_definitions.yaml#/state"}, "submitter_id": {"type": ["string", "null"]}, "type": {"type": "string"}, "updated_datetime": {"$ref": "_definitions.yaml#/datetime"}}, "required": ["submitter_id", "type"], "submittable": true, "systemProperties": ["id", "project_id", "created_datetime", "updated_datetime", "state"], "title": "Practitioner", "type": "object", "uniqueKeys": [["id"], ["project_id", "submitter_id"]], "validators": null}, "MedicationRequest.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "additionalProperties": false, "category": "Clinical", "description": " Ordering of medication for patient or group.\n \n An order or request for both supply of the medication and the instructions\n for administration of the medication to a patient. The resource is called\n \"MedicationRequest\" rather than \"MedicationPrescription\" or\n \"MedicationOrder\" to generalize the use across inpatient and outpatient\n settings, including care plans, etc., and to harmonize with workflow\n patterns.\n ", "id": "MedicationRequest", "links": [{"backref": "MedicationRequests", "label": "Patients", "multiplicity": "many_to_many", "name": "Patients", "required": true, "target_type": "Patient"}, {"backref": "MedicationRequests", "label": "Encounters", "multiplicity": "many_to_many", "name": "Encounters", "required": false, "target_type": "Encounter"}, {"backref": "MedicationRequests", "label": "Medications", "multiplicity": "many_to_many", "name": "Medications", "required": false, "target_type": "Medication"}], "namespace": "http://hl7.org/fhir", "program": "*", "project": "*", "properties": {"authoredOn": {"description": "When request was initially authored.", "type": ["string", "null"]}, "created_datetime": {"$ref": "_definitions.yaml#/datetime"}, "dosageInstruction_additionalInstruction_coding_sct": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "dosageInstruction_additionalInstruction_text": {"description": "Plain text representation of the concept.", "type": ["string", "null"]}, "dosageInstruction_asNeededBoolean": {"description": "Take \"as needed\" (for x).", "type": ["boolean", "null"]}, "dosageInstruction_doseAndRate_doseQuantity_value": {"description": "Numerical value (with implicit precision).", "type": ["number", "null"]}, "dosageInstruction_doseAndRate_type_coding_dose_rate_type": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "dosageInstruction_sequence": {"description": "The order of the dosage instructions.", "type": ["number", "null"]}, "dosageInstruction_text": {"description": "Free text dosage instructions e.g. SIG.", "type": ["string", "null"]}, "dosageInstruction_timing_repeat_frequency": {"description": "Event occurs frequency times per period.", "type": ["number", "null"]}, "dosageInstruction_timing_repeat_period": {"description": "Event occurs frequency times per period.", "type": ["number", "null"]}, "dosageInstruction_timing_repeat_periodUnit": {"description": "s | min | h | d | wk | mo | a - unit of time (UCUM).", "type": ["string", "null"]}, "encounter_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "id": {"description": "", "type": ["string", "null"]}, "intent": {"description": "Whether the request is a proposal, plan, or an original order.. http://hl7.org/fhir/CodeSystem/medicationrequest-intent", "enum": ["proposal", "plan", "order", "original-order", "reflex-order", "filler-order", "instance-order", "option"], "term": {"description": "Whether the request is a proposal, plan, or an original order.. http://hl7.org/fhir/CodeSystem/medicationrequest-intent", "termDef": {"cde_id": "http://hl7.org/fhir/CodeSystem/medicationrequest-intent", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/CodeSystem/medicationrequest-intent", "term_url": "http://hl7.org/fhir/CodeSystem/medicationrequest-intent"}}}, "medicationCodeableConcept_coding_rxnorm": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "medicationCodeableConcept_text": {"description": "Plain text representation of the concept.", "type": ["string", "null"]}, "medicationReference_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "meta_profile": {"description": "Profiles this resource claims to conform to.", "type": ["string", "null"]}, "project_id": {"$ref": "_definitions.yaml#/project_id"}, "reasonReference_0_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "reasonReference_1_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "reasonReference_2_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "reasonReference_3_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "reasonReference_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "requester_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "requester_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "resourceType": {"description": "One of the resource types defined as part of this version of FHIR", "type": ["string", "null"]}, "state": {"$ref": "_definitions.yaml#/state"}, "status": {"description": "A code specifying the current state of the order. Generally, this will be active or completed state.. http://hl7.org/fhir/CodeSystem/medicationrequest-status", "enum": ["active", "on-hold", "cancelled", "completed", "entered-in-error", "stopped", "draft", "unknown"], "term": {"description": "A code specifying the current state of the order. Generally, this will be active or completed state.. http://hl7.org/fhir/CodeSystem/medicationrequest-status", "termDef": {"cde_id": "http://hl7.org/fhir/CodeSystem/medicationrequest-status", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/CodeSystem/medicationrequest-status", "term_url": "http://hl7.org/fhir/CodeSystem/medicationrequest-status"}}}, "subject_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "submitter_id": {"type": ["string", "null"]}, "type": {"type": "string"}, "updated_datetime": {"$ref": "_definitions.yaml#/datetime"}}, "required": ["submitter_id", "type", "status", "intent"], "submittable": true, "systemProperties": ["id", "project_id", "created_datetime", "updated_datetime", "state"], "title": "MedicationRequest", "type": "object", "uniqueKeys": [["id"], ["project_id", "submitter_id"]], "validators": null}, "_program.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "id": "program", "title": "Program", "type": "object", "category": "administrative", "program": "*", "project": "*", "description": "A broad framework of goals to be achieved. (NCIt C52647)\n", "additionalProperties": false, "submittable": false, "validators": null, "systemProperties": ["id"], "required": ["name", "dbgap_accession_number"], "uniqueKeys": [["id"], ["name"]], "links": [], "properties": {"type": {"type": "string"}, "id": {"$ref": "_definitions.yaml#/UUID", "systemAlias": "node_id"}, "name": {"type": "string", "description": "Full name/title of the program."}, "dbgap_accession_number": {"type": "string", "description": "The dbgap accession number provided for the program."}}}, "_project.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "id": "project", "title": "Project", "type": "object", "program": "*", "project": "*", "category": "administrative", "description": "Any specifically defined piece of work that is undertaken or attempted to meet a single requirement. (NCIt C47885)\n", "additionalProperties": false, "submittable": true, "validators": null, "systemProperties": ["id", "state", "released", "releasable", "intended_release_date"], "required": ["code", "name", "dbgap_accession_number", "programs"], "uniqueKeys": [["id"], ["code"]], "links": [{"name": "programs", "backref": "projects", "label": "member_of", "target_type": "program", "multiplicity": "many_to_one", "required": true}], "constraints": null, "properties": {"type": {"type": "string"}, "id": {"$ref": "_definitions.yaml#/UUID", "systemAlias": "node_id", "description": "UUID for the project."}, "name": {"type": "string", "description": "Display name/brief description for the project."}, "code": {"type": "string", "description": "Unique identifier for the project."}, "investigator_name": {"description": "Name of the principal investigator for the project.", "type": "string"}, "investigator_affiliation": {"description": "The investigator's affiliation with respect to a research institution.", "type": "string"}, "date_collected": {"description": "The date or date range in which the project data was collected.", "type": "string"}, "availability_type": {"description": "Is the project open or restricted?", "enum": ["Open", "Restricted"]}, "availability_mechanism": {"description": "Mechanism by which the project will be made avilable.", "type": "string"}, "support_source": {"description": "The name of source providing support/grant resources.", "type": "string"}, "support_id": {"description": "The ID of the source providing support/grant resources.", "type": "string"}, "programs": {"$ref": "_definitions.yaml#/to_one", "description": "Indicates that the project is logically part of the indicated project.\n"}, "state": {"description": "The possible states a project can be in. All but `open` are\nequivalent to some type of locked state.\n", "default": "open", "enum": ["open", "review", "submitted", "processing", "closed", "legacy"]}, "released": {"description": "To release a project is to tell the GDC to include all submitted\nentities in the next GDC index.\n", "default": false, "type": "boolean"}, "releasable": {"description": "A project can only be released by the user when `releasable` is true.\n", "default": false, "type": "boolean"}, "intended_release_date": {"description": "Tracks a Project's intended release date.", "type": "string", "format": "date-time"}, "dbgap_accession_number": {"type": "string", "description": "The dbgap accession number provided for the project."}}}, "_definitions.yaml": {"id": "_definitions", "UUID": {"term": {"$ref": "_terms.yaml#/UUID"}, "type": "string", "pattern": "^[a-fA-F0-9]{8}-[a-fA-F0-9]{4}-[a-fA-F0-9]{4}-[a-fA-F0-9]{4}-[a-fA-F0-9]{12}$"}, "parent_uuids": {"type": "array", "minItems": 1, "items": {"$ref": "#/UUID"}, "uniqueItems": true}, "foreign_key_project": {"type": "object", "additionalProperties": true, "properties": {"id": {"$ref": "#/UUID"}, "code": {"type": "string"}}}, "to_one_project": {"anyOf": [{"type": "array", "items": {"$ref": "#/foreign_key_project", "minItems": 1, "maxItems": 1}}, {"$ref": "#/foreign_key_project"}]}, "to_many_project": {"anyOf": [{"type": "array", "items": {"$ref": "#/foreign_key_project", "minItems": 1}}, {"$ref": "#/foreign_key_project"}]}, "foreign_key": {"type": "object", "additionalProperties": true, "properties": {"id": {"$ref": "#/UUID"}, "submitter_id": {"type": "string"}}}, "to_one": {"anyOf": [{"type": "array", "items": {"$ref": "#/foreign_key", "minItems": 1, "maxItems": 1}}, {"$ref": "#/foreign_key"}]}, "to_many": {"anyOf": [{"type": "array", "items": {"$ref": "#/foreign_key", "minItems": 1}}, {"$ref": "#/foreign_key"}]}, "datetime": {"oneOf": [{"type": "string", "format": "date-time"}, {"type": "null"}], "term": {"$ref": "_terms.yaml#/datetime"}}, "file_name": {"type": "string", "term": {"$ref": "_terms.yaml#/file_name"}}, "file_size": {"type": "integer", "term": {"$ref": "_terms.yaml#/file_size"}}, "file_format": {"type": "string", "term": {"$ref": "_terms.yaml#/file_format"}}, "ga4gh_drs_uri": {"type": "string", "term": {"$ref": "_terms.yaml#/ga4gh_drs_uri"}}, "md5sum": {"type": "string", "term": {"$ref": "_terms.yaml#/md5sum"}, "pattern": "^[a-f0-9]{32}$"}, "object_id": {"type": "string", "description": "The GUID of the object in the index service."}, "release_state": {"description": "Release state of an entity.", "default": "unreleased", "enum": ["unreleased", "released", "redacted"]}, "data_bundle_state": {"description": "State of a data bundle.", "default": "submitted", "enum": ["submitted", "validated", "error", "released", "suppressed", "redacted"]}, "data_file_error_type": {"term": {"$ref": "_terms.yaml#/data_file_error_type"}, "enum": ["file_size", "file_format", "md5sum"]}, "state": {"term": {"$ref": "_terms.yaml#/state"}, "default": "validated", "downloadable": ["uploaded", "md5summed", "validating", "validated", "error", "invalid", "released"], "public": ["live"], "oneOf": [{"enum": ["uploading", "uploaded", "md5summing", "md5summed", "validating", "error", "invalid", "suppressed", "redacted", "live"]}, {"enum": ["validated", "submitted", "released"]}]}, "file_state": {"term": {"$ref": "_terms.yaml#/file_state"}, "default": "registered", "enum": ["registered", "uploading", "uploaded", "validating", "validated", "submitted", "processing", "processed", "released", "error"]}, "qc_metrics_state": {"term": {"$ref": "_terms.yaml#/qc_metric_state"}, "enum": ["FAIL", "PASS", "WARN"]}, "project_id": {"type": "string", "term": {"$ref": "_terms.yaml#/project_id"}}, "data_file_properties": {"$ref": "#/ubiquitous_properties", "file_name": {"$ref": "#/file_name"}, "file_size": {"$ref": "#/file_size"}, "file_format": {"$ref": "#/file_format"}, "md5sum": {"$ref": "#/md5sum"}, "object_id": {"$ref": "#/object_id"}, "file_state": {"$ref": "#/file_state"}, "error_type": {"$ref": "#/data_file_error_type"}, "ga4gh_drs_uri": {"$ref": "#/ga4gh_drs_uri"}}, "workflow_properties": {"$ref": "#/ubiquitous_properties", "workflow_link": {"description": "Link to Github hash for the CWL workflow used.", "type": "string"}, "workflow_version": {"description": "Major version for a GDC workflow.", "type": "string"}, "workflow_start_datetime": {"$ref": "#/datetime"}, "workflow_end_datetime": {"$ref": "#/datetime"}}, "ubiquitous_properties": {"type": {"type": "string"}, "id": {"$ref": "#/UUID", "systemAlias": "node_id"}, "submitter_id": {"type": ["string"], "description": "A project-specific identifier for a node. This property is the calling card/nickname/alias for a unit of submission. It can be used in place of the UUID for identifying or recalling a node.\n"}, "state": {"$ref": "#/state"}, "project_id": {"$ref": "#/project_id"}, "created_datetime": {"$ref": "#/datetime"}, "updated_datetime": {"$ref": "#/datetime"}}}, "DiagnosticReport.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "additionalProperties": false, "category": "Clinical", "description": " A Diagnostic report - a combination of request information, atomic results,\n images, interpretation, as well as formatted reports.\n \n The findings and interpretation of diagnostic tests performed on patients,\n groups of patients, devices, and locations, and/or specimens derived from\n these. The report includes clinical context such as requesting and provider\n information, and some mix of atomic results, images, textual and coded\n interpretations, and formatted representation of diagnostic reports.\n ", "id": "DiagnosticReport", "links": [{"backref": "DiagnosticReports", "label": "Patients", "multiplicity": "many_to_many", "name": "Patients", "required": true, "target_type": "Patient"}, {"backref": "DiagnosticReports", "label": "Encounters", "multiplicity": "many_to_many", "name": "Encounters", "required": false, "target_type": "Encounter"}, {"backref": "DiagnosticReports", "label": "Organizations", "multiplicity": "many_to_many", "name": "Organizations", "required": false, "target_type": "Organization"}, {"backref": "DiagnosticReports", "label": "Practitioners", "multiplicity": "many_to_many", "name": "Practitioners", "required": false, "target_type": "Practitioner"}], "namespace": "http://hl7.org/fhir", "program": "*", "project": "*", "properties": {"category_coding_0_code": {"description": "Symbol in syntax defined by the system.", "type": ["string", "null"]}, "category_coding_0_display": {"description": "Representation defined by the system.", "type": ["string", "null"]}, "category_coding_0_system": {"description": "Identity of the terminology system.", "type": ["string", "null"]}, "category_coding_1_code": {"description": "Symbol in syntax defined by the system.", "type": ["string", "null"]}, "category_coding_1_display": {"description": "Representation defined by the system.", "type": ["string", "null"]}, "category_coding_1_system": {"description": "Identity of the terminology system.", "type": ["string", "null"]}, "category_coding_v2_0074": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "code_coding_0_code": {"description": "Symbol in syntax defined by the system.", "type": ["string", "null"]}, "code_coding_0_display": {"description": "Representation defined by the system.", "type": ["string", "null"]}, "code_coding_0_system": {"description": "Identity of the terminology system.", "type": ["string", "null"]}, "code_coding_1_code": {"description": "Symbol in syntax defined by the system.", "type": ["string", "null"]}, "code_coding_1_display": {"description": "Representation defined by the system.", "type": ["string", "null"]}, "code_coding_1_system": {"description": "Identity of the terminology system.", "type": ["string", "null"]}, "code_coding_loinc_org": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "code_text": {"description": "Plain text representation of the concept.", "type": ["string", "null"]}, "created_datetime": {"$ref": "_definitions.yaml#/datetime"}, "effectiveDateTime": {"description": "Clinically relevant time/time-period for report.", "type": ["string", "null"]}, "encounter_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "id": {"description": "", "type": ["string", "null"]}, "issued": {"description": "DateTime this version was made.", "type": ["string", "null"]}, "meta_profile": {"description": "Profiles this resource claims to conform to.", "type": ["string", "null"]}, "performer_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "performer_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "presentedForm_contentType": {"description": "Mime type of the content, with charset etc..", "type": ["string", "null"]}, "presentedForm_data": {"description": "Data inline, base64ed.", "type": ["string", "null"]}, "presentedForm_url": {"description": "Uri where the data can be found.", "type": ["string", "null"]}, "project_id": {"$ref": "_definitions.yaml#/project_id"}, "resourceType": {"description": "One of the resource types defined as part of this version of FHIR", "type": ["string", "null"]}, "result_0_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_0_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_10_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_10_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_11_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_11_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_12_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_12_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_13_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_13_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_14_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_14_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_15_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_15_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_16_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_16_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_17_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_17_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_18_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_18_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_19_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_19_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_1_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_1_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_20_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_20_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_21_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_21_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_22_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_22_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_23_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_23_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_24_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_24_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_25_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_25_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_26_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_26_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_27_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_27_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_28_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_28_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_29_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_29_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_2_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_2_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_30_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_30_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_31_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_31_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_32_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_32_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_33_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_33_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_34_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_34_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_35_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_35_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_36_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_36_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_37_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_37_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_38_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_38_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_39_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_39_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_3_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_3_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_40_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_40_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_41_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_41_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_42_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_42_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_43_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_43_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_44_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_44_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_45_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_45_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_46_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_46_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_47_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_47_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_48_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_48_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_49_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_49_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_4_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_4_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_50_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_50_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_5_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_5_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_6_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_6_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_7_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_7_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_8_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_8_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_9_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_9_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "result_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "result_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "specimen_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "state": {"$ref": "_definitions.yaml#/state"}, "status": {"description": "The status of the diagnostic report.. http://hl7.org/fhir/diagnostic-report-status", "enum": ["registered", "partial", "preliminary", "final", "amended", "corrected", "appended", "cancelled", "entered-in-error", "unknown"], "term": {"description": "The status of the diagnostic report.. http://hl7.org/fhir/diagnostic-report-status", "termDef": {"cde_id": "http://hl7.org/fhir/diagnostic-report-status", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/diagnostic-report-status", "term_url": "http://hl7.org/fhir/diagnostic-report-status"}}}, "subject_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "submitter_id": {"type": ["string", "null"]}, "type": {"type": "string"}, "updated_datetime": {"$ref": "_definitions.yaml#/datetime"}}, "required": ["submitter_id", "type", "status"], "submittable": true, "systemProperties": ["id", "project_id", "created_datetime", "updated_datetime", "state"], "title": "DiagnosticReport", "type": "object", "uniqueKeys": [["id"], ["project_id", "submitter_id"]], "validators": null}, "_settings.yaml": {"enable_case_cache": false, "_dict_commit": "520a25999fd183f6c5b7ddef2980f3e839517da5", "_dict_version": "0.2.1-9-g520a259"}, "Task.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "additionalProperties": false, "category": "Analysis", "description": " A task to be performed.\n ", "id": "Task", "links": [{"backref": "Tasks", "label": "Patients", "multiplicity": "many_to_many", "name": "Patients", "required": true, "target_type": "Patient"}, {"backref": "Tasks", "label": "Specimen", "multiplicity": "many_to_many", "name": "Specimen", "required": true, "target_type": "Specimen"}, {"backref": "Tasks", "label": "DocumentReferences", "multiplicity": "many_to_many", "name": "DocumentReferences", "required": true, "target_type": "DocumentReference"}], "namespace": "http://hl7.org/fhir", "program": "*", "project": "*", "properties": {"created_datetime": {"$ref": "_definitions.yaml#/datetime"}, "focus_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "for_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "id": {"description": "", "type": ["string", "null"]}, "input_type_coding_": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "input_valueReference_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "intent": {"description": "unknown | proposal | plan | order | original-order | reflex-order | filler-order | instance-order | option.", "type": ["string"]}, "output_type_coding_": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "output_valueReference_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "project_id": {"$ref": "_definitions.yaml#/project_id"}, "resourceType": {"description": "One of the resource types defined as part of this version of FHIR", "type": ["string", "null"]}, "state": {"$ref": "_definitions.yaml#/state"}, "status": {"description": "The current status of the task.. http://hl7.org/fhir/task-status", "enum": ["draft", "requested", "received", "accepted", "rejected", "ready", "cancelled", "in-progress", "on-hold", "failed", "completed", "entered-in-error"], "term": {"description": "The current status of the task.. http://hl7.org/fhir/task-status", "termDef": {"cde_id": "http://hl7.org/fhir/task-status", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/task-status", "term_url": "http://hl7.org/fhir/task-status"}}}, "submitter_id": {"type": ["string", "null"]}, "text_div": {"description": "Limited xhtml content.", "type": ["string"]}, "text_status": {"description": "The status of the narrative - whether it's entirely generated (from just the defined data or the extensions too), or whether a human authored it and it may contain additional data.. http://hl7.org/fhir/narrative-status", "enum": ["generated", "extensions", "additional", "empty"], "term": {"description": "The status of the narrative - whether it's entirely generated (from just the defined data or the extensions too), or whether a human authored it and it may contain additional data.. http://hl7.org/fhir/narrative-status", "termDef": {"cde_id": "http://hl7.org/fhir/narrative-status", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/narrative-status", "term_url": "http://hl7.org/fhir/narrative-status"}}}, "type": {"type": "string"}, "updated_datetime": {"$ref": "_definitions.yaml#/datetime"}}, "required": ["submitter_id", "type", "text_status", "text_div", "status", "intent"], "submittable": true, "systemProperties": ["id", "project_id", "created_datetime", "updated_datetime", "state"], "title": "Task", "type": "object", "uniqueKeys": [["id"], ["project_id", "submitter_id"]], "validators": null}, "Condition.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "additionalProperties": false, "category": "Clinical", "description": " Detailed information about conditions, problems or diagnoses.\n \n A clinical condition, problem, diagnosis, or other event, situation, issue,\n or clinical concept that has risen to a level of concern.\n ", "id": "Condition", "links": [{"backref": "Conditions", "label": "Patients", "multiplicity": "many_to_many", "name": "Patients", "required": true, "target_type": "Patient"}, {"backref": "Conditions", "label": "Encounters", "multiplicity": "many_to_many", "name": "Encounters", "required": true, "target_type": "Encounter"}], "namespace": "http://hl7.org/fhir", "program": "*", "project": "*", "properties": {"abatementDateTime": {"description": "When in resolution/remission.", "type": ["string", "null"]}, "category_coding_condition_category": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "clinicalStatus_coding_condition_clinical": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "code_coding_sct": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "code_text": {"description": "Plain text representation of the concept.", "type": ["string", "null"]}, "created_datetime": {"$ref": "_definitions.yaml#/datetime"}, "encounter_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "id": {"description": "", "type": ["string", "null"]}, "meta_profile": {"description": "Profiles this resource claims to conform to.", "type": ["string", "null"]}, "onsetDateTime": {"description": "Estimated or actual date, date-time, or age.", "type": ["string", "null"]}, "project_id": {"$ref": "_definitions.yaml#/project_id"}, "recordedDate": {"description": "Date record was first recorded.", "type": ["string", "null"]}, "resourceType": {"description": "One of the resource types defined as part of this version of FHIR", "type": ["string", "null"]}, "state": {"$ref": "_definitions.yaml#/state"}, "subject_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "submitter_id": {"type": ["string", "null"]}, "type": {"type": "string"}, "updated_datetime": {"$ref": "_definitions.yaml#/datetime"}, "verificationStatus_coding_condition_ver_status": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}}, "required": ["submitter_id", "type"], "submittable": true, "systemProperties": ["id", "project_id", "created_datetime", "updated_datetime", "state"], "title": "Condition", "type": "object", "uniqueKeys": [["id"], ["project_id", "submitter_id"]], "validators": null}, "Location.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "additionalProperties": false, "category": "Administrative", "description": " Details and position information for a physical place.\n \n Details and position information for a physical place where services are\n provided and resources and participants may be stored, found, contained, or\n accommodated.\n ", "id": "Location", "links": [{"backref": "Locations", "label": "Organizations", "multiplicity": "many_to_many", "name": "Organizations", "required": true, "target_type": "Organization"}], "namespace": "http://hl7.org/fhir", "program": "*", "project": "*", "properties": {"address_city": {"description": "Name of city, town etc..", "type": ["string", "null"]}, "address_line": {"description": "Street name, number, direction & P.O. Box etc..", "type": ["string", "null"]}, "address_postalCode": {"description": "Postal code for area.", "type": ["string", "null"]}, "address_state": {"description": "Sub-unit of country (abbreviations ok).", "type": ["string", "null"]}, "created_datetime": {"$ref": "_definitions.yaml#/datetime"}, "id": {"description": "", "type": ["string", "null"]}, "identifier_name": {"description": "Unique code or number identifying the location to its users.", "type": ["string", "null"]}, "managingOrganization_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "position_latitude": {"description": "Latitude with WGS84 datum.", "type": ["number"]}, "position_longitude": {"description": "Longitude with WGS84 datum.", "type": ["number"]}, "project_id": {"$ref": "_definitions.yaml#/project_id"}, "resourceType": {"description": "One of the resource types defined as part of this version of FHIR", "type": ["string", "null"]}, "state": {"$ref": "_definitions.yaml#/state"}, "submitter_id": {"type": ["string", "null"]}, "type": {"type": "string"}, "updated_datetime": {"$ref": "_definitions.yaml#/datetime"}}, "required": ["submitter_id", "type", "position_longitude", "position_latitude"], "submittable": true, "systemProperties": ["id", "project_id", "created_datetime", "updated_datetime", "state"], "title": "Location", "type": "object", "uniqueKeys": [["id"], ["project_id", "submitter_id"]], "validators": null}, "Encounter.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "additionalProperties": false, "category": "Clinical", "description": " An interaction during which services are provided to the patient.\n \n An interaction between a patient and healthcare provider(s) for the purpose\n of providing healthcare service(s) or assessing the health status of a\n patient.\n ", "id": "Encounter", "links": [{"backref": "Encounters", "label": "Patients", "multiplicity": "many_to_many", "name": "Patients", "required": true, "target_type": "Patient"}], "namespace": "http://hl7.org/fhir", "program": "*", "project": "*", "properties": {"class_v3_ActCode": {"description": "Classification of patient encounter.", "type": ["string"]}, "created_datetime": {"$ref": "_definitions.yaml#/datetime"}, "hospitalization_dischargeDisposition_coding_patient_discharge": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "hospitalization_dischargeDisposition_text": {"description": "Plain text representation of the concept.", "type": ["string", "null"]}, "id": {"description": "", "type": ["string", "null"]}, "identifier_synthea": {"description": "Identifier(s) by which this encounter is known.", "type": ["string", "null"]}, "location_location_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "location_location_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "meta_profile": {"description": "Profiles this resource claims to conform to.", "type": ["string", "null"]}, "participant_individual_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "participant_individual_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "participant_period_end": {"description": "End time with inclusive boundary, if not ongoing.", "type": ["string", "null"]}, "participant_period_start": {"description": "Starting time with inclusive boundary.", "type": ["string", "null"]}, "participant_type_coding_v3_ParticipationType": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "participant_type_text": {"description": "Plain text representation of the concept.", "type": ["string", "null"]}, "period_end": {"description": "End time with inclusive boundary, if not ongoing.", "type": ["string", "null"]}, "period_start": {"description": "Starting time with inclusive boundary.", "type": ["string", "null"]}, "project_id": {"$ref": "_definitions.yaml#/project_id"}, "reasonCode_coding_sct": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "resourceType": {"description": "One of the resource types defined as part of this version of FHIR", "type": ["string", "null"]}, "serviceProvider_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "serviceProvider_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "state": {"$ref": "_definitions.yaml#/state"}, "status": {"description": "None. http://hl7.org/fhir/encounter-status", "enum": ["planned", "arrived", "triaged", "in-progress", "onleave", "finished", "cancelled", "entered-in-error", "unknown"], "term": {"description": "None. http://hl7.org/fhir/encounter-status", "termDef": {"cde_id": "http://hl7.org/fhir/encounter-status", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/encounter-status", "term_url": "http://hl7.org/fhir/encounter-status"}}}, "subject_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "subject_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "submitter_id": {"type": ["string", "null"]}, "type": {"type": "string"}, "type_coding_sct": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "type_text": {"description": "Plain text representation of the concept.", "type": ["string", "null"]}, "updated_datetime": {"$ref": "_definitions.yaml#/datetime"}}, "required": ["submitter_id", "type", "status", "class_v3_ActCode"], "submittable": true, "systemProperties": ["id", "project_id", "created_datetime", "updated_datetime", "state"], "title": "Encounter", "type": "object", "uniqueKeys": [["id"], ["project_id", "submitter_id"]], "validators": null}, "Procedure.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "additionalProperties": false, "category": "Clinical", "description": " An action that is being or was performed on a patient.\n \n An action that is or was performed on or for a patient. This can be a\n physical intervention like an operation, or less invasive like long term\n services, counseling, or hypnotherapy.\n ", "id": "Procedure", "links": [{"backref": "Procedures", "label": "Patients", "multiplicity": "many_to_many", "name": "Patients", "required": true, "target_type": "Patient"}, {"backref": "Procedures", "label": "Encounters", "multiplicity": "many_to_many", "name": "Encounters", "required": false, "target_type": "Encounter"}], "namespace": "http://hl7.org/fhir", "program": "*", "project": "*", "properties": {"code_coding_sct": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "code_text": {"description": "Plain text representation of the concept.", "type": ["string", "null"]}, "created_datetime": {"$ref": "_definitions.yaml#/datetime"}, "encounter_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "id": {"description": "", "type": ["string", "null"]}, "location_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "location_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "meta_profile": {"description": "Profiles this resource claims to conform to.", "type": ["string", "null"]}, "performedPeriod_end": {"description": "End time with inclusive boundary, if not ongoing.", "type": ["string", "null"]}, "performedPeriod_start": {"description": "Starting time with inclusive boundary.", "type": ["string", "null"]}, "project_id": {"$ref": "_definitions.yaml#/project_id"}, "reasonReference_0_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "reasonReference_0_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "reasonReference_1_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "reasonReference_1_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "reasonReference_2_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "reasonReference_2_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "reasonReference_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "reasonReference_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "resourceType": {"description": "One of the resource types defined as part of this version of FHIR", "type": ["string", "null"]}, "state": {"$ref": "_definitions.yaml#/state"}, "status": {"description": "A code specifying the state of the procedure. Generally, this will be the in-progress or completed state.. http://hl7.org/fhir/event-status", "enum": ["preparation", "in-progress", "not-done", "on-hold", "stopped", "completed", "entered-in-error", "unknown"], "term": {"description": "A code specifying the state of the procedure. Generally, this will be the in-progress or completed state.. http://hl7.org/fhir/event-status", "termDef": {"cde_id": "http://hl7.org/fhir/event-status", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/event-status", "term_url": "http://hl7.org/fhir/event-status"}}}, "subject_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "submitter_id": {"type": ["string", "null"]}, "type": {"type": "string"}, "updated_datetime": {"$ref": "_definitions.yaml#/datetime"}}, "required": ["submitter_id", "type", "status"], "submittable": true, "systemProperties": ["id", "project_id", "created_datetime", "updated_datetime", "state"], "title": "Procedure", "type": "object", "uniqueKeys": [["id"], ["project_id", "submitter_id"]], "validators": null}, "Organization.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "additionalProperties": false, "category": "Administrative", "description": " A grouping of people or organizations with a common purpose.\n \n A formally or informally recognized grouping of people or organizations\n formed for the purpose of achieving some form of collective action.\n Includes companies, institutions, corporations, departments, community\n groups, healthcare practice groups, payer/insurer, etc.\n ", "id": "Organization", "links": [{"backref": "Organizations", "label": "Organizations", "multiplicity": "many_to_many", "name": "Organizations", "required": false, "target_type": "Organization"}], "namespace": "http://hl7.org/fhir", "program": "*", "project": "*", "properties": {"created_datetime": {"$ref": "_definitions.yaml#/datetime"}, "id": {"description": "", "type": ["string", "null"]}, "identifier_name": {"description": "Identifies this organization across multiple systems.", "type": ["string", "null"]}, "identifier_synthea": {"description": "Identifies this organization across multiple systems.", "type": ["string", "null"]}, "name": {"description": "Name used for the organization.", "type": ["string", "null"]}, "project_id": {"$ref": "_definitions.yaml#/project_id"}, "resourceType": {"description": "One of the resource types defined as part of this version of FHIR", "type": ["string", "null"]}, "state": {"$ref": "_definitions.yaml#/state"}, "submitter_id": {"type": ["string", "null"]}, "telecom_system": {"description": "Telecommunications form for contact point - what communications system is required to make use of the contact.. http://hl7.org/fhir/contact-point-system", "enum": ["phone", "fax", "email", "pager", "url", "sms", "other"], "term": {"description": "Telecommunications form for contact point - what communications system is required to make use of the contact.. http://hl7.org/fhir/contact-point-system", "termDef": {"cde_id": "http://hl7.org/fhir/contact-point-system", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/contact-point-system", "term_url": "http://hl7.org/fhir/contact-point-system"}}}, "telecom_value": {"description": "The actual contact point details.", "type": ["string", "null"]}, "type": {"type": "string"}, "updated_datetime": {"$ref": "_definitions.yaml#/datetime"}}, "required": ["submitter_id", "type"], "submittable": true, "systemProperties": ["id", "project_id", "created_datetime", "updated_datetime", "state"], "title": "Organization", "type": "object", "uniqueKeys": [["id"], ["project_id", "submitter_id"]], "validators": null}, "_terms.yaml": {"id": "_terms", "data_category": {"description": "Broad categorization of the contents of the data file.\n"}, "data_file_error_type": {"description": "Type of error for the data file object.\n"}, "data_format": {"description": "Format of the data files.\n"}, "data_type": {"description": "Specific content type of the data file. (CMG)\n"}, "datetime": {"description": "A combination of date and time of day in the form [-]CCYY-MM-DDThh:mm:ss[Z|(+|-)hh:mm]\n"}, "encoding": {"description": "Version of ASCII encoding of quality values found in the file.\n", "termDef": {"term": "Encoding", "source": "FastQC", "cde_id": null, "cde_version": null, "term_url": "http://www.bioinformatics.babraham.ac.uk/projects/fastqc/Help/3%20Analysis%20Modules/1%20Basic%20Statistics.html"}}, "file_format": {"description": "The format of the data file object.\n"}, "file_name": {"description": "The name (or part of a name) of a file (of any type).\n"}, "file_size": {"description": "The size of the data file (object) in bytes.\n"}, "file_state": {"description": "The current state of the data file object.\n"}, "md5sum": {"description": "The 128-bit hash value expressed as a 32 digit hexadecimal number used as a file's digital fingerprint.\n"}, "project_id": {"description": "Unique ID for any specific defined piece of work that is undertaken or attempted to meet a single requirement.\n"}, "state": {"description": "The current state of the object.\n"}, "UUID": {"description": "A 128-bit identifier. Depending on the mechanism used to generate it, it is either guaranteed to be different from all other UUIDs/GUIDs generated until 3400 AD or extremely likely to be different. Its relatively small size lends itself well to sorting, ordering, and hashing of all sorts, storing in databases, simple allocation, and ease of programming in general.\n", "termDef": {"term": "Universally Unique Identifier", "source": "NCIt", "cde_id": "C54100", "cde_version": null, "term_url": "https://ncit.nci.nih.gov/ncitbrowser/ConceptReport.jsp?dictionary=NCI_Thesaurus&version=16.02d&ns=NCI_Thesaurus&code=C54100"}}, "microsatellite_instability_abnormal": {"description": "The yes/no indicator to signify the status of a tumor for microsatellite instability.\n", "termDef": {"term": "Microsatellite Instability Occurrence Indicator", "source": "caDSR", "cde_id": 3123142, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3123142&version=1.0"}}, "morphology": {"description": "The third edition of the International Classification of Diseases for Oncology, published in 2000 used principally in tumor and cancer registries for coding the site (topography) and the histology (morphology) of neoplasms. The study of the structure of the cells and their arrangement to constitute tissues and, finally, the association among these to form organs. In pathology, the microscopic process of identifying normal and abnormal morphologic characteristics in tissues, by employing various cytochemical and immunocytochemical stains. A system of numbered categories for representation of data.\n", "termDef": {"term": "International Classification of Diseases for Oncology, Third Edition ICD-O-3 Histology Code", "source": "caDSR", "cde_id": 3226275, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3226275&version=1.0"}}, "new_event_anatomic_site": {"description": "Text term to specify the anatomic location of the return of tumor after treatment.\n", "termDef": {"term": "New Neoplasm Event Occurrence Anatomic Site", "source": "caDSR", "cde_id": 3108271, "cde_version": 2.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3108271&version=2.0"}}, "new_event_type": {"description": "Text term to identify a new tumor event.\n", "termDef": {"term": "New Neoplasm Event Type", "source": "caDSR", "cde_id": 3119721, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3119721&version=1.0"}}, "normal_tumor_genotype_snp_match": {"description": "Text term that represents whether or not the genotype of the normal tumor matches or if the data is not available.\n", "termDef": {"term": "Normal Tumor Genotype Match Indicator", "source": "caDSR", "cde_id": 4588156, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=4588156&version=1.0"}}, "number_proliferating_cells": {"description": "Numeric value that represents the count of proliferating cells determined during pathologic review of the sample slide(s).\n", "termDef": {"term": "Pathology Review Slide Proliferating Cell Count", "source": "caDSR", "cde_id": 5432636, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432636&version=1.0"}}, "oct_embedded": {"description": "Indicator of whether or not the sample was embedded in Optimal Cutting Temperature (OCT) compound.\n", "termDef": {"term": "Tissue Sample Optimal Cutting Temperature Compound Embedding Indicator", "source": "caDSR", "cde_id": 5432538, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432538&version=1.0"}}, "years_smoked": {"description": "Numeric value (or unknown) to represent the number of years a person has been smoking (HARMONIZED). If the number of years is greater than 89, see 'years_smoked_gt89'.\n", "termDef": {"term": "Person Smoking Duration Year Count", "source": "caDSR", "cde_id": 3137957, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3137957&version=1.0"}}, "percent_eosinophil_infiltration": {"description": "Numeric value to represent the percentage of infiltration by eosinophils in a tumor sample or specimen.\n", "termDef": {"term": "Specimen Eosinophilia Percentage Value", "source": "caDSR", "cde_id": 2897700, "cde_version": 2.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2897700&version=2.0"}}, "percent_gc_content": {"description": "The overall %GC of all bases in all sequences.\n", "termDef": {"term": "%GC", "source": "FastQC", "cde_id": null, "cde_version": null, "term_url": "http://www.bioinformatics.babraham.ac.uk/projects/fastqc/Help/3%20Analysis%20Modules/1%20Basic%20Statistics.html"}}, "percent_granulocyte_infiltration": {"description": "Numeric value to represent the percentage of infiltration by granulocytes in a tumor sample or specimen.\n", "termDef": {"term": "Specimen Granulocyte Infiltration Percentage Value", "source": "caDSR", "cde_id": 2897705, "cde_version": 2.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2897705&version=2.0"}}, "percent_inflam_infiltration": {"description": "Numeric value to represent local response to cellular injury, marked by capillary dilatation, edema and leukocyte infiltration; clinically, inflammation is manifest by reddness, heat, pain, swelling and loss of function, with the need to heal damaged tissue.\n", "termDef": {"term": "Specimen Inflammation Change Percentage Value", "source": "caDSR", "cde_id": 2897695, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2897695&version=1.0"}}, "percent_lymphocyte_infiltration": {"description": "Numeric value to represent the percentage of infiltration by lymphocytes in a solid tissue normal sample or specimen.\n", "termDef": {"term": "Specimen Lymphocyte Infiltration Percentage Value", "source": "caDSR", "cde_id": 2897710, "cde_version": 2.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2897710&version=2.0"}}, "percent_monocyte_infiltration": {"description": "Numeric value to represent the percentage of monocyte infiltration in a sample or specimen.\n", "termDef": {"term": "Specimen Monocyte Infiltration Percentage Value", "source": "caDSR", "cde_id": 5455535, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5455535&version=1.0"}}, "percent_necrosis": {"description": "Numeric value to represent the percentage of cell death in a malignant tumor sample or specimen.\n", "termDef": {"term": "Malignant Neoplasm Necrosis Percentage Value", "source": "caDSR", "cde_id": 2841237, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2841237&version=1.0"}}, "percent_neutrophil_infiltration": {"description": "Numeric value to represent the percentage of infiltration by neutrophils in a tumor sample or specimen.\n", "termDef": {"term": "Malignant Neoplasm Neutrophil Infiltration Percentage Cell Value", "source": "caDSR", "cde_id": 2841267, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2841267&version=1.0"}}, "percent_normal_cells": {"description": "Numeric value to represent the percentage of normal cell content in a malignant tumor sample or specimen.\n", "termDef": {"term": "Malignant Neoplasm Normal Cell Percentage Value", "source": "caDSR", "cde_id": 2841233, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2841233&version=1.0"}}, "percent_stromal_cells": {"description": "Numeric value to represent the percentage of reactive cells that are present in a malignant tumor sample or specimen but are not malignant such as fibroblasts, vascular structures, etc.\n", "termDef": {"term": "Malignant Neoplasm Stromal Cell Percentage Value", "source": "caDSR", "cde_id": 2841241, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2841241&version=1.0"}}, "percent_tumor_cells": {"description": "Numeric value that represents the percentage of infiltration by granulocytes in a sample.\n", "termDef": {"term": "Specimen Tumor Cell Percentage Value", "source": "caDSR", "cde_id": 5432686, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432686&version=1.0"}}, "percent_tumor_nuclei": {"description": "Numeric value to represent the percentage of tumor nuclei in a malignant neoplasm sample or specimen.\n", "termDef": {"term": "Malignant Neoplasm Neoplasm Nucleus Percentage Cell Value", "source": "caDSR", "cde_id": 2841225, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2841225&version=1.0"}}, "perineural_invasion_present": {"description": "a yes/no indicator to ask if perineural invasion or infiltration of tumor or cancer is present.\n", "termDef": {"term": "Tumor Perineural Invasion Ind", "source": "caDSR", "cde_id": 64181, "cde_version": 3.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=64181&version=3.0"}}, "platform": {"description": "Name of the platform used to obtain data.\n"}, "portion_number": {"description": "Numeric value that represents the sequential number assigned to a portion of the sample.\n", "termDef": {"term": "Biospecimen Portion Sequence Number", "source": "caDSR", "cde_id": 5432711, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432711&version=1.0"}}, "portion_weight": {"description": "Numeric value that represents the sample portion weight, measured in milligrams.\n", "termDef": {"term": "Biospecimen Portion Weight Milligram Value", "source": "caDSR", "cde_id": 5432593, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432593&version=1.0"}}, "preservation_method": {"description": "Text term that represents the method used to preserve the sample.\n", "termDef": {"term": "Tissue Sample Preservation Method Type", "source": "caDSR", "cde_id": 5432521, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432521&version=1.0"}}, "prior_malignancy": {"description": "Text term to describe the patient's history of prior cancer diagnosis and the spatial location of any previous cancer occurrence.\n", "termDef": {"term": "Prior Cancer Diagnosis Occurrence Description Text", "source": "caDSR", "cde_id": 3382736, "cde_version": 2.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3382736&version=2.0"}}, "prior_treatment": {"description": "A yes/no/unknown/not applicable indicator related to the administration of therapeutic agents received before the body specimen was collected.\n", "termDef": {"term": "Therapeutic Procedure Prior Specimen Collection Administered Yes No Unknown Not Applicable Indicator", "source": "caDSR", "cde_id": 4231463, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=4231463&version=1.0"}}, "progesterone_receptor_percent_positive_ihc": {"description": "Classification to represent Progesterone Receptor Positive results expressed as a percentage value.\n", "termDef": {"term": "Progesterone Receptor Level Cell Percentage Category", "source": "caDSR", "cde_id": 3128342, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3128342&version=1.0"}}, "progesterone_receptor_result_ihc": {"description": "Text term to represent the overall result of Progresterone Receptor (PR) testing.\n", "termDef": {"term": "Breast Carcinoma Progesterone Receptor Status", "source": "caDSR", "cde_id": 2957357, "cde_version": 2.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2957357&version=2.0"}}, "progression_or_recurrence": {"description": "Yes/No/Unknown indicator to identify whether a patient has had a new tumor event after initial treatment.\n", "termDef": {"term": "New Neoplasm Event Post Initial Therapy Indicator", "source": "caDSR", "cde_id": 3121376, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3121376&version=1.0"}}, "qc_metric_state": {"description": "State classification given by FASTQC for the metric. Metric specific details about the states are available on their website.\n", "termDef": {"term": "QC Metric State", "source": "FastQC", "cde_id": null, "cde_version": null, "term_url": "http://www.bioinformatics.babraham.ac.uk/projects/fastqc/Help/3%20Analysis%20Modules/"}}, "race": {"description": "An arbitrary classification of a taxonomic group that is a division of a species (HARMONIZED). It usually arises as a consequence of geographical isolation within a species and is characterized by shared heredity, physical attributes and behavior, and in the case of humans, by common history, nationality, or geographic distribution. The provided values are based on the categories defined by the U.S. Office of Management and Business and used by the U.S. Census Bureau. (GTEx)\n", "termDef": {"term": "Race Category Text", "source": "caDSR", "cde_id": 2192199, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2192199&version=1.0"}}, "read_length": {"description": "The length of the reads.\n"}, "read_group_name": {"description": "The name of the read group.\n"}, "relationship_age_at_diagnosis": {"description": "The age (in years) when the patient's relative was first diagnosed.\n", "termDef": {"term": "Relative Diagnosis Age Value", "source": "caDSR", "cde_id": 5300571, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5300571&version=1.0"}}, "relationship_to_proband": {"description": "The subgroup that describes the state of connectedness between members of the unit of society organized around kinship ties.\n", "termDef": {"term": "Family Member Relationship Type", "source": "caDSR", "cde_id": 2690165, "cde_version": 2.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2690165&version=2.0"}}, "relationship_type": {"description": "The subgroup that describes the state of connectedness between members of the unit of society organized around kinship ties.\n", "termDef": {"term": "Family Member Relationship Type", "source": "caDSR", "cde_id": 2690165, "cde_version": 2.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2690165&version=2.0"}}, "relative_with_cancer_history": {"description": "Indicator to signify whether or not an individual's biological relative has been diagnosed with another type of cancer.\n", "termDef": {"term": "Other Cancer Biological Relative History Indicator", "source": "caDSR", "cde_id": 3901752, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3901752&version=1.0"}}, "residual_disease": {"description": "Text terms to describe the status of a tissue margin following surgical resection.\n", "termDef": {"term": "Surgical Margin Resection Status", "source": "caDSR", "cde_id": 2608702, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2608702&version=1.0"}}, "RIN": {"description": "A numerical assessment of the integrity of RNA based on the entire electrophoretic trace of the RNA sample including the presence or absence of degradation products.\n", "termDef": {"term": "Biospecimen RNA Integrity Number Value", "source": "caDSR", "cde_id": 5278775, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5278775&version=1.0"}}, "sample_type": {"description": "Text term to describe the source of a biospecimen used for a laboratory test.\n", "termDef": {"term": "Specimen Type Collection Biospecimen Type", "source": "caDSR", "cde_id": 3111302, "cde_version": 2.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3111302&version=2.0"}}, "sample_type_id": {"description": "The accompanying sample type id for the sample type.\n"}, "section_location": {"description": "Tissue source of the slide.\n"}, "sequencing_center": {"description": "Name of the center that provided the sequence files.\n"}, "shortest_dimension": {"description": "Numeric value that represents the shortest dimension of the sample, measured in millimeters.\n", "termDef": {"term": "Tissue Sample Short Dimension Millimeter Measurement", "source": "caDSR", "cde_id": 5432603, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432603&version=1.0"}}, "site_of_resection_or_biopsy": {"description": "The third edition of the International Classification of Diseases for Oncology, published in 2000, used principally in tumor and cancer registries for coding the site (topography) and the histology (morphology) of neoplasms. The description of an anatomical region or of a body part. Named locations of, or within, the body. A system of numbered categories for representation of data.\n", "termDef": {"term": "International Classification of Diseases for Oncology, Third Edition ICD-O-3 Site Code", "source": "caDSR", "cde_id": 3226281, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3226281&version=1.0"}}, "size_selection_range": {"description": "Range of size selection.\n"}, "smoking_history": {"description": "Category describing current smoking status and smoking history as self-reported by a patient.\n", "termDef": {"term": "Smoking History", "source": "caDSR", "cde_id": 2181650, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2181650&version=1.0"}}, "smoking_intensity": {"description": "Numeric computed value to represent lifetime tobacco exposure defined as number of cigarettes smoked per day x number of years smoked divided by 20\n", "termDef": {"term": "Person Cigarette Smoking History Pack Year Value", "source": "caDSR", "cde_id": 2955385, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2955385&version=1.0"}}, "source_center": {"description": "Name of the center that provided the item.\n"}, "spectrophotometer_method": {"description": "Name of the method used to determine the concentration of purified nucleic acid within a solution.\n", "termDef": {"term": "Purification Nucleic Acid Solution Concentration Determination Method Type", "source": "caDSR", "cde_id": 3008378, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3008378&version=1.0"}}, "spike_ins_fasta": {"description": "Name of the FASTA file that contains the spike-in sequences.\n"}, "spike_ins_concentration": {"description": "Spike in concentration.\n"}, "target_capture_kit_name": {"description": "Name of Target Capture Kit.\n"}, "target_capture_kit_vendor": {"description": "Vendor of Target Capture Kit.\n"}, "target_capture_kit_catalog_number": {"description": "Catalog of Target Capture Kit.\n"}, "target_capture_kit_version": {"description": "Version of Target Capture Kit.\n"}, "target_capture_kit_target_region": {"description": "Target Capture Kit BED file.\n"}, "therapeutic_agents": {"description": "Text identification of the individual agent(s) used as part of a prior treatment regimen.\n", "termDef": {"term": "Prior Therapy Regimen Text", "source": "caDSR", "cde_id": 2975232, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2975232&version=1.0"}}, "time_between_clamping_and_freezing": {"description": "Numeric representation of the elapsed time between the surgical clamping of blood supply and freezing of the sample, measured in minutes.\n", "termDef": {"term": "Tissue Sample Clamping and Freezing Elapsed Minute Time", "source": "caDSR", "cde_id": 5432611, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432611&version=1.0"}}, "time_between_excision_and_freezing": {"description": "Numeric representation of the elapsed time between the excision and freezing of the sample, measured in minutes.\n", "termDef": {"term": "Tissue Sample Excision and Freezing Elapsed Minute Time", "source": "caDSR", "cde_id": 5432612, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432612&version=1.0"}}, "tissue_or_organ_of_origin": {"description": "Text term that describes the anatomic site of the tumor or disease.\n", "termDef": {"term": "Tumor Disease Anatomic Site", "source": "caDSR", "cde_id": 3427536, "cde_version": 3.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3427536&version=3.0"}}, "tissue_type": {"description": "Text term that represents a description of the kind of tissue collected with respect to disease status or proximity to tumor tissue.\n", "termDef": {"term": "Tissue Sample Description Type", "source": "caDSR", "cde_id": 5432687, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432687&version=1.0"}}, "to_trim_adapter_sequence": {"description": "Does the user suggest adapter trimming?\n"}, "tobacco_smoking_onset_year": {"description": "The year in which the participant began smoking.\n", "termDef": {"term": "Started Smoking Year", "source": "caDSR", "cde_id": 2228604, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2228604&version=1.0"}}, "tobacco_smoking_quit_year": {"description": "The year in which the participant quit smoking.\n", "termDef": {"term": "Stopped Smoking Year", "source": "caDSR", "cde_id": 2228610, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2228610&version=1.0"}}, "tobacco_smoking_status": {"description": "Category describing current smoking status and smoking history as self-reported by a patient.\n", "termDef": {"term": "Patient Smoking History Category", "source": "caDSR", "cde_id": 2181650, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2181650&version=1.0"}}, "total_sequences": {"description": "A count of the total number of sequences processed.\n", "termDef": {"term": "Total Sequences", "source": "FastQC", "cde_id": null, "cde_version": null, "term_url": "http://www.bioinformatics.babraham.ac.uk/projects/fastqc/Help/3%20Analysis%20Modules/1%20Basic%20Statistics.html"}}, "treatment_anatomic_site": {"description": "The anatomic site or field targeted by a treatment regimen or single agent therapy.\n", "termDef": {"term": "Treatment Anatomic Site", "source": null, "cde_id": null, "cde_version": null, "term_url": null}}, "treatment_intent_type": {"description": "Text term to identify the reason for the administration of a treatment regimen. [Manually-curated]\n", "termDef": {"term": "Treatment Regimen Intent Type", "source": "caDSR", "cde_id": 2793511, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2793511&version=1.0"}}, "treatment_or_therapy": {"description": "A yes/no/unknown/not applicable indicator related to the administration of therapeutic agents received before the body specimen was collected.\n", "termDef": {"term": "Therapeutic Procedure Prior Specimen Collection Administered Yes No Unknown Not Applicable Indicator", "source": "caDSR", "cde_id": 4231463, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=4231463&version=1.0"}}, "treatment_outcome": {"description": "Text term that describes the patient\u00bfs final outcome after the treatment was administered.\n", "termDef": {"term": "Treatment Outcome Type", "source": "caDSR", "cde_id": 5102383, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5102383&version=1.0"}}, "treatment_type": {"description": "Text term that describes the kind of treatment administered.\n", "termDef": {"term": "Treatment Method Type", "source": "caDSR", "cde_id": 5102381, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5102381&version=1.0"}}, "tumor_grade": {"description": "Numeric value to express the degree of abnormality of cancer cells, a measure of differentiation and aggressiveness.\n", "termDef": {"term": "Neoplasm Histologic Grade", "source": "caDSR", "cde_id": 2785839, "cde_version": 2.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2785839&version=2.0"}}, "tumor_code": {"description": "Diagnostic tumor code of the tissue sample source.\n"}, "tumor_code_id": {"description": "BCR-defined id code for the tumor sample.\n"}, "tumor_descriptor": {"description": "Text that describes the kind of disease present in the tumor specimen as related to a specific timepoint.\n", "termDef": {"term": "Tumor Tissue Disease Description Type", "source": "caDSR", "cde_id": 3288124, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3288124&version=1.0"}}, "tumor_stage": {"description": "The extent of a cancer in the body. Staging is usually based on the size of the tumor, whether lymph nodes contain cancer, and whether the cancer has spread from the original site to other parts of the body. The accepted values for tumor_stage depend on the tumor site, type, and accepted staging system. These items should accompany the tumor_stage value as associated metadata.\n", "termDef": {"term": "Tumor Stage", "source": "NCIt", "cde_id": "C16899", "cde_version": null, "term_url": "https://ncit.nci.nih.gov/ncitbrowser/pages/concept_details.jsf?dictionary=NCI%20Thesaurus&code=C16899"}}, "vascular_invasion_present": {"description": "The yes/no indicator to ask if large vessel or venous invasion was detected by surgery or presence in a tumor specimen.\n", "termDef": {"term": "Tumor Vascular Invasion Ind-3", "source": "caDSR", "cde_id": 64358, "cde_version": 3.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=64358&version=3.0"}}, "vital_status": {"description": "The survival state of the person registered on the protocol.\n", "termDef": {"term": "Patient Vital Status", "source": "caDSR", "cde_id": 5, "cde_version": 5.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5&version=5.0"}}, "weight": {"description": "The weight of the patient measured in lbs (HARMONIZED).\n", "termDef": {"term": "Patient Weight Measurement", "source": "caDSR", "cde_id": 651, "cde_version": 4.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=651&version=4.0"}}, "well_number": {"description": "Numeric value that represents the the well location within a plate for the analyte or aliquot from the sample.\n", "termDef": {"term": "Biospecimen Analyte or Aliquot Plate Well Number", "source": "caDSR", "cde_id": 5432613, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432613&version=1.0"}}, "workflow_type": {"description": "Generic name for the workflow used to analyze a data set.\n"}, "year_of_birth": {"description": "Numeric value to represent the calendar year in which an individual was born.\n", "termDef": {"term": "Year Birth Date Number", "source": "caDSR", "cde_id": 2896954, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2896954&version=1.0"}}, "year_of_diagnosis": {"description": "Numeric value to represent the year of an individual's initial pathologic diagnosis of cancer.\n", "termDef": {"term": "Year of initial pathologic diagnosis", "source": "caDSR", "cde_id": 2896960, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2896960&version=1.0"}}, "year_of_death": {"description": "Numeric value to represent the year of the death of an individual.\n", "termDef": {"term": "Year Death Number", "source": "caDSR", "cde_id": 2897030, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2897030&version=1.0"}}, "years_smoked_gt89": {"description": "Indicate whether the numeric value to represent the number of years a person has been smoking (HARMONIZED) is greater than 89 years.\n", "termDef": {"term": "Person Smoking Duration Year Count", "source": "caDSR", "cde_id": 3137957, "cde_version": 1.0, "term_url": "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3137957&version=1.0"}}, "ga4gh_drs_uri": {"description": "DRS URI as defined by GA4GH DRS spec for pointers to file objects.\n"}}, "ImagingStudy.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "additionalProperties": false, "category": "Clinical", "description": " A set of images produced in single study (one or more series of references\n images).\n \n Representation of the content produced in a DICOM imaging study. A study\n comprises a set of series, each of which includes a set of Service-Object\n Pair Instances (SOP Instances - images or other data) acquired or produced\n in a common context. A series is of only one modality (e.g. X-ray, CT, MR,\n ultrasound), but a study may have multiple series of different modalities.\n ", "id": "ImagingStudy", "links": [{"backref": "ImagingStudies", "label": "Patients", "multiplicity": "many_to_many", "name": "Patients", "required": true, "target_type": "Patient"}, {"backref": "ImagingStudies", "label": "Encounters", "multiplicity": "many_to_many", "name": "Encounters", "required": false, "target_type": "Encounter"}], "namespace": "http://hl7.org/fhir", "program": "*", "project": "*", "properties": {"created_datetime": {"$ref": "_definitions.yaml#/datetime"}, "encounter_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "id": {"description": "", "type": ["string", "null"]}, "identifier_urn_ietf_rfc_3986": {"description": "Identifiers for the whole study.", "type": ["string", "null"]}, "location_display": {"description": "Text alternative for the resource.", "type": ["string", "null"]}, "location_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "numberOfInstances": {"description": "Number of Study Related Instances.", "type": ["number", "null"]}, "numberOfSeries": {"description": "Number of Study Related Series.", "type": ["number", "null"]}, "project_id": {"$ref": "_definitions.yaml#/project_id"}, "resourceType": {"description": "One of the resource types defined as part of this version of FHIR", "type": ["string", "null"]}, "series_bodySite_sct": {"description": "Body part examined.", "type": ["string", "null"]}, "series_instance_number": {"description": "The number of this instance in the series.", "type": ["number", "null"]}, "series_instance_sopClass_urn_ietf_rfc_3986": {"description": "DICOM class type.", "type": ["string"]}, "series_instance_title": {"description": "Description of instance.", "type": ["string", "null"]}, "series_instance_uid": {"description": "DICOM SOP Instance UID.", "type": ["string"]}, "series_modality_sect_CID_29_html": {"description": "The modality of the instances in the series.", "type": ["string"]}, "series_number": {"description": "Numeric identifier of this series.", "type": ["number", "null"]}, "series_numberOfInstances": {"description": "Number of Series Related Instances.", "type": ["number", "null"]}, "series_started": {"description": "When the series started.", "type": ["string", "null"]}, "series_uid": {"description": "DICOM Series Instance UID for the series.", "type": ["string"]}, "started": {"description": "When the study was started.", "type": ["string", "null"]}, "state": {"$ref": "_definitions.yaml#/state"}, "status": {"description": "The current state of the ImagingStudy.. http://hl7.org/fhir/imagingstudy-status", "enum": ["registered", "available", "cancelled", "entered-in-error", "unknown"], "term": {"description": "The current state of the ImagingStudy.. http://hl7.org/fhir/imagingstudy-status", "termDef": {"cde_id": "http://hl7.org/fhir/imagingstudy-status", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/imagingstudy-status", "term_url": "http://hl7.org/fhir/imagingstudy-status"}}}, "subject_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "submitter_id": {"type": ["string", "null"]}, "type": {"type": "string"}, "updated_datetime": {"$ref": "_definitions.yaml#/datetime"}}, "required": ["submitter_id", "type", "status", "series_uid", "series_modality_sect_CID_29_html", "series_instance_uid", "series_instance_sopClass_urn_ietf_rfc_3986"], "submittable": true, "systemProperties": ["id", "project_id", "created_datetime", "updated_datetime", "state"], "title": "ImagingStudy", "type": "object", "uniqueKeys": [["id"], ["project_id", "submitter_id"]], "validators": null}, "ResearchStudy.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "additionalProperties": false, "category": "Administrative", "description": " Investigation to increase healthcare-related patient-independent knowledge.\n \n A process where a researcher or organization plans and then executes a\n series of steps intended to increase the field of healthcare-related\n knowledge. This includes studies of safety, efficacy, comparative\n effectiveness and other information about medications, devices, therapies\n and other interventional and investigative techniques. A ResearchStudy\n involves the gathering of information about human or animal subjects.\n ", "id": "ResearchStudy", "links": [{"backref": "ResearchStudies", "label": "ResearchStudies", "multiplicity": "many_to_many", "name": "ResearchStudies", "required": false, "target_type": "ResearchStudy"}, {"backref": "ResearchStudies", "label": "Practitioners", "multiplicity": "many_to_many", "name": "Practitioners", "required": false, "target_type": "Practitioner"}, {"backref": "ResearchStudies", "label": "Organizations", "multiplicity": "many_to_many", "name": "Organizations", "required": false, "target_type": "Organization"}, {"backref": "ResearchStudies", "label": "projects", "multiplicity": "many_to_many", "name": "projects", "required": true, "target_type": "project"}], "namespace": "http://hl7.org/fhir", "program": "*", "project": "*", "properties": {"condition_0_coding_sct": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "condition_0_text": {"description": "Plain text representation of the concept.", "type": ["string", "null"]}, "condition_1_coding_sct": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "condition_1_text": {"description": "Plain text representation of the concept.", "type": ["string", "null"]}, "created_datetime": {"$ref": "_definitions.yaml#/datetime"}, "description": {"description": "What this is study doing.", "type": ["string", "null"]}, "id": {"description": "", "type": ["string", "null"]}, "project_id": {"$ref": "_definitions.yaml#/project_id"}, "resourceType": {"description": "One of the resource types defined as part of this version of FHIR", "type": ["string", "null"]}, "state": {"$ref": "_definitions.yaml#/state"}, "status": {"description": "The current state of the study.. http://hl7.org/fhir/research-study-status", "enum": ["active", "administratively-completed", "approved", "closed-to-accrual", "closed-to-accrual-and-intervention", "completed", "disapproved", "in-review", "temporarily-closed-to-accrual", "temporarily-closed-to-accrual-and-intervention", "withdrawn"], "term": {"description": "The current state of the study.. http://hl7.org/fhir/research-study-status", "termDef": {"cde_id": "http://hl7.org/fhir/research-study-status", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/research-study-status", "term_url": "http://hl7.org/fhir/research-study-status"}}}, "submitter_id": {"type": ["string", "null"]}, "title": {"description": "Name for this study.", "type": ["string", "null"]}, "type": {"type": "string"}, "updated_datetime": {"$ref": "_definitions.yaml#/datetime"}}, "required": ["submitter_id", "type", "status"], "submittable": true, "systemProperties": ["id", "project_id", "created_datetime", "updated_datetime", "state"], "title": "ResearchStudy", "type": "object", "uniqueKeys": [["id"], ["project_id", "submitter_id"]], "validators": null}, "Specimen.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "additionalProperties": false, "category": "Biospecimen", "description": " Sample for analysis.\n \n A sample to be used for analysis.\n ", "id": "Specimen", "links": [{"backref": "Specimen", "label": "Patients", "multiplicity": "many_to_many", "name": "Patients", "required": true, "target_type": "Patient"}], "namespace": "http://hl7.org/fhir", "program": "*", "project": "*", "properties": {"created_datetime": {"$ref": "_definitions.yaml#/datetime"}, "id": {"description": "", "type": ["string", "null"]}, "project_id": {"$ref": "_definitions.yaml#/project_id"}, "resourceType": {"description": "One of the resource types defined as part of this version of FHIR", "type": ["string", "null"]}, "state": {"$ref": "_definitions.yaml#/state"}, "subject_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "submitter_id": {"type": ["string", "null"]}, "text_div": {"description": "Limited xhtml content.", "type": ["string"]}, "text_status": {"description": "The status of the narrative - whether it's entirely generated (from just the defined data or the extensions too), or whether a human authored it and it may contain additional data.. http://hl7.org/fhir/narrative-status", "enum": ["generated", "extensions", "additional", "empty"], "term": {"description": "The status of the narrative - whether it's entirely generated (from just the defined data or the extensions too), or whether a human authored it and it may contain additional data.. http://hl7.org/fhir/narrative-status", "termDef": {"cde_id": "http://hl7.org/fhir/narrative-status", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/narrative-status", "term_url": "http://hl7.org/fhir/narrative-status"}}}, "type": {"type": "string"}, "updated_datetime": {"$ref": "_definitions.yaml#/datetime"}}, "required": ["submitter_id", "type", "text_status", "text_div"], "submittable": true, "systemProperties": ["id", "project_id", "created_datetime", "updated_datetime", "state"], "title": "Specimen", "type": "object", "uniqueKeys": [["id"], ["project_id", "submitter_id"]], "validators": null}, "Medication.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "additionalProperties": false, "category": "Clinical", "description": " Definition of a Medication.\n \n This resource is primarily used for the identification and definition of a\n medication for the purposes of prescribing, dispensing, and administering a\n medication as well as for making statements about medication use.\n ", "id": "Medication", "links": [], "namespace": "http://hl7.org/fhir", "program": "*", "project": "*", "properties": {"code_coding_rxnorm": {"description": "Code defined by a terminology system.", "type": ["string", "null"]}, "code_text": {"description": "Plain text representation of the concept.", "type": ["string", "null"]}, "created_datetime": {"$ref": "_definitions.yaml#/datetime"}, "id": {"description": "", "type": ["string", "null"]}, "meta_profile": {"description": "Profiles this resource claims to conform to.", "type": ["string", "null"]}, "project_id": {"$ref": "_definitions.yaml#/project_id"}, "resourceType": {"description": "One of the resource types defined as part of this version of FHIR", "type": ["string", "null"]}, "state": {"$ref": "_definitions.yaml#/state"}, "status": {"description": "A code to indicate if the medication is in active use.. http://hl7.org/fhir/CodeSystem/medication-status", "enum": ["active", "inactive", "entered-in-error"], "term": {"description": "A code to indicate if the medication is in active use.. http://hl7.org/fhir/CodeSystem/medication-status", "termDef": {"cde_id": "http://hl7.org/fhir/CodeSystem/medication-status", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/CodeSystem/medication-status", "term_url": "http://hl7.org/fhir/CodeSystem/medication-status"}}}, "submitter_id": {"type": ["string", "null"]}, "type": {"type": "string"}, "updated_datetime": {"$ref": "_definitions.yaml#/datetime"}}, "required": ["submitter_id", "type"], "submittable": true, "systemProperties": ["id", "project_id", "created_datetime", "updated_datetime", "state"], "title": "Medication", "type": "object", "uniqueKeys": [["id"], ["project_id", "submitter_id"]], "validators": null}, "ResearchSubject.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "additionalProperties": false, "category": "Administrative", "description": " Physical entity which is the primary unit of interest in the study.\n \n A physical entity which is the primary unit of operational and/or\n administrative interest in a study.\n ", "id": "ResearchSubject", "links": [{"backref": "ResearchSubjects", "label": "Patients", "multiplicity": "many_to_many", "name": "Patients", "required": true, "target_type": "Patient"}, {"backref": "ResearchSubjects", "label": "ResearchStudies", "multiplicity": "many_to_many", "name": "ResearchStudies", "required": true, "target_type": "ResearchStudy"}], "namespace": "http://hl7.org/fhir", "program": "*", "project": "*", "properties": {"created_datetime": {"$ref": "_definitions.yaml#/datetime"}, "id": {"description": "", "type": ["string", "null"]}, "individual_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "meta_source": {"description": "Identifies where the resource comes from.", "type": ["string", "null"]}, "project_id": {"$ref": "_definitions.yaml#/project_id"}, "resourceType": {"description": "One of the resource types defined as part of this version of FHIR", "type": ["string", "null"]}, "state": {"$ref": "_definitions.yaml#/state"}, "status": {"description": "The current state of the subject.. http://hl7.org/fhir/research-subject-status", "enum": ["candidate", "eligible", "follow-up", "ineligible", "not-registered", "off-study", "on-study", "on-study-intervention", "on-study-observation", "pending-on-study", "potential-candidate", "screening", "withdrawn"], "term": {"description": "The current state of the subject.. http://hl7.org/fhir/research-subject-status", "termDef": {"cde_id": "http://hl7.org/fhir/research-subject-status", "cde_version": null, "source": "fhir", "strength": "required", "term": "http://hl7.org/fhir/research-subject-status", "term_url": "http://hl7.org/fhir/research-subject-status"}}}, "study_reference": {"description": "Literal reference, Relative, internal or absolute URL.", "type": ["string", "null"]}, "submitter_id": {"type": ["string", "null"]}, "type": {"type": "string"}, "updated_datetime": {"$ref": "_definitions.yaml#/datetime"}}, "required": ["submitter_id", "type", "status"], "submittable": true, "systemProperties": ["id", "project_id", "created_datetime", "updated_datetime", "state"], "title": "ResearchSubject", "type": "object", "uniqueKeys": [["id"], ["project_id", "submitter_id"]], "validators": null}, "core_metadata_collection.yaml": {"$schema": "http://json-schema.org/draft-04/schema#", "id": "core_metadata_collection", "title": "Core Metadata Collection", "type": "object", "nodeTerms": null, "namespace": "https://data.midrc.org", "category": "administrative", "program": "*", "project": "*", "description": "A collection of data files in a project.", "additionalProperties": false, "submittable": true, "validators": null, "systemProperties": ["id", "project_id", "state", "created_datetime", "updated_datetime"], "links": [{"name": "projects", "backref": "core_metadata_collections", "label": "data_from", "target_type": "project", "multiplicity": "many_to_one", "required": true}], "required": ["submitter_id", "type", "projects"], "uniqueKeys": [["id"], ["project_id", "submitter_id"]], "properties": {"$ref": "_definitions.yaml#/ubiquitous_properties", "contributor": {"description": "An entity responsible for making contributions to the resource. Examples of a Contributor include a person, an organization, or a service. Typically, the name of a Contributor should be used to indicate the entity.", "type": "string"}, "coverage": {"description": "The spatial or temporal topic of the resource, the spatial applicability of the resource, or the jurisdiction under which the resource is relevant. Spatial topic and spatial applicability may be a named place or a location specified by its geographic coordinates. Temporal topic may be a named period, date, or date range. A jurisdiction may be a named administrative entity or a geographic place to which the resource applies. Recommended best practice is to use a controlled vocabulary such as the Thesaurus of Geographic Names [TGN] (http-//www.getty.edu/research/tools/vocabulary/tgn/index.html). Where appropriate, named places or time periods can be used in preference to numeric identifiers such as sets of coordinates or date ranges.", "type": "string"}, "creator": {"description": "An entity primarily responsible for making the resource. Examples of a Creator include a person, an organization, or a service. Typically, the name of a Creator should be used to indicate the entity.", "type": "string"}, "data_type": {"description": "The nature or genre of the resource. Recommended best practice is to use a controlled vocabulary such as the DCMI Type Vocabulary [DCMITYPE]. To describe the file format, physical medium, or dimensions of the resource, use the Format element.", "type": "string"}, "date": {"description": "The date the collection of data was created.", "type": "string"}, "description": {"description": "An account of the resource. Description may include but is not limited to- an abstract, a table of contents, a graphical representation, or a free-text account of the resource.", "type": "string"}, "format": {"description": "The file format, physical medium, or dimensions of the resource. Examples of dimensions include size and duration. Recommended best practice is to use a controlled vocabulary such as the list of Internet Media Types [MIME] (http-//www.iana.org/assignments/media-types/).", "type": "string"}, "language": {"description": "A language of the resource. Recommended best practice is to use a controlled vocabulary such as RFC 4646 (http-//www.ietf.org/rfc/rfc4646.txt).", "type": "string"}, "publisher": {"description": "An entity responsible for making the resource available. Examples of a Publisher include a person, an organization, or a service. Typically, the name of a Publisher should be used to indicate the entity.", "type": "string"}, "relation": {"description": "A related resource. Recommended best practice is to identify the related resource by means of a string conforming to a formal identification system.", "type": "string"}, "rights": {"description": "Information about rights held in and over the resource. Typically, rights information includes a statement about various property rights associated with the resource, including intellectual property rights.", "type": "string"}, "source": {"description": "A related resource from which the described resource is derived. The described resource may be derived from the related resource in whole or in part. Recommended best practice is to identify the related resource by means of a string conforming to a formal identification system.", "type": "string"}, "subject": {"description": "The topic of the resource. Typically, the subject will be represented using keywords, key phrases, or classification codes. Recommended best practice is to use a controlled vocabulary.", "type": "string"}, "title": {"description": "A name given to the resource. Typically, a Title will be a name by which the resource is formally known.", "type": "string"}, "projects": {"description": "The project code for the record in the project node that this core_metadata_collection belongs to, i.e., a link to the parent node.", "$ref": "_definitions.yaml#/to_one_project"}}}}
\ No newline at end of file
diff --git a/datadictionary/gdcdictionary/schemas/README.md b/datadictionary/gdcdictionary/schemas/README.md
deleted file mode 100644
index aa820df4..00000000
--- a/datadictionary/gdcdictionary/schemas/README.md
+++ /dev/null
@@ -1,24 +0,0 @@
-Proposed additional keywords
-============================
-
-The schemas defined here follow jsonschema as closely as possbile,
-introducing new keywords as needed.
-
-systemAlias
------------
-
-For implementation. Allows properties to be stored as different
-keywords. The property listed in the properties section is what the
-user will refer to it, and the systemAlias value is what it will be
-stored in the database as.
-
-systemProperties
----------------
-
-The property keys listed under systemProperties are properties that
-the submitter is not allowed to update.
-
-parentType
----------------
-
-The type of object that the parent relationship points to.
diff --git a/datadictionary/gdcdictionary/schemas/_core_metadata_collection.yaml b/datadictionary/gdcdictionary/schemas/_core_metadata_collection.yaml
new file mode 100644
index 00000000..7d97b1b5
--- /dev/null
+++ b/datadictionary/gdcdictionary/schemas/_core_metadata_collection.yaml
@@ -0,0 +1,140 @@
+$schema: "http://json-schema.org/draft-04/schema#"
+
+#
+# included because of windmill portal.
+# portal wont build unless this entity exists
+#
+
+id: "core_metadata_collection"
+title: Core Metadata Collection
+type: object
+nodeTerms: null
+namespace: https://data.midrc.org
+category: administrative
+program: '*'
+project: '*'
+description: A collection of data files in a project. DEPRECATED - included for portal compatibility
+additionalProperties: false
+submittable: true
+validators: null
+
+systemProperties:
+ - id
+ - project_id
+ - state
+ - created_datetime
+ - updated_datetime
+
+links:
+ - name: projects
+ backref: core_metadata_collections
+ label: data_from
+ target_type: project
+ multiplicity: many_to_one
+ required: true
+
+required:
+ - submitter_id
+ - type
+ - projects
+
+uniqueKeys:
+ - [id]
+ - [project_id, submitter_id]
+
+properties:
+
+ $ref: "_definitions.yaml#/ubiquitous_properties"
+
+ contributor:
+ description: An entity responsible for making contributions to the resource. Examples
+ of a Contributor include a person, an organization, or a service. Typically,
+ the name of a Contributor should be used to indicate the entity.
+ type: string
+
+ coverage:
+ description: The spatial or temporal topic of the resource, the spatial applicability
+ of the resource, or the jurisdiction under which the resource is relevant. Spatial
+ topic and spatial applicability may be a named place or a location specified
+ by its geographic coordinates. Temporal topic may be a named period, date, or
+ date range. A jurisdiction may be a named administrative entity or a geographic
+ place to which the resource applies. Recommended best practice is to use a controlled
+ vocabulary such as the Thesaurus of Geographic Names [TGN] (http-//www.getty.edu/research/tools/vocabulary/tgn/index.html).
+ Where appropriate, named places or time periods can be used in preference to
+ numeric identifiers such as sets of coordinates or date ranges.
+ type: string
+
+ creator:
+ description: An entity primarily responsible for making the resource. Examples
+ of a Creator include a person, an organization, or a service. Typically, the
+ name of a Creator should be used to indicate the entity.
+ type: string
+
+ data_type:
+ description: The nature or genre of the resource. Recommended best practice is
+ to use a controlled vocabulary such as the DCMI Type Vocabulary [DCMITYPE].
+ To describe the file format, physical medium, or dimensions of the resource,
+ use the Format element.
+ type: string
+
+ date:
+ description: The date the collection of data was created.
+ type: string
+
+ description:
+ description: An account of the resource. Description may include but is not limited
+ to- an abstract, a table of contents, a graphical representation, or a free-text
+ account of the resource.
+ type: string
+
+ format:
+ description: The file format, physical medium, or dimensions of the resource.
+ Examples of dimensions include size and duration. Recommended best practice
+ is to use a controlled vocabulary such as the list of Internet Media Types [MIME]
+ (http-//www.iana.org/assignments/media-types/).
+ type: string
+
+ language:
+ description: A language of the resource. Recommended best practice is to use a
+ controlled vocabulary such as RFC 4646 (http-//www.ietf.org/rfc/rfc4646.txt).
+ type: string
+
+ publisher:
+ description: An entity responsible for making the resource available. Examples
+ of a Publisher include a person, an organization, or a service. Typically, the
+ name of a Publisher should be used to indicate the entity.
+ type: string
+
+ relation:
+ description: A related resource. Recommended best practice is to identify the
+ related resource by means of a string conforming to a formal identification
+ system.
+ type: string
+
+ rights:
+ description: Information about rights held in and over the resource. Typically,
+ rights information includes a statement about various property rights associated
+ with the resource, including intellectual property rights.
+ type: string
+
+ source:
+ description: A related resource from which the described resource is derived.
+ The described resource may be derived from the related resource in whole or
+ in part. Recommended best practice is to identify the related resource by means
+ of a string conforming to a formal identification system.
+ type: string
+
+ subject:
+ description: The topic of the resource. Typically, the subject will be represented
+ using keywords, key phrases, or classification codes. Recommended best practice
+ is to use a controlled vocabulary.
+ type: string
+
+ title:
+ description: A name given to the resource. Typically, a Title will be a name by
+ which the resource is formally known.
+ type: string
+
+ projects:
+ description: The project code for the record in the project node that this core_metadata_collection belongs to, i.e., a link to the parent node.
+ $ref: "_definitions.yaml#/to_one_project"
\ No newline at end of file
diff --git a/datadictionary/gdcdictionary/schemas/_definitions.yaml b/datadictionary/gdcdictionary/schemas/_definitions.yaml
index 5308ae2d..1af65b9d 100644
--- a/datadictionary/gdcdictionary/schemas/_definitions.yaml
+++ b/datadictionary/gdcdictionary/schemas/_definitions.yaml
@@ -18,8 +18,8 @@ foreign_key_project:
# Allow true here because we can have other unique keys defined on
# a target type
additionalProperties: true
- # Can either use 'id' which are Gen3 IDs (UUID) or 'code'
- # which is the user defined ID for project
+ #Can either use 'id' which are Gen3 IDs (UUID) or 'code'
+ #which is the user defined ID for project
properties:
id:
$ref: "#/UUID"
@@ -48,8 +48,8 @@ foreign_key:
# Allow true here because we can have other unique keys defined on
# a target type
additionalProperties: True
- # Can either use 'id' which are GDC IDs (UUID) or 'submitter_id'
- # which are user defined IDs ("submitter IDs in the backend")
+ #Can either use 'id' which are GDC IDs (UUID) or 'submitter_id'
+ #which are user defined IDs ("submitter IDs in the backend")
properties:
id:
$ref: "#/UUID"
@@ -88,7 +88,7 @@ file_name:
file_size:
type: integer
- term:
+ term:
$ref: "_terms.yaml#/file_size"
file_format:
@@ -96,11 +96,16 @@ file_format:
term:
$ref: "_terms.yaml#/file_format"
+ga4gh_drs_uri:
+ type: string
+ term:
+ $ref: "_terms.yaml#/ga4gh_drs_uri"
+
md5sum:
type: string
- pattern: "^[a-f0-9]{32}$"
term:
$ref: "_terms.yaml#/md5sum"
+ pattern: "^[a-f0-9]{32}$"
object_id:
type: string
@@ -135,7 +140,7 @@ data_file_error_type:
state:
term:
- $ref: "_terms.yaml#/state"
+ $ref: "_terms.yaml#/state"
default: validated
downloadable:
- uploaded
@@ -166,7 +171,7 @@ state:
file_state:
term:
- $ref: "_terms.yaml#/file_state"
+ $ref: "_terms.yaml#/file_state"
default: registered
enum:
- registered
@@ -194,49 +199,26 @@ project_id:
$ref: "_terms.yaml#/project_id"
data_file_properties:
- id:
- $ref: "#/UUID"
- systemAlias: node_id
- submitter_id:
- type:
- - string
- - "null"
- description: "The file ID assigned by the submitter." # TOREVIEW
+ $ref: "#/ubiquitous_properties"
file_name:
$ref: "#/file_name"
file_size:
$ref: "#/file_size"
+ file_format:
+ $ref: "#/file_format"
md5sum:
- $ref: "#/md5sum"
- file_state:
- $ref: "#/file_state"
+ $ref: "#/md5sum"
object_id:
$ref: "#/object_id"
- state:
- $ref: "#/state"
+ file_state:
+ $ref: "#/file_state"
error_type:
$ref: "#/data_file_error_type"
- state_comment:
- type: string
- description: >
- Optional comment about why the file is in the
- current state, mainly for invalid state.
- project_id:
- $ref: "#/project_id"
- created_datetime:
- $ref: "#/datetime"
- updated_datetime:
- $ref: "#/datetime"
+ ga4gh_drs_uri:
+ $ref: "#/ga4gh_drs_uri"
workflow_properties:
- id:
- $ref: "#/UUID"
- systemAlias: node_id
- submitter_id:
- type:
- - string
- - "null"
- description: "The file ID assigned by the submitter." # TOREVIEW
+ $ref: "#/ubiquitous_properties"
workflow_link:
description: "Link to Github hash for the CWL workflow used."
type: string
@@ -247,14 +229,6 @@ workflow_properties:
$ref: "#/datetime"
workflow_end_datetime:
$ref: "#/datetime"
- state:
- $ref: "#/state"
- project_id:
- $ref: "#/project_id"
- created_datetime:
- $ref: "#/datetime"
- updated_datetime:
- $ref: "#/datetime"
ubiquitous_properties:
type:
@@ -276,3 +250,15 @@ ubiquitous_properties:
$ref: "#/datetime"
updated_datetime:
$ref: "#/datetime"
+
+
+fhir_resource:
+ type: object
+ # Allow true here because we can have other unique keys defined on
+ # a target type
+ additionalProperties: true
+ properties:
+ id:
+ $ref: "#/UUID"
+ resource_type:
+ type: string
diff --git a/datadictionary/gdcdictionary/schemas/program.yaml b/datadictionary/gdcdictionary/schemas/_program.yaml
similarity index 100%
rename from datadictionary/gdcdictionary/schemas/program.yaml
rename to datadictionary/gdcdictionary/schemas/_program.yaml
diff --git a/datadictionary/gdcdictionary/schemas/project.yaml b/datadictionary/gdcdictionary/schemas/_project.yaml
similarity index 100%
rename from datadictionary/gdcdictionary/schemas/project.yaml
rename to datadictionary/gdcdictionary/schemas/_project.yaml
diff --git a/datadictionary/gdcdictionary/schemas/_terms.yaml b/datadictionary/gdcdictionary/schemas/_terms.yaml
index 0ca296ba..44ee3c19 100644
--- a/datadictionary/gdcdictionary/schemas/_terms.yaml
+++ b/datadictionary/gdcdictionary/schemas/_terms.yaml
@@ -1,624 +1,26 @@
id: _terms
-28s_16s_ribosomal_rna_ratio:
- description: >
- The 28S/18S ribosomal RNA band ratio used to assess the quality of total RNA.
- termDef:
- term: "28s/18s Ribosomal RNA Ratio"
- source: null
- cde_id: null
- cde_version: null
- term_url: null
-
-a260_a280_ratio:
- description: >
- Numeric value that represents the sample ratio of nucleic acid absorbance at 260 nm and 280 nm,
- used to determine a measure of DNA purity.
- termDef:
- term: Nucleic Acid Absorbance at 260 And Absorbance at 280 DNA Purity Ratio Value
- source: caDSR
- cde_id: 5432595
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432595&version=1.0"
-
-adapter_name: # TOREVIEW
- description: >
- Name of the sequencing adapter.
-
-adapter_sequence: # TOREVIEW
- description: >
- Base sequence of the sequencing adapter.
-
-age_at_diagnosis:
- description: >
- Age at the time of diagnosis expressed in number of days since birth.
- termDef:
- term: Patient Diagnosis Age Day Value
- source: caDSR
- cde_id: 3225640
- cde_version: 2.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3225640&version=2.0"
-
-ajcc_clinical_m:
- description: >
- Extent of the distant metastasis for the cancer based on evidence obtained from clinical
- assessment parameters determined prior to treatment.
- termDef:
- term: Neoplasm American Joint Committee on Cancer Clinical Distant Metastasis M Stage
- source: caDSR
- cde_id: 3440331
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3440331&version=1.0"
-
-ajcc_clinical_n:
- description: >
- Extent of the regional lymph node involvement for the cancer based on evidence obtained from
- clinical assessment parameters determined prior to treatment.
- termDef:
- term: Neoplasm American Joint Committee on Cancer Clinical Regional Lymph Node N Stage
- source: caDSR
- cde_id: 3440330
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3440330&version=1.0"
-
-ajcc_clinical_stage:
- description: >
- Stage group determined from clinical information on the tumor (T), regional node (N) and
- metastases (M) and by grouping cases with similar prognosis for cancer.
- termDef:
- term: Neoplasm American Joint Committee on Cancer Clinical Group Stage
- source: caDSR
- cde_id: 3440332
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3440332&version=1.0"
-
-ajcc_clinical_t:
- description: >
- Extent of the primary cancer based on evidence obtained from clinical assessment parameters
- determined prior to treatment.
- termDef:
- term: Neoplasm American Joint Committee on Cancer Clinical Primary Tumor T Stage
- source: caDSR
- cde_id: 3440328
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3440328&version=1.0"
-
-ajcc_pathologic_m:
- description: >
- Code to represent the defined absence or presence of distant spread or metastases (M) to
- locations via vascular channels or lymphatics beyond the regional lymph nodes, using
- criteria established by the American Joint Committee on Cancer (AJCC).
- termDef:
- term: American Joint Committee on Cancer Metastasis Stage Code
- source: caDSR
- cde_id: 3045439
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3045439&version=1.0"
-
-ajcc_pathologic_n:
- description: >
- The codes that represent the stage of cancer based on the nodes present (N stage) according
- to criteria based on multiple editions of the AJCC's Cancer Staging Manual.
- termDef:
- term: Neoplasm Disease Lymph Node Stage American Joint Committee on Cancer Code
- source: caDSR
- cde_id: 3203106
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3203106&version=1.0"
-
-ajcc_pathologic_stage:
- description: >
- The extent of a cancer, especially whether the disease has spread from the original site to
- other parts of the body based on AJCC staging criteria.
- termDef:
- term: Neoplasm Disease Stage American Joint Committee on Cancer Code
- source: caDSR
- cde_id: 3203222
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3203222&version=1.0"
-
-ajcc_pathologic_t:
- description: >
- Code of pathological T (primary tumor) to define the size or contiguous extension of the
- primary tumor (T), using staging criteria from the American Joint Committee on Cancer
- (AJCC).
- termDef:
- term: American Joint Committee on Cancer Tumor Stage Code
- source: caDSR
- cde_id: 3045435
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3045435&version=1.0"
-
-alcohol_history:
- description: >
- A response to a question that asks whether the participant has consumed at least 12 drinks of
- any kind of alcoholic beverage in their lifetime.
- termDef:
- term: Alcohol Lifetime History Indicator
- source: caDSR
- cde_id: 2201918
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2201918&version=1.0"
-
-alcohol_intensity:
- description: >
- Category to describe the patient's current level of alcohol use as self-reported by the patient.
- termDef:
- term: Person Self-Report Alcoholic Beverage Exposure Category
- source: caDSR
- cde_id: 3457767
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3457767&version=1.0"
-
-aliquot_quantity:
- description: >
- The quantity in micrograms (ug) of the aliquot(s) derived from the analyte(s) shipped for
- sequencing and characterization.
- termDef:
- term: Biospecimen Aliquot Quantity
- source: null
- cde_id: null
- cde_version: null
- term_url: null
-
-aliquot_volume:
- description: >
- The volume in microliters (ml) of the aliquot(s) derived from the analyte(s) shipped for
- sequencing and characterization.
- termDef:
- term: Biospecimen Aliquot Volume
- source: null
- cde_id: null
- cde_version: null
- term_url: null
-
-amount: # TOREVIEW
- description: >
- Weight in grams or volume in mL.
-
-analyte_quantity:
- description: >
- The quantity in micrograms (ug) of the analyte(s) derived from the analyte(s) shipped for
- sequencing and characterization.
- termDef:
- term: Biospecimen Analyte Quantity
- source: null
- cde_id: null
- cde_version: null
- term_url: null
-
-analyte_type:
- description: >
- Text term that represents the kind of molecular specimen analyte.
- termDef:
- term: Molecular Specimen Type Text Name
- source: caDSR
- cde_id: 2513915
- cde_version: 2.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2513915&version=2.0"
-
-analyte_type_id:
- description: >
- A single letter code used to identify a type of molecular analyte.
- termDef:
- term: Molecular Analyte Identification Code
- source: caDSR
- cde_id: 5432508
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432508&version=1.0"
-
-analyte_volume:
- description: >
- The volume in microliters (ml) of the analyte(s) derived from the analyte(s) shipped for
- sequencing and characterization.
- termDef:
- term: Biospecimen Analyte Volume
- source: null
- cde_id: null
- cde_version: null
- term_url: null
-
-ann_arbor_b_symptoms:
- description: >
- Text term to signify whether lymphoma B-symptoms are present as noted in the patient's medical
- record.
- termDef:
- term: Lymphoma B-Symptoms Medical Record Documented Indicator
- source: caDSR
- cde_id: 2902402
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2902402&version=1.0"
-
-ann_arbor_clinical_stage:
- description: >
- The classification of the clinically confirmed anatomic disease extent of lymphoma (Hodgkin's
- and Non-Hodgkins) based on the Ann Arbor Staging System.
- termDef:
- term: Ann Arbor Clinical Stage
- source: null
- cde_id: null
- cde_version: null
- term_url: null
-
-ann_arbor_extranodal_involvement:
- description: >
- Indicator that identifies whether a patient with malignant lymphoma has lymphomatous involvement
- of an extranodal site.
- termDef:
- term: Lymphomatous Extranodal Site Involvement Indicator
- source: caDSR
- cde_id: 3364582
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3364582&version=1.0"
-
-ann_arbor_pathologic_stage:
- description: >
- The classification of the pathologically confirmed anatomic disease extent of lymphoma
- (Hodgkin's and Non-Hodgkins) based on the Ann Arbor Staging System.
- termDef:
- term: Ann Arbor Pathologic Stage
- source: null
- cde_id: null
- cde_version: null
- term_url: null
-
-ann_arbor_tumor_stage:
- description: >
- The classification of the anatomic disease extent of lymphoma (Hodgkin's and Non-Hodgkins) based
- on the Ann Arbor Staging System.
- termDef:
- term: Ann Arbor Tumor Stage
- source: null
- cde_id: null
- cde_version: null
- term_url: null
-
-base_caller_name: # TOREVIEW
- description: >
- Name of the base caller.
-
-base_caller_version: # TOREVIEW
- description: >
- Version of the base caller.
-
-biomarker_name:
- description: >
- The name of the biomarker being tested for this specimen and set of test results.
- termDef:
- term: Biomarker Name
- source: caDSR
- cde_id: 5473
- cde_version: 11.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5473&version=2.31"
-
-biomarker_result:
- description: >
- Text term to define the results of genetic testing.
- termDef:
- term: Laboratory Procedure Genetic Abnormality Test Result Type
- source: caDSR
- cde_id: 3234680
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3234680&version=1.0"
-
-biomarker_test_method:
- description: >
- Text descriptor of a molecular analysis method used for an individual.
- termDef:
- term: Disease Detection Molecular Analysis Method Type
- source: caDSR
- cde_id: 3121575
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3121575&version=1.0"
-
-biospecimen_anatomic_site:
- description: >
- Text term that represents the name of the primary disease site of the submitted tumor sample.
- termDef:
- term: Submitted Tumor Sample Primary Anatomic Site
- source: caDSR
- cde_id: 4742851
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=4742851&version=1.0"
-
-bmi:
- description: >
- The body mass divided by the square of the body height expressed in units of kg/m^2.
- termDef:
- term: Body Mass Index (BMI)
- source: caDSR
- cde_id: 4973892
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=4973892&version=1.0"
-
-burkitt_lymphoma_clinical_variant:
- description: >
- Burkitt's lymphoma categorization based on clinical features that differ from other forms of the
- same disease.
- termDef:
- term: Burkitt Lymphoma Clinical Variant Type
- source: caDSR
- cde_id: 3770421
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3770421&version=1.0"
-
-cause_of_death:
- description: >
- Text term to identify the cause of death for a patient.
- termDef:
- term: Patient Death Reason
- source: caDSR
- cde_id: 2554674
- cde_version: 3.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2554674&version=3.0"
-
-cea_level_preoperative:
- description: >
- Numeric value of the Carcinoembryonic antigen or CEA at the time before surgery. [Manually-
- curated]
- termDef:
- term: Preoperative Carcinoembryonic Antigen Result Value
- source: caDSR
- cde_id: 2716510
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2716510&version=1.0"
-
-cigarettes_per_day:
- description: >
- The average number of cigarettes smoked per day.
- termDef:
- term: Smoking Use Average Number
- source: caDSR
- cde_id: 2001716
- cde_version: 4.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2001716&version=4.0"
-
-circumferential_resection_margin:
- description: >
- A value in millimeters indicating the measured length between a malignant lesion of the
- colon or rectum and the nearest radial (or circumferential) border of tissue removed during
- cancer surgery.
- termDef:
- term: Colorectal Surgical Margin Circumferential Distance Measurement
- source: caDSR
- cde_id: 64202
- cde_version: 3.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=64202&version=3.0"
-
-classification_of_tumor:
- description: >
- Text that describes the kind of disease present in the tumor specimen as related to a specific
- timepoint.
- termDef:
- term: Tumor Tissue Disease Description Type
- source: caDSR
- cde_id: 3288124
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3288124&version=1.0"
-
-colon_polyps_history:
- description: >
- Yes/No indicator to describe if the subject had a previous history of colon polyps as noted
- in the history/physical or previous endoscopic report (s).
- termDef:
- term: Colon Carcinoma Polyp Occurrence Indicator
- source: caDSR
- cde_id: 3107197
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3107197&version=1.0"
-
-composition:
- description: >
- Text term that represents the cellular composition of the sample.
- termDef:
- term: Biospecimen Cellular Composition Type
- source: caDSR
- cde_id: 5432591
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432591&version=1.0"
-
-concentration:
- description: >
- Numeric value that represents the concentration of an analyte or aliquot extracted from the
- sample or sample portion, measured in milligrams per milliliter.
- termDef:
- term: Biospecimen Analyte or Aliquot Extracted Concentration Milligram per Milliliter Value
- source: caDSR
- cde_id: 5432594
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432594&version=1.0"
-
-creation_datetime:
- description: >
- The datetime of portion creation encoded as seconds from epoch.
- termDef:
- term: Biospecimen Portion Creation Seconds Date/Time
- source: caDSR
- cde_id: 5432592
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432592&version=1.0"
-
-current_weight:
- description: >
- Numeric value that represents the current weight of the sample, measured in milligrams.
- termDef:
- term: Tissue Sample Current Weight Milligram Value
- source: caDSR
- cde_id: 5432606
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432606&version=1.0"
-
data_category: # TOREVIEW
- description: >
- Broad categorization of the contents of the data file.
-
-data_file_error_type: # TOREVIEW
- description: >
- Type of error for the data file object.
-
-data_format: # TOREVIEW
- description: >
- Format of the data files.
-
-data_type: # TOREVIEW
- description: >
- Specific content type of the data file.
-
-datetime:
- description: >
- A combination of date and time of day in the form [-]CCYY-MM-DDThh:mm:ss[Z|(+|-)hh:mm]
-
-days_to_birth:
- description: >
- Time interval from a person's date of birth to the date of initial pathologic diagnosis,
- represented as a calculated negative number of days.
- termDef:
- term: Person Birth Date Less Initial Pathologic Diagnosis Date Calculated Day Value
- source: caDSR
- cde_id: 3008233
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3008233&version=1.0"
-
-days_to_collection:
- description: >
- Time interval from the date of biospecimen collection to the date of initial pathologic
- diagnosis, represented as a calculated number of days.
- termDef:
- term: Biospecimen Collection Date Less Initial Pathologic Diagnosis Date Calculated Day Value
- source: caDSR
- cde_id: 3008340
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3008340&version=1.0"
-
-days_to_death:
- description: >
- Time interval from a person's date of death to the date of initial pathologic diagnosis,
- represented as a calculated number of days.
- termDef:
- term: Death Less Initial Pathologic Diagnosis Date Calculated Day Value
- source: caDSR
- cde_id: 3165475
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3165475&version=1.0"
-
-days_to_hiv_diagnosis:
- description: >
- Time interval from the date of the initial pathologic diagnosis to the date of human
- immunodeficiency diagnosis, represented as a calculated number of days.
- termDef:
- term: Human Immunodeficiency Virus Diagnosis Subtract Initial Pathologic Diagnosis Time Duration Day Calculation Value
- source: caDSR
- cde_id: 4618491
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=4618491&version=1.0"
-
-days_to_last_follow_up:
- description: >
- Time interval from the date of last follow up to the date of initial pathologic diagnosis,
- represented as a calculated number of days.
- termDef:
- term: Last Communication Contact Less Initial Pathologic Diagnosis Date Calculated Day Value
- source: caDSR
- cde_id: 3008273
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3008273&version=1.0"
-
-days_to_last_known_disease_status:
- description: >
- Time interval from the date of last follow up to the date of initial pathologic diagnosis,
- represented as a calculated number of days.
- termDef:
- term: Last Communication Contact Less Initial Pathologic Diagnosis Date Calculated Day Value
- source: caDSR
- cde_id: 3008273
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3008273&version=1.0"
-
-days_to_new_event:
- description: >
- Time interval from the date of new tumor event including progression, recurrence and new
- primary malignacies to the date of initial pathologic diagnosis, represented as a calculated
- number of days.
- termDef:
- term: New Tumor Event Less Initial Pathologic Diagnosis Date Calculated Day Value
- source: caDSR
- cde_id: 3392464
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3392464&version=1.0"
-
-days_to_recurrence:
- description: >
- Time interval from the date of new tumor event including progression, recurrence and new primary
- malignancies to the date of initial pathologic diagnosis, represented as a calculated number of
- days.
- termDef:
- term: New Tumor Event Less Initial Pathologic Diagnosis Date Calculated Day Value
- source: caDSR
- cde_id: 3392464
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3392464&version=1.0"
-
-days_to_sample_procurement:
- description: >
- The number of days from the date the patient was diagnosed to the date of the procedure that
- produced the sample.
-
-days_to_treatment:
- description: >
- Number of days from date of initial pathologic diagnosis that treatment began.
- termDef:
- term: Days to Treatment Start
- source: null
- cde_id: null
- cde_version: null
- term_url: null
+ description: >
+ Broad categorization of the contents of the data file.
-days_to_treatment_end:
+data_file_error_type: # TOREVIEW
description: >
- Time interval from the date of the initial pathologic diagnosis to the date of treatment end,
- represented as a calculated number of days.
- termDef:
- term: Treatment End Subtract First Pathologic Diagnosis Day Calculation Value
- source: caDSR
- cde_id: 5102431
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5102431&version=1.0"
+ Type of error for the data file object.
-days_to_treatment_start:
+data_format: # TOREVIEW
description: >
- Time interval from the date of the initial pathologic diagnosis to the start of treatment,
- represented as a calculated number of days.
- termDef:
- term: Treatment Start Subtract First Pathologic Diagnosis Time Day Calculation Value
- source: caDSR
- cde_id: 5102411
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5102411&version=1.0"
+ Format of the data files.
-diagnosis_pathologically_confirmed:
+data_type: # TOREVIEW
description: >
- The histologic description of tissue or cells confirmed by a pathology review of frozen or
- formalin fixed slide(s) completed after the diagnostic pathology review of the tumor sample used
- to extract analyte(s).
- termDef:
- term: Post-Diagnostic Pathology Review Confirmation
- source: null
- cde_id: null
- cde_version: null
- term_url: null
+ Specific content type of the data file. (CMG)
-dlco_ref_predictive_percent:
+datetime:
description: >
- The value, as a percentage of predicted lung volume, measuring the amount of carbon monoxide
- detected in a patient's lungs.
- termDef:
- term: Lung Carbon Monoxide Diffusing Capability Test Assessment Predictive Value Percentage Value
- source: caDSR
- cde_id: 2180255
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2180255&version=1.0"
+ A combination of date and time of day in the form [-]CCYY-MM-DDThh:mm:ss[Z|(+|-)hh:mm]
-encoding:
+encoding:
description: >
Version of ASCII encoding of quality values found in the file.
termDef:
@@ -628,105 +30,9 @@ encoding:
cde_version: null
term_url: "http://www.bioinformatics.babraham.ac.uk/projects/fastqc/Help/3%20Analysis%20Modules/1%20Basic%20Statistics.html"
-estrogen_receptor_percent_positive_ihc:
- description: >
- Classification to represent ER Positive results expressed as a percentage value.
- termDef:
- term: ER Level Cell Percentage Category
- source: caDSR
- cde_id: 3128341
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3128341&version=1.0"
-
-estrogen_receptor_result_ihc:
- description: >
- Text term to represent the overall result of Estrogen Receptor (ER) testing.
- termDef:
- term: Breast Carcinoma Estrogen Receptor Status
- source: caDSR
- cde_id: 2957359
- cde_version: 2.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2957359&version=2.0"
-
-ethnicity:
- description: >
- An individual's self-described social and cultural grouping, specifically whether an individual
- describes themselves as Hispanic or Latino. The provided values are based on the categories
- defined by the U.S. Office of Management and Business and used by the U.S. Census Bureau.
- termDef:
- term: Ethnic Group Category Text
- source: caDSR
- cde_id: 2192217
- cde_version: 2.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2192217&version=2.0"
-
-experiment_name: # TOREVIEW
- description: >
- Submitter-defined name for the experiment.
-
-experimental_strategy: # TOREVIEW
- description: >
- The sequencing strategy used to generate the data file.
-
-fastq_name: # TOREVIEW
- description: >
- Names of FASTQs.
-
-fev1_ref_post_bronch_percent:
- description: >
- The percentage comparison to a normal value reference range of the volume of air that a
- patient can forcibly exhale from the lungs in one second post-bronchodilator.
- termDef:
- term: Post Bronchodilator Lung Forced Expiratory Volume 1 Test Lab Percentage Value
- source: caDSR
- cde_id: 3302948
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3302948&version=1.0"
-
-fev1_ref_pre_bronch_percent:
- description: >
- The percentage comparison to a normal value reference range of the volume of air that a
- patient can forcibly exhale from the lungs in one second pre-bronchodilator.
- termDef:
- term: Pre Bronchodilator Lung Forced Expiratory Volume 1 Test Lab Percentage Value
- source: caDSR
- cde_id: 3302947
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3302947&version=1.0"
-
-fev1_fvc_post_bronch_percent:
- description: >
- Percentage value to represent result of Forced Expiratory Volume in 1 second (FEV1) divided
- by the Forced Vital Capacity (FVC) post-bronchodilator.
- termDef:
- term: Post Bronchodilator FEV1/FVC Percent Value
- source: caDSR
- cde_id: 3302956
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3302956&version=1.0"
-
-fev1_fvc_pre_bronch_percent:
- description: >
- Percentage value to represent result of Forced Expiratory Volume in 1 second (FEV1) divided
- by the Forced Vital Capacity (FVC) pre-bronchodilator.
- termDef:
- term: Pre Bronchodilator FEV1/FVC Percent Value
- source: caDSR
- cde_id: 3302955
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3302955&version=1.0"
-
-figo_stage:
+file_format:
description: >
- The extent of a cervical or endometrial cancer within the body, especially whether the
- disease has spread from the original site to other parts of the body, as described by the
- International Federation of Gynecology and Obstetrics (FIGO) stages.
- termDef:
- term: Gynecologic Tumor Grouping Cervical Endometrial FIGO 2009 Stage
- source: caDSR
- cde_id: 3225684
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3225684&version=1.0"
+ The format of the data file object.
file_name: # TOREVIEW
description: >
@@ -740,278 +46,33 @@ file_state: # TOREVIEW
description: >
The current state of the data file object.
-flow_cell_barcode: # TOREVIEW
- description: >
- Flow Cell Barcode.
-
-freezing_method:
- description: >
- Text term that represents the method used for freezing the sample.
- termDef:
- term: Tissue Sample Freezing Method Type
- source: caDSR
- cde_id: 5432607
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432607&version=1.0"
-
-gender:
- description: >
- Text designations that identify gender. Gender is described as the assemblage of properties that
- distinguish people on the basis of their societal roles. [Explanatory Comment 1: Identification
- of gender is based upon self-report and may come from a form, questionnaire, interview, etc.]
- termDef:
- term: Person Gender Text Type
- source: caDSR
- cde_id: 2200604
- cde_version: 3.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2200604&version=3.0"
-
-height:
- description: >
- The height of the patient in centimeters.
- termDef:
- term: Patient Height Measurement
- source: caDSR
- cde_id: 649
- cde_version: 4.1
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=649&version=4.1"
-
-her2_erbb2_percent_positive_ihc:
- description: >
- Classification to represent the number of positive HER2/ERBB2 cells in a specimen or sample.
- termDef:
- term: HER2 ERBB Positive Finding Cell Percentage Category
- source: caDSR
- cde_id: 3086980
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3086980&version=1.0"
-
-her2_erbb2_result_fish:
- description: >
- the type of outcome for HER2 as determined by an in situ hybridization (ISH) assay.
- termDef:
- term: Laboratory Procedure HER2/neu in situ Hybridization Outcome Type
- source: caDSR
- cde_id: 2854089
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2854089&version=1.0"
-
-her2_erbb2_result_ihc:
- description: >
- Text term to signify the result of the medical procedure that involves testing a sample of
- blood or tissue for HER2 by histochemical localization of immunoreactive substances using
- labeled antibodies as reagents.
- termDef:
- term: Laboratory Procedure HER2/neu Immunohistochemistry Receptor Status
- source: caDSR
- cde_id: 2957563
- cde_version: 2.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2957563&version=2.0"
-
-hiv_positive:
- description: >
- Text term to signify whether a physician has diagnosed HIV infection in a patient.
- termDef:
- term: Physician Diagnosed HIV Infection Personal Medical History Yes No Not Applicable Indicator
- source: caDSR
- cde_id: 4030799
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=4030799&version=1.0"
-
-hpv_positive_type:
- description: >
- Text classification to represent the strain or type of human papillomavirus identified in an
- individual.
- termDef:
- term: Human Papillomavirus Type
- source: caDSR
- cde_id: 2922649
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2922649&version=1.0"
-
-hpv_status:
- description: >
- The findings of the oncogenic HPV.
- termDef:
- term: Oncogenic Human Papillomavirus Result Type
- source: caDSR
- cde_id: 2230033
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2230033&version=1.0"
-
-includes_spike_ins: # TOREVIEW
- description: >
- Spike-in included?
-
-initial_weight:
- description: >
- Numeric value that represents the initial weight of the sample, measured in milligrams.
- termDef:
- term: Tissue Sample Initial Weight Milligram Value
- source: caDSR
- cde_id: 5432605
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432605&version=1.0"
-
-instrument_model:
- description: >
- Numeric value that represents the sample dimension that is greater than the shortest
- dimension and less than the longest dimension, measured in millimeters.
- termDef:
- term: Tissue Sample Intermediate Dimension Millimeter Measurement
- source: caDSR
- cde_id: 5432604
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432604&version=1.0"
-
-intermediate_dimension: # TOREVIEW
- description: >
- Intermediate dimension of the sample, in millimeters.
-
-is_ffpe:
- description: >
- Indicator to signify whether or not the tissue sample was fixed in formalin and embedded in
- paraffin (FFPE).
- termDef:
- term: Specimen Processing Formalin Fixed Paraffin Embedded Tissue Indicator
- source: caDSR
- cde_id: 4170557
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=4170557&version=1.0"
-
-is_paired_end: # TOREVIEW
- description: >
- Are the reads paired end?
-
-last_known_disease_status:
- description: >
- Text term that describes the last known state or condition of an individual's neoplasm.
- termDef:
- term: Person Last Known Neoplasm Status
- source: caDSR
- cde_id: 5424231
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2759550&version=1.0"
-
-laterality:
- description: >
- For tumors in paired organs, designates the side on which the cancer originates.
- termDef:
- term: Primary Tumor Laterality
- source: caDSR
- cde_id: 827
- cde_version: 3.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=827&version=3.0"
-
-ldh_level_at_diagnosis:
- description: >
- The 2 decimal place numeric laboratory value measured, assigned or computed related to the
- assessment of lactate dehydrogenase in a specimen.
- termDef:
- term: Laboratory Procedure Lactate Dehydrogenase Result Integer::2 Decimal Place Value
- source: caDSR
- cde_id: 2798766
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2798766&version=1.0"
-
-ldh_normal_range_upper:
- description: >
- The top value of the range of statistical characteristics that are supposed to represent
- accepted standard, non-pathological pattern for lactate dehydrogenase (units not specified).
- termDef:
- term: Laboratory Procedure Lactate Dehydrogenase Result Upper Limit of Normal Value
- source: caDSR
- cde_id: 2597015
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2597015&version=1.0"
-
-library_strategy: # TOREVIEW
- description: >
- Library strategy.
-
-library_selection: # TOREVIEW
- description: >
- Library Selection Method
-
-library_name: # TOREVIEW
- description: >
- Name of the library.
-
-library_preparation_kit_name: # TOREVIEW
- description: >
- Name of Library Preparation Kit
-
-library_preparation_kit_vendor: # TOREVIEW
- description: >
- Vendor of Library Preparation Kit
-
-library_preparation_kit_catalog_number: # TOREVIEW
- description: >
- Catalog of Library Preparation Kit
-
-library_preparation_kit_version: # TOREVIEW
- description: >
- Version of Library Preparation Kit
-
-library_strand: # TOREVIEW
- description: >
- Library stranded-ness.
-
-longest_dimension:
+md5sum: # TOREVIEW
description: >
- Numeric value that represents the longest dimension of the sample, measured in millimeters.
- termDef:
- term: Tissue Sample Longest Dimension Millimeter Measurement
- source: caDSR
- cde_id: 5432602
- cde_version: 1.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432602&version=1.0"
+ The 128-bit hash value expressed as a 32 digit hexadecimal number used as a file's digital
+ fingerprint.
-lymph_nodes_positive:
+project_id: # TOREVIEW
description: >
- The number of lymph nodes involved with disease as determined by pathologic examination.
- termDef:
- term: Lymph Node(s) Positive Number
- source: caDSR
- cde_id: 89
- cde_version: 3.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=89&version=3.0"
+ Unique ID for any specific defined piece of work that is undertaken or attempted to
+ meet a single requirement.
-lymphatic_invasion_present:
+state: # TOREVIEW
description: >
- A yes/no indicator to ask if small or thin-walled vessel invasion is present, indicating
- lymphatic involvement
- termDef:
- term: Lymphatic/Small vessel Invasion Ind
- source: caDSR
- cde_id: 64171
- cde_version: 3.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=64171&version=3.0"
+ The current state of the object.
-method_of_diagnosis:
- description: >
- The method used to initially the patient's diagnosis.
- termDef:
- term: Method of Diagnosis
- source: null
- cde_id: null
- cde_version: null
- term_url: null
-method_of_sample_procurement:
+UUID:
description: >
- The method used to procure the sample used to extract analyte(s).
+ A 128-bit identifier. Depending on the mechanism used to generate it, it is either guaranteed to
+ be different from all other UUIDs/GUIDs generated until 3400 AD or extremely likely to be
+ different. Its relatively small size lends itself well to sorting, ordering, and hashing of all
+ sorts, storing in databases, simple allocation, and ease of programming in general.
termDef:
- term: Method of Sample Procurement
- source: null
- cde_id: null
+ term: Universally Unique Identifier
+ source: NCIt
+ cde_id: C54100
cde_version: null
- term_url: null
-
-md5sum: # TOREVIEW
- description: >
- The 128-bit hash value expressed as a 32 digit hexadecimal number used as a file's digital
- fingerprint.
+ term_url: "https://ncit.nci.nih.gov/ncitbrowser/ConceptReport.jsp?dictionary=NCI_Thesaurus&version=16.02d&ns=NCI_Thesaurus&code=C54100"
microsatellite_instability_abnormal:
description: >
@@ -1091,10 +152,9 @@ oct_embedded:
cde_version: 1.0
term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432538&version=1.0"
-pack_years_smoked:
+years_smoked:
description: >
- Numeric computed value to represent lifetime tobacco exposure defined as number of cigarettes
- smoked per day x number of years smoked divided by 20.
+ Number of years the Donor smoked. (GTEx)
termDef:
term: Person Cigarette Smoking History Pack Year Value
source: caDSR
@@ -1275,15 +335,6 @@ preservation_method:
cde_version: 1.0
term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5432521&version=1.0"
-primary_diagnosis:
- description: >
- Text term for the structural pattern of cancer cells used to define a microscopic diagnosis.
- termDef:
- term: Neoplasm Histologic Type Name
- source: caDSR
- cde_id: 3081934
- cde_version: 3.0
- term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3081934&version=3.0"
prior_malignancy:
description: >
@@ -1339,10 +390,6 @@ progression_or_recurrence:
cde_version: 1.0
term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3121376&version=1.0"
-project_id: # TOREVIEW
- description: >
- Unique ID for any specific defined piece of work that is undertaken or attempted to
- meet a single requirement.
qc_metric_state:
description: >
@@ -1357,11 +404,11 @@ qc_metric_state:
race:
description: >
- An arbitrary classification of a taxonomic group that is a division of a species. It usually
+ An arbitrary classification of a taxonomic group that is a division of a species (HARMONIZED). It usually
arises as a consequence of geographical isolation within a species and is characterized by
shared heredity, physical attributes and behavior, and in the case of humans, by common history,
nationality, or geographic distribution. The provided values are based on the categories defined
- by the U.S. Office of Management and Business and used by the U.S. Census Bureau.
+ by the U.S. Office of Management and Business and used by the U.S. Census Bureau. (GTEx)
termDef:
term: Race Category Text
source: caDSR
@@ -1387,6 +434,17 @@ relationship_age_at_diagnosis:
cde_version: 1.0
term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=5300571&version=1.0"
+relationship_to_proband:
+ description: >
+ The subgroup that describes the state of connectedness between members of the unit of society
+ organized around kinship ties.
+ termDef:
+ term: Family Member Relationship Type
+ source: caDSR
+ cde_id: 2690165
+ cde_version: 2.0
+ term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=2690165&version=2.0"
+
relationship_type:
description: >
The subgroup that describes the state of connectedness between members of the unit of society
@@ -1524,9 +582,6 @@ spike_ins_concentration: # TOREVIEW
description: >
Spike in concentration.
-state: # TOREVIEW
- description: >
- The current state of the object.
target_capture_kit_name: # TOREVIEW
description: >
@@ -1741,19 +796,6 @@ tumor_stage:
cde_version: null
term_url: "https://ncit.nci.nih.gov/ncitbrowser/pages/concept_details.jsf?dictionary=NCI%20Thesaurus&code=C16899"
-UUID:
- description: >
- A 128-bit identifier. Depending on the mechanism used to generate it, it is either guaranteed to
- be different from all other UUIDs/GUIDs generated until 3400 AD or extremely likely to be
- different. Its relatively small size lends itself well to sorting, ordering, and hashing of all
- sorts, storing in databases, simple allocation, and ease of programming in general.
- termDef:
- term: Universally Unique Identifier
- source: NCIt
- cde_id: C54100
- cde_version: null
- term_url: "https://ncit.nci.nih.gov/ncitbrowser/ConceptReport.jsp?dictionary=NCI_Thesaurus&version=16.02d&ns=NCI_Thesaurus&code=C54100"
-
vascular_invasion_present:
description: >
The yes/no indicator to ask if large vessel or venous invasion was detected by surgery or
@@ -1777,7 +819,7 @@ vital_status:
weight:
description: >
- The weight of the patient measured in kilograms.
+ The weight of the patient measured in lbs (HARMONIZED).
termDef:
term: Patient Weight Measurement
source: caDSR
@@ -1832,10 +874,24 @@ year_of_death:
years_smoked:
description: >
- Numeric value (or unknown) to represent the number of years a person has been smoking.
+ Numeric value (or unknown) to represent the number of years a person has been smoking (HARMONIZED). If the number of years is greater than 89, see 'years_smoked_gt89'.
+ termDef:
+ term: Person Smoking Duration Year Count
+ source: caDSR
+ cde_id: 3137957
+ cde_version: 1.0
+ term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3137957&version=1.0"
+
+years_smoked_gt89:
+ description: >
+ Indicate whether the numeric value to represent the number of years a person has been smoking (HARMONIZED) is greater than 89 years.
termDef:
term: Person Smoking Duration Year Count
source: caDSR
cde_id: 3137957
cde_version: 1.0
term_url: "https://cdebrowser.nci.nih.gov/CDEBrowser/search?elementDetails=9&FirstTimer=0&PageId=ElementDetailsGroup&publicId=3137957&version=1.0"
+
+ga4gh_drs_uri:
+ description: >
+ DRS URI as defined by GA4GH DRS spec for pointers to file objects.
diff --git a/datadictionary/gdcdictionary/schemas/acknowledgement.yaml b/datadictionary/gdcdictionary/schemas/acknowledgement.yaml
deleted file mode 100644
index cee5d3d9..00000000
--- a/datadictionary/gdcdictionary/schemas/acknowledgement.yaml
+++ /dev/null
@@ -1,61 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "acknowledgement"
-title: Acknowledgement
-type: object
-namespace: http://gdc.nci.nih.gov
-category: administrative
-program: '*'
-project: '*'
-description: "Acknowledgement of an individual involved in a project."
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - state
- - created_datetime
- - updated_datetime
-
-links:
- - name: projects
- backref: acknowledgements
- label: contribute_to
- target_type: project
- multiplicity: many_to_many
- required: true
-
-required:
- - submitter_id
- - type
- - projects
-
-uniqueKeys:
- - [ id ]
- - [ project_id, submitter_id ]
-
-properties:
- type:
- enum: [ "acknowledgement" ]
- id:
- $ref: "_definitions.yaml#/UUID"
- systemAlias: node_id
- state:
- $ref: "_definitions.yaml#/state"
- submitter_id:
- type:
- - string
- - "null"
- acknowledgee:
- description: "The indvidiual or group being acknowledged by the project."
- type: string
- projects:
- $ref: "_definitions.yaml#/to_many_project"
- project_id:
- type: string
- created_datetime:
- $ref: "_definitions.yaml#/datetime"
- updated_datetime:
- $ref: "_definitions.yaml#/datetime"
diff --git a/datadictionary/gdcdictionary/schemas/aligned_reads_index.yaml b/datadictionary/gdcdictionary/schemas/aligned_reads_index.yaml
deleted file mode 100644
index e09ee3ed..00000000
--- a/datadictionary/gdcdictionary/schemas/aligned_reads_index.yaml
+++ /dev/null
@@ -1,77 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "aligned_reads_index"
-title: Aligned Reads Index
-type: object
-namespace: http://gdc.nci.nih.gov
-category: index_file
-program: '*'
-project: '*'
-description: "Data file containing the index for a set of aligned reads."
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - created_datetime
- - updated_datetime
- - state
- - file_state
- - error_type
-
-links:
- - exclusive: false
- required: true
- subgroup:
- - name: submitted_aligned_reads_files
- backref: aligned_reads_indexes
- label: derived_from
- target_type: submitted_aligned_reads
- multiplicity: one_to_one
- required: false
- - name: core_metadata_collections
- backref: aligned_reads_indexes
- label: data_from
- target_type: core_metadata_collection
- multiplicity: many_to_many
- required: false
-
-required:
- - submitter_id
- - type
- - file_name
- - file_size
- - md5sum
- - data_category
- - data_type
- - data_format
-
-uniqueKeys:
- - [ id ]
- - [ project_id, submitter_id ]
-
-properties:
- $ref: "_definitions.yaml#/data_file_properties"
- type:
- enum: [ "aligned_reads_index" ]
- data_category:
- term:
- $ref: "_terms.yaml#/data_category"
- enum:
- - Sequencing Data
- - Sequencing Reads
- - Raw Sequencing Data
- data_type:
- term:
- $ref: "_terms.yaml#/data_type"
- enum: [ "Aligned Reads Index" ]
- data_format:
- term:
- $ref: "_terms.yaml#/data_format"
- enum: [ "BAI" ]
- submitted_aligned_reads_files:
- $ref: "_definitions.yaml#/to_one"
- core_metadata_collections:
- $ref: "_definitions.yaml#/to_many"
diff --git a/datadictionary/gdcdictionary/schemas/aliquot.yaml b/datadictionary/gdcdictionary/schemas/aliquot.yaml
deleted file mode 100644
index 078cb17a..00000000
--- a/datadictionary/gdcdictionary/schemas/aliquot.yaml
+++ /dev/null
@@ -1,103 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "aliquot"
-title: Aliquot
-type: object
-category: biospecimen
-program: '*'
-project: '*'
-description: >
- Pertaining to a portion of the whole; any one of two or more samples of something, of the same
- volume or weight.
-additionalProperties: false
-submittable: true
-validators: []
-
-systemProperties:
- - id
- - project_id
- - state
- - created_datetime
- - updated_datetime
-
-required:
- - submitter_id
- - type
- - samples
-
-uniqueKeys:
- - [id]
- - [project_id, submitter_id]
-
-links:
- - name: samples
- backref: aliquots
- label: derived_from
- multiplicity: many_to_many
- target_type: sample
- required: true
-
-constraints: null
-
-# Aliquot properties
-properties:
- type:
- type: string
- id:
- $ref: "_definitions.yaml#/UUID"
- systemAlias: node_id
- state:
- $ref: "_definitions.yaml#/state"
- submitter_id:
- type:
- - string
- - "null"
- description: >
- The legacy barcode used before prior to the use
- UUIDs. For TCGA this is bcraliquotbarcode.
- aliquot_quantity:
- term:
- $ref: "_terms.yaml#/aliquot_quantity"
- type: number
- aliquot_volume:
- term:
- $ref: "_terms.yaml#/aliquot_volume"
- type: number
- amount:
- term:
- $ref: "_terms.yaml#/amount"
- type: number
- analyte_type:
- term:
- $ref: "_terms.yaml#/analyte_type"
- type: string
- analyte_type_id:
- term:
- $ref: "_terms.yaml#/analyte_type_id"
- enum:
- - D
- - E
- - G
- - H
- - R
- - S
- - T
- - W
- - X
- - Y
- concentration:
- term:
- $ref: "_terms.yaml#/concentration"
- type: number
- project_id:
- $ref: "_definitions.yaml#/project_id"
- source_center:
- term:
- $ref: "_terms.yaml#/source_center"
- type: string
- samples:
- $ref: "_definitions.yaml#/to_one"
- created_datetime:
- $ref: "_definitions.yaml#/datetime"
- updated_datetime:
- $ref: "_definitions.yaml#/datetime"
diff --git a/datadictionary/gdcdictionary/schemas/case.yaml b/datadictionary/gdcdictionary/schemas/case.yaml
deleted file mode 100644
index 3db55f5a..00000000
--- a/datadictionary/gdcdictionary/schemas/case.yaml
+++ /dev/null
@@ -1,71 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "case"
-title: Case
-type: object
-namespace: http://gdc.nci.nih.gov
-category: administrative
-program: '*'
-project: '*'
-description: >
- The collection of all data related to a specific subject in the
- context of a specific experiment.
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - created_datetime
- - updated_datetime
- - state
-
-links:
- - name: experiments
- backref: cases
- label: member_of
- target_type: experiment
- multiplicity: many_to_one
- required: true
-
-required:
- - submitter_id
- - type
- - experiments
-
-uniqueKeys:
- - [id]
- - [project_id, submitter_id]
-
-# Case properties
-properties:
- type:
- type: string
- id:
- $ref: "_definitions.yaml#/UUID"
- systemAlias: node_id
- state:
- $ref: "_definitions.yaml#/state"
- submitter_id:
- type:
- - string
- - "null"
- consent_codes:
- type: array
- items:
- type: string
- primary_site:
- description: "Primary site for the case."
- type: string
- disease_type:
- description: "Name of the disease for the case."
- type: string
- experiments:
- $ref: "_definitions.yaml#/to_one"
- project_id:
- $ref: "_definitions.yaml#/project_id"
- created_datetime:
- $ref: "_definitions.yaml#/datetime"
- updated_datetime:
- $ref: "_definitions.yaml#/datetime"
diff --git a/datadictionary/gdcdictionary/schemas/clinical_test.yaml b/datadictionary/gdcdictionary/schemas/clinical_test.yaml
deleted file mode 100644
index 0ac0376f..00000000
--- a/datadictionary/gdcdictionary/schemas/clinical_test.yaml
+++ /dev/null
@@ -1,242 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "clinical_test"
-title: Clinical Test
-type: object
-namespace: http://gdc.nci.nih.gov
-category: clinical
-project: '*'
-program: '*'
-description: >
- Metadata concerning any clinical tests used in relation to a case diagnosis.
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - created_datetime
- - updated_datetime
- - state
-
-links:
- - name: cases
- backref: clinical_tests
- label: performed_for
- target_type: case
- multiplicity: many_to_one
- required: true
- - name: diagnoses
- backref: clinical_tests
- label: relates_to
- target_type: diagnosis
- multiplicity: many_to_many
- required: false
-
-required:
- - submitter_id
- - type
- - biomarker_name
- - biomarker_result
- - biomarker_test_method
- - cases
-
-uniqueKeys:
- - [id]
- - [project_id, submitter_id]
-
-properties:
- type:
- enum: [ "clinical_test" ]
-
- id:
- $ref: "_definitions.yaml#/UUID"
- systemAlias: node_id
-
- state:
- $ref: "_definitions.yaml#/state"
-
- submitter_id:
- type:
- - string
- - "null"
-
- biomarker_name:
- term:
- $ref: "_terms.yaml#/biomarker_name"
- type: string
-
- biomarker_result:
- term:
- $ref: "_terms.yaml#/biomarker_result"
- enum:
- - Amplification
- - Gain
- - Loss
- - Normal
- - Other
- - Translocation
- - Not Reported
- - Not Allowed To Collect
- - Pending
-
- biomarker_test_method:
- term:
- $ref: "_terms.yaml#/biomarker_test_method"
- enum:
- - Cytogenetics
- - FISH
- - IHC
- - Karyotype
- - NGS
- - Nuclear Staining
- - Other
- - RT-PCR
- - Southern
- - Not Reported
- - Not Allowed To Collect
- - Pending
-
- cea_level_preoperative:
- term:
- $ref: "_terms.yaml#/cea_level_preoperative"
- type: number
-
- dlco_ref_predictive_percent:
- term:
- $ref: "_terms.yaml#/dlco_ref_predictive_percent"
- type: number
-
- estrogen_receptor_percent_positive_ihc:
- term:
- $ref: "_terms.yaml#/estrogen_receptor_percent_positive_ihc"
- enum:
- - <1%
- - 1-10%
- - 11-20%
- - 21-30%
- - 31-40%
- - 41-50%
- - 51-60%
- - 61-70%
- - 71-80%
- - 81-90%
- - 91-100%
-
- estrogen_receptor_result_ihc:
- term:
- $ref: "_terms.yaml#/estrogen_receptor_result_ihc"
- enum:
- - Negative
- - Not Performed
- - Positive
- - Unknown
-
- fev1_ref_post_bronch_percent:
- term:
- $ref: "_terms.yaml#/fev1_ref_post_bronch_percent"
- type: number
-
- fev1_ref_pre_bronch_percent:
- term:
- $ref: "_terms.yaml#/fev1_ref_pre_bronch_percent"
- type: number
-
- fev1_fvc_post_bronch_percent:
- term:
- $ref: "_terms.yaml#/fev1_fvc_post_bronch_percent"
- type: number
-
- fev1_fvc_pre_bronch_percent:
- term:
- $ref: "_terms.yaml#/fev1_fvc_pre_bronch_percent"
- type: number
-
- her2_erbb2_percent_positive_ihc:
- term:
- $ref: "_terms.yaml#/her2_erbb2_percent_positive_ihc"
- enum:
- - <1%
- - 1-10%
- - 11-20%
- - 21-30%
- - 31-40%
- - 41-50%
- - 51-60%
- - 61-70%
- - 71-80%
- - 81-90%
- - 91-100%
-
- her2_erbb2_result_fish:
- term:
- $ref: "_terms.yaml#/her2_erbb2_result_fish"
- enum:
- - Negative
- - Not Performed
- - Positive
- - Unknown
-
- her2_erbb2_result_ihc:
- term:
- $ref: "_terms.yaml#/her2_erbb2_result_ihc"
- enum:
- - Negative
- - Not Performed
- - Positive
- - Unknown
-
- ldh_level_at_diagnosis:
- term:
- $ref: "_terms.yaml#/ldh_level_at_diagnosis"
- type: number
-
- ldh_normal_range_upper:
- term:
- $ref: "_terms.yaml#/ldh_normal_range_upper"
- type: number
-
- microsatellite_instability_abnormal:
- term:
- $ref: "_terms.yaml#/microsatellite_instability_abnormal"
- enum:
- - "Yes"
- - "No"
- - Unknown
-
- progesterone_receptor_percent_positive_ihc:
- term:
- $ref: "_terms.yaml#/progesterone_receptor_percent_positive_ihc"
- enum:
- - <1%
- - 1-10%
- - 11-20%
- - 21-30%
- - 31-40%
- - 41-50%
- - 51-60%
- - 61-70%
- - 71-80%
- - 81-90%
- - 91-100%
-
- progesterone_receptor_result_ihc:
- term:
- $ref: "_terms.yaml#/progesterone_receptor_result_ihc"
- enum:
- - Negative
- - Not Performed
- - Positive
- - Unknown
-
- cases:
- $ref: "_definitions.yaml#/to_one"
- diagnoses:
- $ref: "_definitions.yaml#/to_many"
- project_id:
- $ref: "_definitions.yaml#/project_id"
- created_datetime:
- $ref: "_definitions.yaml#/datetime"
- updated_datetime:
- $ref: "_definitions.yaml#/datetime"
diff --git a/datadictionary/gdcdictionary/schemas/condition.yaml b/datadictionary/gdcdictionary/schemas/condition.yaml
new file mode 100644
index 00000000..90c07afc
--- /dev/null
+++ b/datadictionary/gdcdictionary/schemas/condition.yaml
@@ -0,0 +1,468 @@
+$schema: http://json-schema.org/draft-04/schema#
+additionalProperties: false
+category: Clinical
+description: " Detailed information about conditions, problems or diagnoses.\n \
+ \ \n A clinical condition, problem, diagnosis, or other event, situation, issue,\n\
+ \ or clinical concept that has risen to a level of concern.\n http://hl7.org/fhir/condition"
+id: condition
+links:
+- backref: conditions
+ label: Condition_encounter_encounter
+ multiplicity: many_to_many
+ name: encounter_encounter
+ required: false
+ target_type: encounter
+- backref: conditions
+ label: Condition_subject_patient
+ multiplicity: many_to_many
+ name: subject_patient
+ required: false
+ target_type: patient
+namespace: http://hl7.org/fhir
+program: '*'
+project: '*'
+properties:
+ abatementDateTime:
+ description: When in resolution/remission.
+ type:
+ - string
+ - 'null'
+ abatementPeriod_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ abatementPeriod_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ abatementRange_high_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ abatementRange_high_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ abatementRange_high_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ abatementRange_high_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ abatementRange_high_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ abatementRange_low_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ abatementRange_low_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ abatementRange_low_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ abatementRange_low_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ abatementRange_low_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ abatementString:
+ description: When in resolution/remission.
+ type:
+ - string
+ - 'null'
+ bodySite_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ bodySite_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ bodySite_0_coding_0_system:
+ description: Anatomical location, if relevant.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ category_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ category_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ category_0_coding_0_system:
+ description: A category assigned to the condition.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ clinicalStatus_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ clinicalStatus_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ clinicalStatus_coding_0_system:
+ description: The clinical status of the condition.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ code_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ code_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ code_coding_0_system:
+ description: Identification of the condition, problem or diagnosis.. Code defined
+ by a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ created_datetime:
+ $ref: _definitions.yaml#/datetime
+ evidence_0_code_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ evidence_0_code_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ evidence_0_code_0_coding_0_system:
+ description: Manifestation/symptom.. Code defined by a terminology system.. Identity
+ of the terminology system.
+ type:
+ - string
+ - 'null'
+ evidence_0_detail_0_reference:
+ description: Supporting information found elsewhere.. Literal reference, Relative,
+ internal or absolute URL.
+ type:
+ - string
+ - 'null'
+ evidence_0_detail_0_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ id:
+ description: Logical id of this artifact.
+ type:
+ - string
+ - 'null'
+ identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ identifier_0_use:
+ description: External Ids for this condition.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: External Ids for this condition.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ language:
+ description: Language of the resource content.
+ type:
+ - string
+ - 'null'
+ onsetDateTime:
+ description: Estimated or actual date, date-time, or age.
+ type:
+ - string
+ - 'null'
+ onsetPeriod_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ onsetPeriod_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ onsetRange_high_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ onsetRange_high_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ onsetRange_high_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ onsetRange_high_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ onsetRange_high_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ onsetRange_low_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ onsetRange_low_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ onsetRange_low_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ onsetRange_low_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ onsetRange_low_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ onsetString:
+ description: Estimated or actual date, date-time, or age.
+ type:
+ - string
+ - 'null'
+ project_id:
+ $ref: _definitions.yaml#/project_id
+ recordedDate:
+ description: Date record was first recorded.
+ type:
+ - string
+ - 'null'
+ resource_type:
+ description: ''
+ type:
+ - string
+ - 'null'
+ severity_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ severity_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ severity_coding_0_system:
+ description: Subjective severity of condition.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ stage_0_assessment_0_reference:
+ description: Formal record of assessment.. Literal reference, Relative, internal
+ or absolute URL.
+ type:
+ - string
+ - 'null'
+ stage_0_assessment_0_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ stage_0_summary_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ stage_0_summary_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ stage_0_summary_coding_0_system:
+ description: Simple summary (disease specific).. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ stage_0_type_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ stage_0_type_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ stage_0_type_coding_0_system:
+ description: Kind of staging.. Code defined by a terminology system.. Identity
+ of the terminology system.
+ type:
+ - string
+ - 'null'
+ state:
+ $ref: _definitions.yaml#/state
+ submitter_id:
+ type:
+ - string
+ - 'null'
+ updated_datetime:
+ $ref: _definitions.yaml#/datetime
+ verificationStatus_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ verificationStatus_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ verificationStatus_coding_0_system:
+ description: The verification status to support the clinical status of the condition..
+ Code defined by a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+required:
+- submitter_id
+- subject
+submittable: true
+systemProperties:
+- id
+- project_id
+- created_datetime
+- updated_datetime
+- state
+title: Condition
+type: object
+uniqueKeys:
+- - id
+- - project_id
+ - submitter_id
+validators: null
diff --git a/datadictionary/gdcdictionary/schemas/core_metadata_collection.yaml b/datadictionary/gdcdictionary/schemas/core_metadata_collection.yaml
deleted file mode 100644
index 1a2bcebc..00000000
--- a/datadictionary/gdcdictionary/schemas/core_metadata_collection.yaml
+++ /dev/null
@@ -1,113 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "core_metadata_collection"
-title: Core Metadata Collection
-type: object
-namespace: https://dcp.bionimbus.org/
-category: administrative
-program: '*'
-project: '*'
-description: >
- Structured description of a collection of several dataset
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - state
- - created_datetime
- - updated_datetime
-
-links:
- - name: projects
- backref: core_metadata_collections
- label: data_from
- target_type: project
- multiplicity: many_to_one
- required: true
-
-uniqueKeys:
- - [id]
- - [project_id, submitter_id]
-
-required:
- - submitter_id
- - type
- - projects
-
-properties:
- $ref: "_definitions.yaml#/ubiquitous_properties"
-
- contributor:
- description: >
- An entity responsible for making contributions to the resource. Examples of a Contributor include a person, an organization, or a service. Typically, the name of a Contributor should be used to indicate the entity.
- type: string
-
- coverage:
- description: >
- The spatial or temporal topic of the resource, the spatial applicability of the resource, or the jurisdiction under which the resource is relevant. Spatial topic and spatial applicability may be a named place or a location specified by its geographic coordinates. Temporal topic may be a named period, date, or date range. A jurisdiction may be a named administrative entity or a geographic place to which the resource applies. Recommended best practice is to use a controlled vocabulary such as the Thesaurus of Geographic Names [TGN] (http://www.getty.edu/research/tools/vocabulary/tgn/index.html). Where appropriate, named places or time periods can be used in preference to numeric identifiers such as sets of coordinates or date ranges.
- type: string
-
- creator:
- description: >
- An entity primarily responsible for making the resource. Examples of a Creator include a person, an organization, or a service. Typically, the name of a Creator should be used to indicate the entity.
- type: string
-
- date:
- $ref: "_definitions.yaml#/datetime"
-
- description:
- description: >
- An account of the resource. Description may include but is not limited to: an abstract, a table of contents, a graphical representation, or a free-text account of the resource.
- type: string
-
- format:
- description: >
- The file format, physical medium, or dimensions of the resource. Examples of dimensions include size and duration. Recommended best practice is to use a controlled vocabulary such as the list of Internet Media Types [MIME] (http://www.iana.org/assignments/media-types/).
- type: string
-
- language:
- description: >
- A language of the resource. Recommended best practice is to use a controlled vocabulary such as RFC 4646 (http://www.ietf.org/rfc/rfc4646.txt).
- type: string
-
- publisher:
- description: >
- An entity responsible for making the resource available. Examples of a Publisher include a person, an organization, or a service. Typically, the name of a Publisher should be used to indicate the entity.
- type: string
-
- relation:
- description: >
- A related resource. Recommended best practice is to identify the related resource by means of a string conforming to a formal identification system.
- type: string
-
- rights:
- description: >
- Information about rights held in and over the resource. Typically, rights information includes a statement about various property rights associated with the resource, including intellectual property rights.
- type: string
-
- source:
- description: >
- A related resource from which the described resource is derived. The described resource may be derived from the related resource in whole or in part. Recommended best practice is to identify the related resource by means of a string conforming to a formal identification system.
- type: string
-
- subject:
- description: >
- The topic of the resource. Typically, the subject will be represented using keywords, key phrases, or classification codes. Recommended best practice is to use a controlled vocabulary.
- type: string
-
- title:
- description: >
- A name given to the resource. Typically, a Title will be a name by which the resource is formally known.
- type: string
-
- data_type:
- description: >
- The nature or genre of the resource. Recommended best practice is to use a controlled vocabulary such as the DCMI Type Vocabulary [DCMITYPE]. To describe the file format, physical medium, or dimensions of the resource, use the Format element.
- type: string
-
- projects:
- $ref: "_definitions.yaml#/to_one_project"
-
diff --git a/datadictionary/gdcdictionary/schemas/demographic.yaml b/datadictionary/gdcdictionary/schemas/demographic.yaml
deleted file mode 100644
index ea8a2b42..00000000
--- a/datadictionary/gdcdictionary/schemas/demographic.yaml
+++ /dev/null
@@ -1,114 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "demographic"
-title: Demographic
-type: object
-namespace: http://gdc.nci.nih.gov
-category: clinical
-program: '*'
-project: '*'
-description: >
- Data for the characterization of the patient by means of segementing the population (e.g.,
- characterization by age, sex, or race).
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - state
- - created_datetime
- - updated_datetime
-
-links:
- - name: cases
- backref: demographics
- label: describes
- target_type: case
- multiplicity: one_to_one
- required: true
-
-required:
- - submitter_id
- - type
- - cases
-
-preferred:
- - year_of_death
-
-uniqueKeys:
- #unclear if want submitter ID for clinical
- - [id]
- - [project_id, submitter_id]
-
-properties:
- type:
- type: string
-
- id:
- $ref: "_definitions.yaml#/UUID"
- systemAlias: node_id
-
- state:
- $ref: "_definitions.yaml#/state"
-
- submitter_id:
- type:
- - string
- - "null"
-
- gender:
- term:
- $ref: "_terms.yaml#/gender"
- enum:
- - female
- - male
- - unknown
- - unspecified
- - not reported
-
- race:
- term:
- $ref: "_terms.yaml#/race"
- enum:
- - white
- - american indian or alaska native
- - black or african american
- - asian
- - native hawaiian or other pacific islander
- - other
- - Unknown
- - not reported
- - not allowed to collect
-
- ethnicity:
- term:
- $ref: "_terms.yaml#/ethnicity"
- enum:
- - hispanic or latino
- - not hispanic or latino
- - Unknown
- - not reported
- - not allowed to collect
-
- year_of_birth:
- term:
- $ref: "_terms.yaml#/year_of_birth"
- type:
- - number
- - "null"
-
- year_of_death:
- term:
- $ref: "_terms.yaml#/year_of_death"
- type: number
-
- cases:
- $ref: "_definitions.yaml#/to_one"
- project_id:
- $ref: "_definitions.yaml#/project_id"
- created_datetime:
- $ref: "_definitions.yaml#/datetime"
- updated_datetime:
- $ref: "_definitions.yaml#/datetime"
diff --git a/datadictionary/gdcdictionary/schemas/diagnosis.yaml b/datadictionary/gdcdictionary/schemas/diagnosis.yaml
deleted file mode 100644
index 28c42408..00000000
--- a/datadictionary/gdcdictionary/schemas/diagnosis.yaml
+++ /dev/null
@@ -1,844 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "diagnosis"
-title: Diagnosis
-type: object
-namespace: http://gdc.nci.nih.gov
-category: clinical
-program: '*'
-project: '*'
-description: >
- Data from the investigation, analysis and recognition of the presence and nature of disease,
- condition, or injury from expressed signs and symptoms; also, the scientific determination of any
- kind; the concise results of such an investigation.
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - state
- - created_datetime
- - updated_datetime
-
-links:
- - name: cases
- backref: diagnoses
- label: describes
- target_type: case
- multiplicity: many_to_one
- required: true
-
-required:
- - submitter_id
- - type
- - age_at_diagnosis
- - days_to_last_follow_up
- - vital_status
- - primary_diagnosis
- - morphology
- - tissue_or_organ_of_origin
- - site_of_resection_or_biopsy
- - classification_of_tumor
- - tumor_stage
- - tumor_grade
- - progression_or_recurrence
- - days_to_recurrence
- - days_to_last_known_disease_status
- - last_known_disease_status
-
-preferred:
- - days_to_birth
- - site_of_resection_or_biopsy
-
-uniqueKeys:
- #unclear if want submitter ID for clinical
- - [id]
- - [project_id, submitter_id]
-
-properties:
- type:
- type: string
-
- id:
- $ref: "_definitions.yaml#/UUID"
- systemAlias: node_id
-
- state:
- $ref: "_definitions.yaml#/state"
-
- submitter_id:
- type:
- - string
- - "null"
-
- age_at_diagnosis:
- term:
- $ref: "_terms.yaml#/age_at_diagnosis"
- type:
- - number
- - "null"
- maximum: 32872
- minimum: 0
-
- ajcc_clinical_m:
- term:
- $ref: "_terms.yaml#/ajcc_clinical_m"
- enum:
- - M0
- - M1
- - M1a
- - M1b
- - M1c
- - MX
- - cM0 (i+)
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- ajcc_clinical_n:
- term:
- $ref: "_terms.yaml#/ajcc_clinical_n"
- enum:
- - N0
- - N0 (i+)
- - N0 (i-)
- - N0 (mol+)
- - N0 (mol-)
- - N1
- - N1a
- - N1b
- - N1bI
- - N1bII
- - N1bIII
- - N1bIV
- - N1c
- - N1mi
- - N2
- - N2a
- - N2b
- - N2c
- - N3
- - N3a
- - N3b
- - N3c
- - N4
- - NX
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- ajcc_clinical_stage:
- term:
- $ref: "_terms.yaml#/ajcc_clinical_stage"
- enum:
- - Stage 0
- - Stage 0a
- - Stage 0is
- - Stage I
- - Stage IA
- - Stage IA1
- - Stage IA2
- - Stage IB
- - Stage IB Cervix
- - Stage IB1
- - Stage IB2
- - Stage II
- - Stage II Cervix
- - Stage IIA
- - Stage IIA Cervix
- - Stage IIB
- - Stage IIC
- - Stage III
- - Stage IIIA
- - Stage IIIB
- - Stage IIIC
- - Stage IS
- - Stage IV
- - Stage IVA
- - Stage IVB
- - Stage IVC
- - Stage Tis
- - Stage X
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- ajcc_clinical_t:
- term:
- $ref: "_terms.yaml#/ajcc_clinical_t"
- enum:
- - T0
- - T1
- - T1a
- - T1a1
- - T1a2
- - T1b
- - T1b1
- - T1b2
- - T1c
- - T1mi
- - T2
- - T2a
- - T2a1
- - T2a2
- - T2b
- - T2c
- - T2d
- - T3
- - T3a
- - T3b
- - T3c
- - T3d
- - T4
- - T4a
- - T4b
- - T4c
- - T4d
- - T4e
- - TX
- - Ta
- - Tis
- - Tis (DCIS)
- - Tis (LCIS)
- - Tis (Paget's)
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- ajcc_pathologic_m:
- term:
- $ref: "_terms.yaml#/ajcc_pathologic_m"
- enum:
- - M0
- - M1
- - M1a
- - M1b
- - M1c
- - M2
- - MX
- - cM0 (i+)
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- ajcc_pathologic_n:
- term:
- $ref: "_terms.yaml#/ajcc_pathologic_n"
- enum:
- - N0
- - N0 (i+)
- - N0 (i-)
- - N0 (mol+)
- - N0 (mol-)
- - N1
- - N1a
- - N1b
- - N1bI
- - N1bII
- - N1bIII
- - N1bIV
- - N1c
- - N1mi
- - N2
- - N2a
- - N2b
- - N2c
- - N3
- - N3a
- - N3b
- - N3c
- - N4
- - NX
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- ajcc_pathologic_stage:
- term:
- $ref: "_terms.yaml#/ajcc_pathologic_stage"
- enum:
- - Stage 0
- - Stage 0a
- - Stage 0is
- - Stage I
- - Stage IA
- - Stage IA1
- - Stage IA2
- - Stage IB
- - Stage IB1
- - Stage IB2
- - Stage IC
- - Stage II
- - Stage IIA
- - Stage IIA1
- - Stage IIA2
- - Stage IIB
- - Stage IIC
- - Stage III
- - Stage IIIA
- - Stage IIIB
- - Stage IIIC
- - Stage IV
- - Stage IVA
- - Stage IVB
- - Stage IVC
- - Stage Tis
- - Stage X
-
- ajcc_pathologic_t:
- term:
- $ref: "_terms.yaml#/ajcc_pathologic_t"
- enum:
- - T0
- - T1
- - T1a
- - T1a1
- - T1a2
- - T1b
- - T1b1
- - T1b2
- - T1c
- - T1mi
- - T2
- - T2a
- - T2a1
- - T2a2
- - T2b
- - T2c
- - T2d
- - T3
- - T3a
- - T3b
- - T3c
- - T3d
- - T4
- - T4a
- - T4b
- - T4c
- - T4d
- - T4e
- - TX
- - Ta
- - Tis
- - Tis (DCIS)
- - Tis (LCIS)
- - Tis (Paget's)
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- ann_arbor_b_symptoms:
- term:
- $ref: "_terms.yaml#/ann_arbor_b_symptoms"
- enum:
- - "Yes"
- - "No"
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- ann_arbor_clinical_stage:
- term:
- $ref: "_terms.yaml#/ann_arbor_clinical_stage"
- enum:
- - Stage I
- - Stage II
- - Stage III
- - Stage IV
-
- ann_arbor_extranodal_involvement:
- term:
- $ref: "_terms.yaml#/ann_arbor_extranodal_involvement"
- enum:
- - "Yes"
- - "No"
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- ann_arbor_pathologic_stage:
- term:
- $ref: "_terms.yaml#/ann_arbor_pathologic_stage"
- enum:
- - Stage I
- - Stage II
- - Stage III
- - Stage IV
-
- burkitt_lymphoma_clinical_variant:
- term:
- $ref: "_terms.yaml#/burkitt_lymphoma_clinical_variant"
- enum:
- - "Endemic"
- - "Immunodeficiency-associated, adult"
- - "Immunodeficiency-associated, pediatric"
- - "Sporadic, adult"
- - "Sporadic, pediatric"
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- cause_of_death:
- term:
- $ref: "_terms.yaml#/cause_of_death"
- enum:
- - Cancer Related
- - Not Cancer Related
- - Unknown
-
- circumferential_resection_margin:
- term:
- $ref: "_terms.yaml#/circumferential_resection_margin"
- type: number
-
- classification_of_tumor:
- term:
- $ref: "_terms.yaml#/classification_of_tumor"
- enum:
- - primary
- - metastasis
- - recurrence
- - other
- - Unknown
- - not reported
- - Not Allowed To Collect
-
- colon_polyps_history:
- term:
- $ref: "_terms.yaml#/colon_polyps_history"
- enum:
- - "Yes"
- - "No"
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- days_to_birth:
- term:
- $ref: "_terms.yaml#/days_to_birth"
- type:
- - number
- - "null"
- maximum: 0
- minimum: -32872
-
- days_to_death:
- term:
- $ref: "_terms.yaml#/days_to_death"
- type: number
- maximum: 32872
- minimum: 0
-
- days_to_hiv_diagnosis:
- term:
- $ref: "_terms.yaml#/days_to_hiv_diagnosis"
- type:
- - number
- - "null"
-
- days_to_last_follow_up:
- term:
- $ref: "_terms.yaml#/days_to_last_follow_up"
- type:
- - number
- - "null"
-
- days_to_last_known_disease_status:
- term:
- $ref: "_terms.yaml#/days_to_last_known_disease_status"
- type:
- - number
- - "null"
-
- days_to_new_event:
- term:
- $ref: "_terms.yaml#/days_to_new_event"
- type:
- - number
- - "null"
-
- days_to_recurrence:
- term:
- $ref: "_terms.yaml#/days_to_recurrence"
- type:
- - number
- - "null"
-
- figo_stage:
- term:
- $ref: "_terms.yaml#/figo_stage"
- enum:
- - Stage 0
- - Stage I
- - Stage IA
- - Stage IA1
- - Stage IA2
- - Stage IB
- - Stage IB1
- - Stage IB2
- - Stage IC
- - Stage II
- - Stage IIA
- - Stage IIA1
- - Stage IIA2
- - Stage IIB
- - Stage III
- - Stage IIIA
- - Stage IIIB
- - Stage IIIC
- - Stage IIIC1
- - Stage IIIC2
- - Stage IV
- - Stage IVA
- - Stage IVB
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- hiv_positive:
- term:
- $ref: "_terms.yaml#/hiv_positive"
- enum:
- - "Yes"
- - "No"
- - Unknown
-
- hpv_positive_type:
- term:
- $ref: "_terms.yaml#/hpv_positive_type"
- enum:
- - HPV 16
- - HPV 18
- - Other HPV type(s)
- - Unknown
-
- hpv_status:
- term:
- $ref: "_terms.yaml#/hpv_status"
- enum:
- - Negative
- - Positive
- - Unknown
-
- last_known_disease_status:
- term:
- $ref: "_terms.yaml#/last_known_disease_status"
- enum:
- - Distant met recurrence/progression
- - Loco-regional recurrence/progression
- - Biochemical evidence of disease without structural correlate
- - Tumor free
- - Unknown tumor status
- - With tumor
- - not reported
- - Not Allowed To Collect
-
- laterality:
- term:
- $ref: "_terms.yaml#/laterality"
- enum:
- - Bilateral
- - Left
- - Right
- - Unknown
-
- ldh_level_at_diagnosis:
- term:
- $ref: "_terms.yaml#/ldh_level_at_diagnosis"
- type:
- - number
- - "null"
-
- ldh_normal_range_upper:
- term:
- $ref: "_terms.yaml#/ldh_normal_range_upper"
- type:
- - number
- - "null"
-
- lymph_nodes_positive:
- term:
- $ref: "_terms.yaml#/lymph_nodes_positive"
- type: integer
-
- lymphatic_invasion_present:
- term:
- $ref: "_terms.yaml#/lymphatic_invasion_present"
- enum:
- - "Yes"
- - "No"
- - Unknown
-
- method_of_diagnosis:
- term:
- $ref: "_terms.yaml#/method_of_diagnosis"
- enum:
- - Autopsy
- - Biopsy
- - Blood Draw
- - Bone Marrow Aspirate
- - Core Biopsy
- - Cytology
- - Debulking
- - Diagnostic Imaging
- - Excisional Biopsy
- - Fine Needle Aspiration
- - Incisional Biopsy
- - Laparoscopy
- - Laparotomy
- - Other
- - Surgical Resection
- - Ultrasound Guided Biopsy
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- morphology:
- term:
- $ref: "_terms.yaml#/morphology"
- type: string
-
- new_event_anatomic_site:
- term:
- $ref: "_terms.yaml#/new_event_anatomic_site"
- enum:
- - Abdomen
- - Adrenal
- - Anus
- - Appendix
- - Ascites/Peritoneum
- - Axillary lymph nodes
- - Bladder
- - Bone
- - Bone Marrow
- - Brain
- - Breast
- - Cervical lymph nodes
- - Cervix
- - Colon
- - Conjunctiva
- - Contralateral Pleura
- - Distant Metastasis
- - Epididymis
- - Epidural
- - Epitrochlear lymph nodes
- - Esophagus
- - Extremities
- - Femoral lymph nodes
- - Gallbladder
- - Gastrointestinal/Abdominal
- - Head & Neck
- - Heart
- - Hilar lymph nodes
- - Hypopharynx
- - Iliac Lymph Node
- - Iliac-common lymph nodes
- - Iliac-external lymph nodes
- - Inguinal lymph nodes
- - Intraocular
- - Ipsilateral Chest Cavity
- - Ipsilateral Chest Wall
- - Ipsilateral Lymph Nodes
- - Ipsilateral Pleura
- - Kidney
- - Large Intestine
- - Larynx
- - Leptomeninges
- - Liver
- - Lung
- - Lymph Node Only
- - Lymph Node(s)
- - Mandible
- - Maxilla
- - Mediastinal Soft Tissue
- - Mediastinal lymph nodes
- - Mediastinal/Intra-thoracic
- - Mesenteric lymph nodes
- - Nasal Soft Tissue
- - Nasopharynx
- - No Known Extranodal Involvement
- - Non-regional / Distant Lymph Nodes
- - Not Applicable
- - Occipital lymph nodes
- - Oral Cavity
- - Oropharynx
- - Other
- - Other Extranodal Site
- - Other, specify
- - Ovary
- - Pancreas
- - Paraaortic lymph nodes
- - Parotid Gland
- - Parotid lymph nodes
- - Pelvis
- - Peri-orbital Soft Tissue
- - Pericardium
- - Perihilar Lymph Node
- - Peripheral Blood
- - Peritoneal Surfaces
- - Pleura/Pleural Effusion
- - Popliteal lymph nodes
- - Prostate
- - Pulmonary
- - Rectum
- - Renal Pelvis
- - Retroperitoneal lymph nodes
- - Retroperitoneum
- - Salivary Gland
- - Sinus
- - Skin
- - Small Intestine
- - Soft Tissue
- - Splenic lymph nodes
- - Stomach
- - Submandibular lymph nodes
- - Supraclavicular lymph nodes
- - Testes
- - Thyroid
- - Trunk
- - Tumor Bed
- - Ureter
- - Urethra
- - Uterus
- - Vulva
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- new_event_type:
- term:
- $ref: "_terms.yaml#/new_event_type"
- enum:
- - Biochemical Evidence of Disease
- - Both Locoregional and Distant Metastasis
- - Distant Metastasis
- - Extrahepatic Recurrence
- - Intrahepatic Recurrence
- - Intrapleural Progression
- - Locoregional (Urothelial tumor event)
- - Locoregional Disease
- - Locoregional Recurrence
- - Metachronous Testicular Tumor
- - Metastatic
- - New Primary Tumor
- - New primary Melanoma
- - No New Tumor Event
- - Not Applicable
- - Progression of Disease
- - Recurrence
- - Regional Lymph Node
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- perineural_invasion_present:
- term:
- $ref: "_terms.yaml#/perineural_invasion_present"
- enum:
- - "Yes"
- - "No"
- - Unknown
-
- primary_diagnosis:
- term:
- $ref: "_terms.yaml#/primary_diagnosis"
- type: string
-
- prior_malignancy:
- term:
- $ref: "_terms.yaml#/prior_malignancy"
- enum:
- - "yes"
- - "no"
- - unknown
- - not reported
- - Not Allowed To Collect
-
- prior_treatment:
- term:
- $ref: "_terms.yaml#/prior_treatment"
- enum:
- - "Yes"
- - "No"
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- progression_or_recurrence:
- term:
- $ref: "_terms.yaml#/progression_or_recurrence"
- enum:
- - "yes"
- - "no"
- - unknown
- - not reported
- - Not Allowed To Collect
-
- residual_disease:
- term:
- $ref: "_terms.yaml#/residual_disease"
- enum:
- - R0
- - R1
- - R2
- - RX
-
- site_of_resection_or_biopsy:
- term:
- $ref: "_terms.yaml#/site_of_resection_or_biopsy"
- type: string
-
- tissue_or_organ_of_origin:
- term:
- $ref: "_terms.yaml#/tissue_or_organ_of_origin"
- type: string
-
- tumor_grade:
- term:
- $ref: "_terms.yaml#/tumor_grade"
- type: string
-
- tumor_stage:
- term:
- $ref: "_terms.yaml#/tumor_stage"
- type: string
-
- vascular_invasion_present:
- term:
- $ref: "_terms.yaml#/vascular_invasion_present"
- enum:
- - "Yes"
- - "No"
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- vital_status:
- term:
- $ref: "_terms.yaml#/vital_status"
- enum:
- - alive
- - dead
- - lost to follow-up
- - unknown
- - not reported
- - Not Allowed To Collect
- - pending # this is from TARGET RT clinical data
-
- year_of_diagnosis:
- term:
- $ref: "_terms.yaml#/year_of_diagnosis"
- type:
- - number
- - "null"
-
- cases:
- $ref: "_definitions.yaml#/to_one"
- project_id:
- $ref: "_definitions.yaml#/project_id"
- created_datetime:
- $ref: "_definitions.yaml#/datetime"
- updated_datetime:
- $ref: "_definitions.yaml#/datetime"
diff --git a/datadictionary/gdcdictionary/schemas/diagnostic_report.yaml b/datadictionary/gdcdictionary/schemas/diagnostic_report.yaml
new file mode 100644
index 00000000..5e1703f3
--- /dev/null
+++ b/datadictionary/gdcdictionary/schemas/diagnostic_report.yaml
@@ -0,0 +1,289 @@
+$schema: http://json-schema.org/draft-04/schema#
+additionalProperties: false
+category: Clinical
+description: " A Diagnostic report - a combination of request information, atomic\
+ \ results,\n images, interpretation, as well as formatted reports.\n \n \
+ \ The findings and interpretation of diagnostic tests performed on patients,\n\
+ \ groups of patients, devices, and locations, and/or specimens derived from\n\
+ \ these. The report includes clinical context such as requesting and provider\n\
+ \ information, and some mix of atomic results, images, textual and coded\n \
+ \ interpretations, and formatted representation of diagnostic reports.\n http://hl7.org/fhir/diagnosticreport"
+id: diagnostic_report
+links:
+- backref: diagnostic_reports
+ label: DiagnosticReport_encounter_encounter
+ multiplicity: many_to_many
+ name: encounter_encounter
+ required: false
+ target_type: encounter
+- backref: diagnostic_reports
+ label: DiagnosticReport_imagingStudy_imaging_study
+ multiplicity: many_to_many
+ name: imagingStudy_imaging_study
+ required: false
+ target_type: imaging_study
+- backref: diagnostic_reports
+ label: DiagnosticReport_performer_practitioner
+ multiplicity: many_to_many
+ name: performer_practitioner
+ required: false
+ target_type: practitioner
+- backref: diagnostic_reports
+ label: DiagnosticReport_performer_organization
+ multiplicity: many_to_many
+ name: performer_organization
+ required: false
+ target_type: organization
+- backref: diagnostic_reports
+ label: DiagnosticReport_performer_practitioner_role
+ multiplicity: many_to_many
+ name: performer_practitioner_role
+ required: false
+ target_type: practitioner_role
+- backref: diagnostic_reports
+ label: DiagnosticReport_result_observation
+ multiplicity: many_to_many
+ name: result_observation
+ required: false
+ target_type: observation
+- backref: diagnostic_reports
+ label: DiagnosticReport_specimen_specimen
+ multiplicity: many_to_many
+ name: specimen_specimen
+ required: false
+ target_type: specimen
+- backref: diagnostic_reports
+ label: DiagnosticReport_subject_patient
+ multiplicity: many_to_many
+ name: subject_patient
+ required: false
+ target_type: patient
+namespace: http://hl7.org/fhir
+program: '*'
+project: '*'
+properties:
+ category_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ category_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ category_0_coding_0_system:
+ description: Service category.. Code defined by a terminology system.. Identity
+ of the terminology system.
+ type:
+ - string
+ - 'null'
+ code_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ code_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ code_coding_0_system:
+ description: Name/Code for this diagnostic report.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ conclusion:
+ description: Clinical conclusion (interpretation) of test results.
+ type:
+ - string
+ - 'null'
+ conclusionCode_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ conclusionCode_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ conclusionCode_0_coding_0_system:
+ description: Codes for the clinical conclusion of test results.. Code defined
+ by a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ created_datetime:
+ $ref: _definitions.yaml#/datetime
+ effectiveDateTime:
+ description: Clinically relevant time/time-period for report.
+ type:
+ - string
+ - 'null'
+ effectivePeriod_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ effectivePeriod_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ id:
+ description: Logical id of this artifact.
+ type:
+ - string
+ - 'null'
+ identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ identifier_0_use:
+ description: Business identifier for report.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: Business identifier for report.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ issued:
+ description: DateTime this version was made.
+ type:
+ - string
+ - 'null'
+ language:
+ description: Language of the resource content.
+ type:
+ - string
+ - 'null'
+ media_0_comment:
+ description: Comment about the image (e.g. explanation).
+ type:
+ - string
+ - 'null'
+ media_0_link_reference:
+ description: Reference to the image source.. Literal reference, Relative, internal
+ or absolute URL.
+ type:
+ - string
+ - 'null'
+ media_0_link_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ presentedForm_0_contentType:
+ description: Mime type of the content, with charset etc..
+ type:
+ - string
+ - 'null'
+ presentedForm_0_creation:
+ description: Date attachment was first created.
+ type:
+ - string
+ - 'null'
+ presentedForm_0_data:
+ description: Data inline, base64ed.
+ type:
+ - string
+ - 'null'
+ presentedForm_0_hash:
+ description: Hash of the data (sha-1, base64ed).
+ type:
+ - string
+ - 'null'
+ presentedForm_0_language:
+ description: Human language of the content (BCP-47).
+ type:
+ - string
+ - 'null'
+ presentedForm_0_size:
+ description: Number of bytes of content (if url provided).
+ type:
+ - number
+ - 'null'
+ presentedForm_0_title:
+ description: Label to display in place of the data.
+ type:
+ - string
+ - 'null'
+ presentedForm_0_url:
+ description: Uri where the data can be found.
+ type:
+ - string
+ - 'null'
+ project_id:
+ $ref: _definitions.yaml#/project_id
+ resource_type:
+ description: ''
+ type:
+ - string
+ - 'null'
+ state:
+ $ref: _definitions.yaml#/state
+ status:
+ description: The status of the diagnostic report.. http://hl7.org/fhir/diagnostic-report-status
+ enum:
+ - registered
+ - partial
+ - preliminary
+ - final
+ - amended
+ - corrected
+ - appended
+ - cancelled
+ - entered-in-error
+ - unknown
+ term:
+ description: The status of the diagnostic report.. http://hl7.org/fhir/diagnostic-report-status
+ termDef:
+ cde_id: http://hl7.org/fhir/diagnostic-report-status
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/diagnostic-report-status
+ term_url: http://hl7.org/fhir/diagnostic-report-status
+ submitter_id:
+ type:
+ - string
+ - 'null'
+ updated_datetime:
+ $ref: _definitions.yaml#/datetime
+required:
+- submitter_id
+- status
+- code
+submittable: true
+systemProperties:
+- id
+- project_id
+- created_datetime
+- updated_datetime
+- state
+title: DiagnosticReport
+type: object
+uniqueKeys:
+- - id
+- - project_id
+ - submitter_id
+validators: null
diff --git a/datadictionary/gdcdictionary/schemas/document_reference.yaml b/datadictionary/gdcdictionary/schemas/document_reference.yaml
new file mode 100644
index 00000000..cb8c8e6c
--- /dev/null
+++ b/datadictionary/gdcdictionary/schemas/document_reference.yaml
@@ -0,0 +1,374 @@
+$schema: http://json-schema.org/draft-04/schema#
+additionalProperties: false
+category: data_file
+description: " A reference to a document.\n \n A reference to a document of\
+ \ any kind for any purpose. Provides metadata\n about the document so that the\
+ \ document can be discovered and managed. The\n scope of a document is any seralized\
+ \ object with a mime-type, so includes\n formal patient centric documents (CDA),\
+ \ cliical notes, scanned paper, and\n non-patient specific documents like policy\
+ \ text.\n http://hl7.org/fhir/documentreference"
+id: document_reference
+links:
+- backref: document_references
+ label: DocumentReference_subject_patient
+ multiplicity: many_to_many
+ name: subject_patient
+ required: false
+ target_type: patient
+namespace: http://hl7.org/fhir
+program: '*'
+project: '*'
+properties:
+ $ref: _definitions.yaml#/data_file_properties
+ category_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ category_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ category_0_coding_0_system:
+ description: Categorization of document.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ content_0_attachment_contentType:
+ description: Mime type of the content, with charset etc..
+ type:
+ - string
+ - 'null'
+ content_0_attachment_creation:
+ description: Date attachment was first created.
+ type:
+ - string
+ - 'null'
+ content_0_attachment_data:
+ description: Data inline, base64ed.
+ type:
+ - string
+ - 'null'
+ content_0_attachment_hash:
+ description: Hash of the data (sha-1, base64ed).
+ type:
+ - string
+ - 'null'
+ content_0_attachment_language:
+ description: Human language of the content (BCP-47).
+ type:
+ - string
+ - 'null'
+ content_0_attachment_size:
+ description: Number of bytes of content (if url provided).
+ type:
+ - number
+ - 'null'
+ content_0_attachment_title:
+ description: Label to display in place of the data.
+ type:
+ - string
+ - 'null'
+ content_0_attachment_url:
+ description: Uri where the data can be found.
+ type:
+ - string
+ - 'null'
+ content_0_format_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ content_0_format_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ content_0_format_system:
+ description: Format/content rules for the document.. Identity of the terminology
+ system.
+ type:
+ - string
+ - 'null'
+ context_encounter_0_reference:
+ description: Context of the document content.. Literal reference, Relative, internal
+ or absolute URL.
+ type:
+ - string
+ - 'null'
+ context_encounter_0_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ context_event_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ context_event_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ context_event_0_coding_0_system:
+ description: Main clinical acts documented.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ context_facilityType_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ context_facilityType_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ context_facilityType_coding_0_system:
+ description: Kind of facility where patient was seen.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ context_period_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ context_period_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ context_practiceSetting_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ context_practiceSetting_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ context_practiceSetting_coding_0_system:
+ description: Additional details about where the content was created (e.g. clinical
+ specialty).. Code defined by a terminology system.. Identity of the terminology
+ system.
+ type:
+ - string
+ - 'null'
+ context_related_0_reference:
+ description: Related identifiers or resources.. Literal reference, Relative, internal
+ or absolute URL.
+ type:
+ - string
+ - 'null'
+ context_related_0_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ context_sourcePatientInfo_reference:
+ description: Patient demographics from source.. Literal reference, Relative, internal
+ or absolute URL.
+ type:
+ - string
+ - 'null'
+ context_sourcePatientInfo_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ created_datetime:
+ $ref: _definitions.yaml#/datetime
+ data_category:
+ term:
+ $ref: _terms.yaml#/data_category
+ type: string
+ data_format:
+ term:
+ $ref: _terms.yaml#/data_format
+ type: string
+ data_type:
+ term:
+ $ref: _terms.yaml#/data_type
+ type: string
+ date:
+ description: When this document reference was created.
+ type:
+ - string
+ - 'null'
+ description:
+ description: Human-readable description.
+ type:
+ - string
+ - 'null'
+ docStatus:
+ description: The status of the underlying document.. http://hl7.org/fhir/composition-status
+ enum:
+ - preliminary
+ - final
+ - amended
+ - entered-in-error
+ term:
+ description: The status of the underlying document.. http://hl7.org/fhir/composition-status
+ termDef:
+ cde_id: http://hl7.org/fhir/composition-status
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/composition-status
+ term_url: http://hl7.org/fhir/composition-status
+ id:
+ description: Logical id of this artifact.
+ type:
+ - string
+ - 'null'
+ identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ identifier_0_use:
+ description: Other identifiers for the document.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: Other identifiers for the document.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ language:
+ description: Language of the resource content.
+ type:
+ - string
+ - 'null'
+ masterIdentifier_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ masterIdentifier_use:
+ description: Master Version Specific Identifier.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: Master Version Specific Identifier.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ masterIdentifier_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ project_id:
+ $ref: _definitions.yaml#/project_id
+ resource_type:
+ description: ''
+ type:
+ - string
+ - 'null'
+ securityLabel_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ securityLabel_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ securityLabel_0_coding_0_system:
+ description: Document security-tags.. Code defined by a terminology system.. Identity
+ of the terminology system.
+ type:
+ - string
+ - 'null'
+ state:
+ $ref: _definitions.yaml#/state
+ status:
+ description: The status of this document reference.. http://hl7.org/fhir/document-reference-status
+ enum:
+ - current
+ - superseded
+ - entered-in-error
+ term:
+ description: The status of this document reference.. http://hl7.org/fhir/document-reference-status
+ termDef:
+ cde_id: http://hl7.org/fhir/document-reference-status
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/document-reference-status
+ term_url: http://hl7.org/fhir/document-reference-status
+ submitter_id:
+ type:
+ - string
+ - 'null'
+ type_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ type_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ type_coding_0_system:
+ description: Kind of document (LOINC if possible).. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ updated_datetime:
+ $ref: _definitions.yaml#/datetime
+required:
+- submitter_id
+- status
+- content
+submittable: true
+systemProperties:
+- id
+- project_id
+- created_datetime
+- updated_datetime
+- state
+title: DocumentReference
+type: object
+uniqueKeys:
+- - id
+- - project_id
+ - submitter_id
+validators: null
diff --git a/datadictionary/gdcdictionary/schemas/encounter.yaml b/datadictionary/gdcdictionary/schemas/encounter.yaml
new file mode 100644
index 00000000..4c0ca7c8
--- /dev/null
+++ b/datadictionary/gdcdictionary/schemas/encounter.yaml
@@ -0,0 +1,563 @@
+$schema: http://json-schema.org/draft-04/schema#
+additionalProperties: false
+category: Clinical
+description: " An interaction during which services are provided to the patient.\n\
+ \ \n An interaction between a patient and healthcare provider(s) for the purpose\n\
+ \ of providing healthcare service(s) or assessing the health status of a\n \
+ \ patient.\n http://hl7.org/fhir/encounter"
+id: encounter
+links:
+- backref: encounters
+ label: Encounter_subject_patient
+ multiplicity: many_to_many
+ name: subject_patient
+ required: false
+ target_type: patient
+namespace: http://hl7.org/fhir
+program: '*'
+project: '*'
+properties:
+ classHistory_0_class_fhir_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ classHistory_0_class_fhir_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ classHistory_0_class_fhir_system:
+ description: inpatient | outpatient | ambulatory | emergency +.. Identity of the
+ terminology system.
+ type:
+ - string
+ - 'null'
+ classHistory_0_period_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ classHistory_0_period_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ class_fhir_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ class_fhir_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ class_fhir_system:
+ description: Classification of patient encounter.. Identity of the terminology
+ system.
+ type:
+ - string
+ - 'null'
+ created_datetime:
+ $ref: _definitions.yaml#/datetime
+ diagnosis_0_condition_reference:
+ description: The diagnosis or procedure relevant to the encounter.. Literal reference,
+ Relative, internal or absolute URL.
+ type:
+ - string
+ - 'null'
+ diagnosis_0_condition_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ diagnosis_0_rank:
+ description: Ranking of the diagnosis (for each role type).
+ type:
+ - number
+ - 'null'
+ diagnosis_0_use_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ diagnosis_0_use_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ diagnosis_0_use_coding_0_system:
+ description: None. Code defined by a terminology system.. Identity of the terminology
+ system.
+ type:
+ - string
+ - 'null'
+ hospitalization_admitSource_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ hospitalization_admitSource_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ hospitalization_admitSource_coding_0_system:
+ description: None. Code defined by a terminology system.. Identity of the terminology
+ system.
+ type:
+ - string
+ - 'null'
+ hospitalization_destination_reference:
+ description: Location/organization to which the patient is discharged.. Literal
+ reference, Relative, internal or absolute URL.
+ type:
+ - string
+ - 'null'
+ hospitalization_destination_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ hospitalization_dietPreference_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ hospitalization_dietPreference_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ hospitalization_dietPreference_0_coding_0_system:
+ description: None. Code defined by a terminology system.. Identity of the terminology
+ system.
+ type:
+ - string
+ - 'null'
+ hospitalization_dischargeDisposition_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ hospitalization_dischargeDisposition_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ hospitalization_dischargeDisposition_coding_0_system:
+ description: None. Code defined by a terminology system.. Identity of the terminology
+ system.
+ type:
+ - string
+ - 'null'
+ hospitalization_origin_reference:
+ description: The location/organization from which the patient came before admission..
+ Literal reference, Relative, internal or absolute URL.
+ type:
+ - string
+ - 'null'
+ hospitalization_origin_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ hospitalization_preAdmissionIdentifier_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ hospitalization_preAdmissionIdentifier_use:
+ description: Pre-admission identifier.. The purpose of this identifier.. http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: Pre-admission identifier.. The purpose of this identifier.. http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ hospitalization_preAdmissionIdentifier_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ hospitalization_reAdmission_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ hospitalization_reAdmission_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ hospitalization_reAdmission_coding_0_system:
+ description: The type of hospital re-admission that has occurred (if any). If
+ the value is absent, then this is not identified as a readmission.. Code defined
+ by a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ hospitalization_specialArrangement_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ hospitalization_specialArrangement_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ hospitalization_specialArrangement_0_coding_0_system:
+ description: Any special requests that have been made for this hospitalization
+ encounter, such as the provision of specific equipment or other things.. Code
+ defined by a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ hospitalization_specialCourtesy_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ hospitalization_specialCourtesy_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ hospitalization_specialCourtesy_0_coding_0_system:
+ description: Special courtesies (VIP, board member).. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ id:
+ description: Logical id of this artifact.
+ type:
+ - string
+ - 'null'
+ identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ identifier_0_use:
+ description: Identifier(s) by which this encounter is known.. The purpose of this
+ identifier.. http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: Identifier(s) by which this encounter is known.. The purpose of
+ this identifier.. http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ language:
+ description: Language of the resource content.
+ type:
+ - string
+ - 'null'
+ length_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ length_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.
+ type:
+ - string
+ - 'null'
+ length_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ length_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ length_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ location_0_location_reference:
+ description: Location the encounter takes place.. Literal reference, Relative,
+ internal or absolute URL.
+ type:
+ - string
+ - 'null'
+ location_0_location_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ location_0_period_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ location_0_period_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ location_0_physicalType_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ location_0_physicalType_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ location_0_physicalType_coding_0_system:
+ description: This will be used to specify the required levels (bed/ward/room/etc.)
+ desired to be recorded to simplify either messaging or query.. Code defined
+ by a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ location_0_status:
+ description: The status of the participants' presence at the specified location
+ during the period specified. If the participant is no longer at the location,
+ then the period will have an end date/time.. http://hl7.org/fhir/encounter-location-status
+ enum:
+ - planned
+ - active
+ - reserved
+ - completed
+ term:
+ description: The status of the participants' presence at the specified location
+ during the period specified. If the participant is no longer at the location,
+ then the period will have an end date/time.. http://hl7.org/fhir/encounter-location-status
+ termDef:
+ cde_id: http://hl7.org/fhir/encounter-location-status
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/encounter-location-status
+ term_url: http://hl7.org/fhir/encounter-location-status
+ participant_0_individual_reference:
+ description: Persons involved in the encounter other than the patient.. Literal
+ reference, Relative, internal or absolute URL.
+ type:
+ - string
+ - 'null'
+ participant_0_individual_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ participant_0_period_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ participant_0_period_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ participant_0_type_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ participant_0_type_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ participant_0_type_0_coding_0_system:
+ description: Role of participant in encounter.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ period_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ period_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ priority_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ priority_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ priority_coding_0_system:
+ description: Indicates the urgency of the encounter.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ project_id:
+ $ref: _definitions.yaml#/project_id
+ reasonCode_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ reasonCode_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ reasonCode_0_coding_0_system:
+ description: Coded reason the encounter takes place.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ resource_type:
+ description: ''
+ type:
+ - string
+ - 'null'
+ serviceType_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ serviceType_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ serviceType_coding_0_system:
+ description: Specific type of service.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ state:
+ $ref: _definitions.yaml#/state
+ status:
+ description: None. http://hl7.org/fhir/encounter-status
+ enum:
+ - planned
+ - arrived
+ - triaged
+ - in-progress
+ - onleave
+ - finished
+ - cancelled
+ - entered-in-error
+ - unknown
+ term:
+ description: None. http://hl7.org/fhir/encounter-status
+ termDef:
+ cde_id: http://hl7.org/fhir/encounter-status
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/encounter-status
+ term_url: http://hl7.org/fhir/encounter-status
+ statusHistory_0_period_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ statusHistory_0_period_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ statusHistory_0_status:
+ description: None. http://hl7.org/fhir/encounter-status
+ enum:
+ - planned
+ - arrived
+ - triaged
+ - in-progress
+ - onleave
+ - finished
+ - cancelled
+ - entered-in-error
+ - unknown
+ term:
+ description: None. http://hl7.org/fhir/encounter-status
+ termDef:
+ cde_id: http://hl7.org/fhir/encounter-status
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/encounter-status
+ term_url: http://hl7.org/fhir/encounter-status
+ submitter_id:
+ type:
+ - string
+ - 'null'
+ type_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ type_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ type_0_coding_0_system:
+ description: Specific type of encounter (e.g. e-mail consultation, surgical day-care,
+ skilled nursing, rehabilitation).. Code defined by a terminology system.. Identity
+ of the terminology system.
+ type:
+ - string
+ - 'null'
+ updated_datetime:
+ $ref: _definitions.yaml#/datetime
+required:
+- submitter_id
+- status
+- class_fhir
+submittable: true
+systemProperties:
+- id
+- project_id
+- created_datetime
+- updated_datetime
+- state
+title: Encounter
+type: object
+uniqueKeys:
+- - id
+- - project_id
+ - submitter_id
+validators: null
diff --git a/datadictionary/gdcdictionary/schemas/experiment.yaml b/datadictionary/gdcdictionary/schemas/experiment.yaml
deleted file mode 100644
index b60cd068..00000000
--- a/datadictionary/gdcdictionary/schemas/experiment.yaml
+++ /dev/null
@@ -1,108 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "experiment"
-title: Experiment
-type: object
-namespace: http://bloodprofilingatlas.org/bpa/
-category: administrative
-program: '*'
-project: '*'
-description: >
- A coordinated set of actions and observations designed to generate data, with the ultimate goal
- of discovery or hypothesis testing.
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - created_datetime
- - updated_datetime
- - state
-
-links:
- - name: projects
- backref: experiments
- label: performed_for
- target_type: project
- multiplicity: many_to_one
- required: true
-
-required:
- - submitter_id
- - type
- - projects
-
-uniqueKeys:
- - [ id ]
- - [ project_id, submitter_id ]
-
-properties:
- type:
- enum: [ "experiment" ]
- id:
- $ref: "_definitions.yaml#/UUID"
- systemAlias: node_id
- state:
- $ref: "_definitions.yaml#/state"
- submitter_id:
- type:
- - string
- - "null"
- number_experimental_group:
- description: "The number denoting this experiment's place within the group within the whole."
- type:
- - integer
- number_samples_per_experimental_group:
- description: "The number of samples contained within this experimental group."
- type:
- - integer
- experimental_description:
- description: "A brief description of the experiment being performed."
- type:
- - string
- experimental_intent:
- description: "Summary of the goals the experiment is designed to discover."
- type:
- - string
- associated_experiment:
- description: "The submitter_id for any experiment with which this experiment is associated, paired, or matched."
- type:
- - string
- type_of_sample:
- description: "String indicator identifying the types of samples as contrived or clinical."
- type:
- - string
- type_of_specimen:
- description: "Broad description of the specimens used in the experiment."
- type:
- - string
- marker_panel_description:
- description: "Brief description of the marker panel used in this experiment."
- type: string
- somatic_mutations_identified:
- description: "Are somatic mutations identified for this experiment?"
- type: boolean
- indels_identified:
- description: "Are indels identified in this experiment?"
- type: boolean
- copy_numbers_identified:
- description: "Are copy number variations identified in this experiment?"
- type: boolean
- type_of_data:
- description: "Is the data raw or processed?"
- enum:
- - Raw
- - Processed
- data_description:
- description: "Brief description of the data being provided for this experiment."
- type: string
- projects:
- $ref: "_definitions.yaml#/to_one_project"
- project_id:
- $ref: "_definitions.yaml#/project_id"
- created_datetime:
- $ref: "_definitions.yaml#/datetime"
- updated_datetime:
- $ref: "_definitions.yaml#/datetime"
diff --git a/datadictionary/gdcdictionary/schemas/experimental_metadata.yaml b/datadictionary/gdcdictionary/schemas/experimental_metadata.yaml
deleted file mode 100644
index cd4f321a..00000000
--- a/datadictionary/gdcdictionary/schemas/experimental_metadata.yaml
+++ /dev/null
@@ -1,77 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "experimental_metadata"
-title: Experimental Metadata
-type: object
-namespace: http://gdc.nci.nih.gov
-category: metadata_file
-project: '*'
-program: '*'
-description: >
- Data file containing the metadata for the experiment performed.
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - created_datetime
- - updated_datetime
- - state
- - file_state
- - error_type
-
-links:
- - exclusive: false
- required: true
- subgroup:
- - name: core_metadata_collections
- backref: experiment_metadata_files
- label: data_from
- target_type: core_metadata_collection
- multiplicity: many_to_many
- required: false
- - name: experiments
- backref: experiment_metadata_files
- label: derived_from
- target_type: experiment
- multiplicity: many_to_many
- required: false
-
-required:
- - submitter_id
- - type
- - file_name
- - file_size
- - md5sum
- - data_category
- - data_type
- - data_format
-
-uniqueKeys:
- - [ id ]
- - [ project_id, submitter_id ]
-
-properties:
- $ref: "_definitions.yaml#/data_file_properties"
- type:
- enum: [ "experimental_metadata" ]
- data_category:
- term:
- $ref: "_terms.yaml#/data_category"
- type:
- - string
- data_type:
- term:
- $ref: "_terms.yaml#/data_type"
- enum: [ "Experimental Metadata" ]
- data_format:
- term:
- $ref: "_terms.yaml#/data_format"
- type:
- - string
- experiments:
- $ref: "_definitions.yaml#/to_one"
- core_metadata_collections:
- $ref: "_definitions.yaml#/to_many"
diff --git a/datadictionary/gdcdictionary/schemas/exposure.yaml b/datadictionary/gdcdictionary/schemas/exposure.yaml
deleted file mode 100644
index 76cfe72c..00000000
--- a/datadictionary/gdcdictionary/schemas/exposure.yaml
+++ /dev/null
@@ -1,132 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "exposure"
-title: Exposure
-type: object
-namespace: http://gdc.nci.nih.gov
-category: clinical
-program: '*'
-project: '*'
-description: >
- Clinically relevant patient information not immediately resulting from genetic predispositions.
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - state
- - created_datetime
- - updated_datetime
-
-required:
- - submitter_id
- - type
-
-links:
- - name: cases
- backref: exposures
- label: describes
- target_type: case
- multiplicity: many_to_one
- required: true
-
-preferred:
- - cigarettes_per_day
- - years_smoked
-
-uniqueKeys:
- #unclear if want submitter ID for clinical
- - [id]
- - [project_id, submitter_id]
-
-properties:
- type:
- enum: [ "exposure" ]
-
- id:
- $ref: "_definitions.yaml#/UUID"
- systemAlias: node_id
-
- state:
- $ref: "_definitions.yaml#/state"
-
- submitter_id:
- type:
- - string
- - "null"
-
- alcohol_history:
- term:
- $ref: "_terms.yaml#/alcohol_history"
- type: string
-
- alcohol_intensity:
- term:
- $ref: "_terms.yaml#/alcohol_intensity"
- type: string
-
- bmi:
- term:
- $ref: "_terms.yaml#/bmi"
- type: number
-
- cigarettes_per_day:
- term:
- $ref: "_terms.yaml#/cigarettes_per_day"
- type: number
-
- height:
- term:
- $ref: "_terms.yaml#/height"
- type: number
-
- pack_years_smoked:
- term:
- $ref: "_terms.yaml#/pack_years_smoked"
- type: number
-
- tobacco_smoking_onset_year:
- term:
- $ref: "_terms.yaml#/tobacco_smoking_onset_year"
- type: integer
-
- tobacco_smoking_quit_year:
- term:
- $ref: "_terms.yaml#/tobacco_smoking_quit_year"
- type: integer
-
- tobacco_smoking_status:
- term:
- $ref: "_terms.yaml#/tobacco_smoking_status"
- enum:
- - "1"
- - "2"
- - "3"
- - "4"
- - "5"
- - "6"
- - "7"
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- weight:
- term:
- $ref: "_terms.yaml#/weight"
- type: number
-
- years_smoked:
- term:
- $ref: "_terms.yaml#/years_smoked"
- type: number
-
- cases:
- $ref: "_definitions.yaml#/to_one"
- project_id:
- $ref: "_definitions.yaml#/project_id"
- created_datetime:
- $ref: "_definitions.yaml#/datetime"
- updated_datetime:
- $ref: "_definitions.yaml#/datetime"
diff --git a/datadictionary/gdcdictionary/schemas/family_history.yaml b/datadictionary/gdcdictionary/schemas/family_history.yaml
deleted file mode 100644
index add3a152..00000000
--- a/datadictionary/gdcdictionary/schemas/family_history.yaml
+++ /dev/null
@@ -1,98 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "family_history"
-title: Family History
-type: object
-namespace: http://gdc.nci.nih.gov
-category: clinical
-program: '*'
-project: '*'
-description: >
- Record of a patient's background regarding cancer events of blood relatives.
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - state
- - created_datetime
- - updated_datetime
-
-required:
- - submitter_id
- - type
-
-links:
- - name: cases
- backref: family_histories
- label: describes
- target_type: case
- multiplicity: many_to_one
- required: true
-
-
-uniqueKeys:
- #unclear if want submitter ID for clinical
- - [id]
- - [project_id, submitter_id]
-
-properties:
- type:
- enum: [ "family_history" ]
-
- id:
- $ref: "_definitions.yaml#/UUID"
- systemAlias: node_id
-
- state:
- $ref: "_definitions.yaml#/state"
-
- submitter_id:
- type:
- - string
- - "null"
-
- relative_with_cancer_history:
- term:
- $ref: "_terms.yaml#/relative_with_cancer_history"
- enum:
- - "yes"
- - "no"
- - unknown
- - not reported
-
- relationship_type:
- term:
- $ref: "_terms.yaml#/relationship_type"
- type: string
-
- relationship_gender:
- term:
- $ref: "_terms.yaml#/gender"
- enum:
- - female
- - male
- - unknown
- - unspecified
- - not reported
-
- relationship_age_at_diagnosis:
- term:
- $ref: "_terms.yaml#/relationship_age_at_diagnosis"
- type: number
-
- relationship_primary_diagnosis:
- term:
- $ref: "_terms.yaml#/primary_diagnosis"
- type: string
-
- cases:
- $ref: "_definitions.yaml#/to_one"
- project_id:
- $ref: "_definitions.yaml#/project_id"
- created_datetime:
- $ref: "_definitions.yaml#/datetime"
- updated_datetime:
- $ref: "_definitions.yaml#/datetime"
diff --git a/datadictionary/gdcdictionary/schemas/imaging_study.yaml b/datadictionary/gdcdictionary/schemas/imaging_study.yaml
new file mode 100644
index 00000000..5ef4786f
--- /dev/null
+++ b/datadictionary/gdcdictionary/schemas/imaging_study.yaml
@@ -0,0 +1,307 @@
+$schema: http://json-schema.org/draft-04/schema#
+additionalProperties: false
+category: Clinical
+description: " A set of images produced in single study (one or more series of references\n\
+ \ images).\n \n Representation of the content produced in a DICOM imaging\
+ \ study. A study\n comprises a set of series, each of which includes a set of\
+ \ Service-Object\n Pair Instances (SOP Instances - images or other data) acquired\
+ \ or produced\n in a common context. A series is of only one modality (e.g.\
+ \ X-ray, CT, MR,\n ultrasound), but a study may have multiple series of different\
+ \ modalities.\n http://hl7.org/fhir/imagingstudy"
+id: imaging_study
+links: []
+namespace: http://hl7.org/fhir
+program: '*'
+project: '*'
+properties:
+ created_datetime:
+ $ref: _definitions.yaml#/datetime
+ description:
+ description: Institution-generated description.
+ type:
+ - string
+ - 'null'
+ id:
+ description: Logical id of this artifact.
+ type:
+ - string
+ - 'null'
+ identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ identifier_0_use:
+ description: Identifiers for the whole study.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: Identifiers for the whole study.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ language:
+ description: Language of the resource content.
+ type:
+ - string
+ - 'null'
+ modality_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ modality_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ modality_0_system:
+ description: All series modality if actual acquisition modalities.. Identity of
+ the terminology system.
+ type:
+ - string
+ - 'null'
+ numberOfInstances:
+ description: Number of Study Related Instances.
+ type:
+ - number
+ - 'null'
+ numberOfSeries:
+ description: Number of Study Related Series.
+ type:
+ - number
+ - 'null'
+ procedureCode_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ procedureCode_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ procedureCode_0_coding_0_system:
+ description: The performed procedure code.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ project_id:
+ $ref: _definitions.yaml#/project_id
+ reasonCode_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ reasonCode_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ reasonCode_0_coding_0_system:
+ description: Why the study was requested.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ resource_type:
+ description: ''
+ type:
+ - string
+ - 'null'
+ series_0_bodySite_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ series_0_bodySite_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ series_0_bodySite_system:
+ description: Body part examined.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ series_0_description:
+ description: A short human readable summary of the series.
+ type:
+ - string
+ - 'null'
+ series_0_endpoint_0_reference:
+ description: Series access endpoint.. Literal reference, Relative, internal or
+ absolute URL.
+ type:
+ - string
+ - 'null'
+ series_0_endpoint_0_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ series_0_instance_0_number:
+ description: The number of this instance in the series.
+ type:
+ - number
+ - 'null'
+ series_0_instance_0_sopClass_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ series_0_instance_0_sopClass_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ series_0_instance_0_sopClass_system:
+ description: DICOM class type.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ series_0_instance_0_title:
+ description: Description of instance.
+ type:
+ - string
+ - 'null'
+ series_0_instance_0_uid:
+ description: DICOM SOP Instance UID.
+ type:
+ - string
+ series_0_laterality_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ series_0_laterality_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ series_0_laterality_system:
+ description: Body part laterality.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ series_0_modality_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ series_0_modality_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ series_0_modality_system:
+ description: The modality of the instances in the series.. Identity of the terminology
+ system.
+ type:
+ - string
+ - 'null'
+ series_0_number:
+ description: Numeric identifier of this series.
+ type:
+ - number
+ - 'null'
+ series_0_numberOfInstances:
+ description: Number of Series Related Instances.
+ type:
+ - number
+ - 'null'
+ series_0_performer_0_actor_reference:
+ description: Who performed the series.. Literal reference, Relative, internal
+ or absolute URL.
+ type:
+ - string
+ - 'null'
+ series_0_performer_0_actor_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ series_0_specimen_0_reference:
+ description: Specimen imaged.. Literal reference, Relative, internal or absolute
+ URL.
+ type:
+ - string
+ - 'null'
+ series_0_specimen_0_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ series_0_started:
+ description: When the series started.
+ type:
+ - string
+ - 'null'
+ series_0_uid:
+ description: DICOM Series Instance UID for the series.
+ type:
+ - string
+ started:
+ description: When the study was started.
+ type:
+ - string
+ - 'null'
+ state:
+ $ref: _definitions.yaml#/state
+ status:
+ description: The current state of the ImagingStudy.. http://hl7.org/fhir/imagingstudy-status
+ enum:
+ - registered
+ - available
+ - cancelled
+ - entered-in-error
+ - unknown
+ term:
+ description: The current state of the ImagingStudy.. http://hl7.org/fhir/imagingstudy-status
+ termDef:
+ cde_id: http://hl7.org/fhir/imagingstudy-status
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/imagingstudy-status
+ term_url: http://hl7.org/fhir/imagingstudy-status
+ submitter_id:
+ type:
+ - string
+ - 'null'
+ updated_datetime:
+ $ref: _definitions.yaml#/datetime
+required:
+- submitter_id
+- status
+- subject
+submittable: true
+systemProperties:
+- id
+- project_id
+- created_datetime
+- updated_datetime
+- state
+title: ImagingStudy
+type: object
+uniqueKeys:
+- - id
+- - project_id
+ - submitter_id
+validators: null
diff --git a/datadictionary/gdcdictionary/schemas/keyword.yaml b/datadictionary/gdcdictionary/schemas/keyword.yaml
deleted file mode 100644
index 99a3a050..00000000
--- a/datadictionary/gdcdictionary/schemas/keyword.yaml
+++ /dev/null
@@ -1,61 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "keyword"
-title: Keyword
-type: object
-namespace: http://gdc.nci.nih.gov
-category: administrative
-program: '*'
-project: '*'
-description: "A keyword for a project."
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - state
- - created_datetime
- - updated_datetime
-
-links:
- - name: projects
- backref: keywords
- label: describe
- target_type: project
- multiplicity: many_to_many
- required: true
-
-required:
- - submitter_id
- - type
- - projects
-
-uniqueKeys:
- - [ id ]
- - [ project_id, submitter_id ]
-
-properties:
- type:
- enum: [ "keyword" ]
- id:
- $ref: "_definitions.yaml#/UUID"
- systemAlias: node_id
- state:
- $ref: "_definitions.yaml#/state"
- submitter_id:
- type:
- - string
- - "null"
- keyword_name:
- description: "The name of the keyword."
- type: string
- projects:
- $ref: "_definitions.yaml#/to_many_project"
- project_id:
- type: string
- created_datetime:
- $ref: "_definitions.yaml#/datetime"
- updated_datetime:
- $ref: "_definitions.yaml#/datetime"
diff --git a/datadictionary/gdcdictionary/schemas/location.yaml b/datadictionary/gdcdictionary/schemas/location.yaml
new file mode 100644
index 00000000..995217c8
--- /dev/null
+++ b/datadictionary/gdcdictionary/schemas/location.yaml
@@ -0,0 +1,239 @@
+$schema: http://json-schema.org/draft-04/schema#
+additionalProperties: false
+category: Administrative
+description: " Details and position information for a physical place.\n \n Details\
+ \ and position information for a physical place where services are\n provided\
+ \ and resources and participants may be stored, found, contained, or\n accommodated.\n\
+ \ http://hl7.org/fhir/location"
+id: location
+links: []
+namespace: http://hl7.org/fhir
+program: '*'
+project: '*'
+properties:
+ alias:
+ description: A list of alternate names that the location is known as, or was known
+ as, in the past.
+ type:
+ - string
+ - 'null'
+ availabilityExceptions:
+ description: Description of availability exceptions.
+ type:
+ - string
+ - 'null'
+ created_datetime:
+ $ref: _definitions.yaml#/datetime
+ description:
+ description: Additional details about the location that could be displayed as
+ further information to identify the location beyond its name.
+ type:
+ - string
+ - 'null'
+ hoursOfOperation_0_allDay:
+ description: The Location is open all day.
+ type:
+ - boolean
+ - 'null'
+ hoursOfOperation_0_closingTime:
+ description: Time that the Location closes.
+ type:
+ - string
+ - 'null'
+ hoursOfOperation_0_daysOfWeek:
+ description: Indicates which days of the week are available between the start
+ and end Times.. http://hl7.org/fhir/days-of-week
+ enum:
+ - mon
+ - tue
+ - wed
+ - thu
+ - fri
+ - sat
+ - sun
+ term:
+ description: Indicates which days of the week are available between the start
+ and end Times.. http://hl7.org/fhir/days-of-week
+ termDef:
+ cde_id: http://hl7.org/fhir/days-of-week
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/days-of-week
+ term_url: http://hl7.org/fhir/days-of-week
+ hoursOfOperation_0_openingTime:
+ description: Time that the Location opens.
+ type:
+ - string
+ - 'null'
+ id:
+ description: Logical id of this artifact.
+ type:
+ - string
+ - 'null'
+ identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ identifier_0_use:
+ description: Unique code or number identifying the location to its users.. The
+ purpose of this identifier.. http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: Unique code or number identifying the location to its users.. The
+ purpose of this identifier.. http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ language:
+ description: Language of the resource content.
+ type:
+ - string
+ - 'null'
+ mode:
+ description: Indicates whether a resource instance represents a specific location
+ or a class of locations.. http://hl7.org/fhir/location-mode
+ enum:
+ - instance
+ - kind
+ term:
+ description: Indicates whether a resource instance represents a specific location
+ or a class of locations.. http://hl7.org/fhir/location-mode
+ termDef:
+ cde_id: http://hl7.org/fhir/location-mode
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/location-mode
+ term_url: http://hl7.org/fhir/location-mode
+ name:
+ description: Name of the location as used by humans.
+ type:
+ - string
+ - 'null'
+ operationalStatus_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ operationalStatus_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ operationalStatus_system:
+ description: The operational status of the location (typically only for a bed/room)..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ physicalType_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ physicalType_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ physicalType_coding_0_system:
+ description: Physical form of the location, e.g. building, room, vehicle, road..
+ Code defined by a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ position_altitude:
+ description: Altitude with WGS84 datum.
+ type:
+ - number
+ - 'null'
+ position_latitude:
+ description: Latitude with WGS84 datum.
+ type:
+ - number
+ position_longitude:
+ description: Longitude with WGS84 datum.
+ type:
+ - number
+ project_id:
+ $ref: _definitions.yaml#/project_id
+ resource_type:
+ description: ''
+ type:
+ - string
+ - 'null'
+ state:
+ $ref: _definitions.yaml#/state
+ status:
+ description: The status property covers the general availability of the resource,
+ not the current value which may be covered by the operationStatus, or by a schedule/slots
+ if they are configured for the location.. http://hl7.org/fhir/location-status
+ enum:
+ - active
+ - suspended
+ - inactive
+ term:
+ description: The status property covers the general availability of the resource,
+ not the current value which may be covered by the operationStatus, or by a
+ schedule/slots if they are configured for the location.. http://hl7.org/fhir/location-status
+ termDef:
+ cde_id: http://hl7.org/fhir/location-status
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/location-status
+ term_url: http://hl7.org/fhir/location-status
+ submitter_id:
+ type:
+ - string
+ - 'null'
+ type_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ type_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ type_0_coding_0_system:
+ description: Type of function performed.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ updated_datetime:
+ $ref: _definitions.yaml#/datetime
+required:
+- submitter_id
+submittable: true
+systemProperties:
+- id
+- project_id
+- created_datetime
+- updated_datetime
+- state
+title: Location
+type: object
+uniqueKeys:
+- - id
+- - project_id
+ - submitter_id
+validators: null
diff --git a/datadictionary/gdcdictionary/schemas/medication.yaml b/datadictionary/gdcdictionary/schemas/medication.yaml
new file mode 100644
index 00000000..09c411ce
--- /dev/null
+++ b/datadictionary/gdcdictionary/schemas/medication.yaml
@@ -0,0 +1,334 @@
+$schema: http://json-schema.org/draft-04/schema#
+additionalProperties: false
+category: Clinical
+description: " Definition of a Medication.\n \n This resource is primarily used\
+ \ for the identification and definition of a\n medication for the purposes of\
+ \ prescribing, dispensing, and administering a\n medication as well as for making\
+ \ statements about medication use.\n http://hl7.org/fhir/medication"
+id: medication
+links: []
+namespace: http://hl7.org/fhir
+program: '*'
+project: '*'
+properties:
+ amount_denominator_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ amount_denominator_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ amount_denominator_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ amount_denominator_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ amount_denominator_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ amount_numerator_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ amount_numerator_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ amount_numerator_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ amount_numerator_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ amount_numerator_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ batch_expirationDate:
+ description: When batch will expire.
+ type:
+ - string
+ - 'null'
+ batch_lotNumber:
+ description: Identifier assigned to batch.
+ type:
+ - string
+ - 'null'
+ code_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ code_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ code_coding_0_system:
+ description: Codes that identify this medication.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ created_datetime:
+ $ref: _definitions.yaml#/datetime
+ form_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ form_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ form_coding_0_system:
+ description: powder | tablets | capsule +.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ id:
+ description: Logical id of this artifact.
+ type:
+ - string
+ - 'null'
+ identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ identifier_0_use:
+ description: Business identifier for this medication.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: Business identifier for this medication.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ ingredient_0_isActive:
+ description: Active ingredient indicator.
+ type:
+ - boolean
+ - 'null'
+ ingredient_0_itemCodeableConcept_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ ingredient_0_itemCodeableConcept_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ ingredient_0_itemCodeableConcept_coding_0_system:
+ description: The actual ingredient or content.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ ingredient_0_itemReference_reference:
+ description: The actual ingredient or content.. Literal reference, Relative, internal
+ or absolute URL.
+ type:
+ - string
+ - 'null'
+ ingredient_0_itemReference_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ ingredient_0_strength_denominator_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ ingredient_0_strength_denominator_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ ingredient_0_strength_denominator_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ ingredient_0_strength_denominator_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ ingredient_0_strength_denominator_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ ingredient_0_strength_numerator_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ ingredient_0_strength_numerator_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ ingredient_0_strength_numerator_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ ingredient_0_strength_numerator_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ ingredient_0_strength_numerator_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ language:
+ description: Language of the resource content.
+ type:
+ - string
+ - 'null'
+ project_id:
+ $ref: _definitions.yaml#/project_id
+ resource_type:
+ description: ''
+ type:
+ - string
+ - 'null'
+ state:
+ $ref: _definitions.yaml#/state
+ status:
+ description: A code to indicate if the medication is in active use.. http://hl7.org/fhir/CodeSystem/medication-status
+ enum:
+ - active
+ - inactive
+ - entered-in-error
+ term:
+ description: A code to indicate if the medication is in active use.. http://hl7.org/fhir/CodeSystem/medication-status
+ termDef:
+ cde_id: http://hl7.org/fhir/CodeSystem/medication-status
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/CodeSystem/medication-status
+ term_url: http://hl7.org/fhir/CodeSystem/medication-status
+ submitter_id:
+ type:
+ - string
+ - 'null'
+ updated_datetime:
+ $ref: _definitions.yaml#/datetime
+required:
+- submitter_id
+submittable: true
+systemProperties:
+- id
+- project_id
+- created_datetime
+- updated_datetime
+- state
+title: Medication
+type: object
+uniqueKeys:
+- - id
+- - project_id
+ - submitter_id
+validators: null
diff --git a/datadictionary/gdcdictionary/schemas/medication_request.yaml b/datadictionary/gdcdictionary/schemas/medication_request.yaml
new file mode 100644
index 00000000..9dae963b
--- /dev/null
+++ b/datadictionary/gdcdictionary/schemas/medication_request.yaml
@@ -0,0 +1,978 @@
+$schema: http://json-schema.org/draft-04/schema#
+additionalProperties: false
+category: Clinical
+description: " Ordering of medication for patient or group.\n \n An order or\
+ \ request for both supply of the medication and the instructions\n for administration\
+ \ of the medication to a patient. The resource is called\n \"MedicationRequest\"\
+ \ rather than \"MedicationPrescription\" or\n \"MedicationOrder\" to generalize\
+ \ the use across inpatient and outpatient\n settings, including care plans, etc.,\
+ \ and to harmonize with workflow\n patterns.\n http://hl7.org/fhir/medicationrequest"
+id: medication_request
+links:
+- backref: medication_requests
+ label: MedicationRequest_encounter_encounter
+ multiplicity: many_to_many
+ name: encounter_encounter
+ required: false
+ target_type: encounter
+- backref: medication_requests
+ label: MedicationRequest_medicationReference_medication
+ multiplicity: many_to_many
+ name: medicationReference_medication
+ required: false
+ target_type: medication
+- backref: medication_requests
+ label: MedicationRequest_subject_patient
+ multiplicity: many_to_many
+ name: subject_patient
+ required: false
+ target_type: patient
+namespace: http://hl7.org/fhir
+program: '*'
+project: '*'
+properties:
+ authoredOn:
+ description: When request was initially authored.
+ type:
+ - string
+ - 'null'
+ category_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ category_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ category_0_coding_0_system:
+ description: Indicates the type of medication request (for example, where the
+ medication is expected to be consumed or administered (i.e. inpatient or outpatient))..
+ Code defined by a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ courseOfTherapyType_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ courseOfTherapyType_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ courseOfTherapyType_coding_0_system:
+ description: The description of the overall patte3rn of the administration of
+ the medication to the patient.. Code defined by a terminology system.. Identity
+ of the terminology system.
+ type:
+ - string
+ - 'null'
+ created_datetime:
+ $ref: _definitions.yaml#/datetime
+ dispenseRequest_dispenseInterval_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_dispenseInterval_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_dispenseInterval_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_dispenseInterval_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_dispenseInterval_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ dispenseRequest_expectedSupplyDuration_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_expectedSupplyDuration_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_expectedSupplyDuration_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_expectedSupplyDuration_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_expectedSupplyDuration_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ dispenseRequest_initialFill_duration_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_initialFill_duration_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_initialFill_duration_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_initialFill_duration_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_initialFill_duration_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ dispenseRequest_initialFill_quantity_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_initialFill_quantity_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ dispenseRequest_initialFill_quantity_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_initialFill_quantity_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_initialFill_quantity_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ dispenseRequest_numberOfRepeatsAllowed:
+ description: Number of refills authorized.
+ type:
+ - number
+ - 'null'
+ dispenseRequest_performer_reference:
+ description: Intended dispenser.. Literal reference, Relative, internal or absolute
+ URL.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_performer_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ dispenseRequest_quantity_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_quantity_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ dispenseRequest_quantity_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_quantity_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_quantity_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ dispenseRequest_validityPeriod_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ dispenseRequest_validityPeriod_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ doNotPerform:
+ description: True if request is prohibiting action.
+ type:
+ - boolean
+ - 'null'
+ dosageInstruction_0_additionalInstruction_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_additionalInstruction_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_additionalInstruction_0_coding_0_system:
+ description: Supplemental instruction or warnings to the patient - e.g. "with
+ meals", "may cause drowsiness".. Code defined by a terminology system.. Identity
+ of the terminology system.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_asNeededBoolean:
+ description: Take "as needed" (for x).
+ type:
+ - boolean
+ - 'null'
+ dosageInstruction_0_asNeededCodeableConcept_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_asNeededCodeableConcept_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_asNeededCodeableConcept_coding_0_system:
+ description: Take "as needed" (for x).. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_doseAndRate_0_doseQuantity_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_doseAndRate_0_doseQuantity_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ dosageInstruction_0_doseAndRate_0_doseQuantity_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_doseAndRate_0_doseQuantity_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_doseAndRate_0_doseQuantity_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ dosageInstruction_0_doseAndRate_0_rateQuantity_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_doseAndRate_0_rateQuantity_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ dosageInstruction_0_doseAndRate_0_rateQuantity_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_doseAndRate_0_rateQuantity_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_doseAndRate_0_rateQuantity_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ dosageInstruction_0_maxDosePerAdministration_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_maxDosePerAdministration_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ dosageInstruction_0_maxDosePerAdministration_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_maxDosePerAdministration_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_maxDosePerAdministration_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ dosageInstruction_0_maxDosePerLifetime_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_maxDosePerLifetime_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ dosageInstruction_0_maxDosePerLifetime_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_maxDosePerLifetime_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_maxDosePerLifetime_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ dosageInstruction_0_maxDosePerPeriod_denominator_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_maxDosePerPeriod_denominator_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ dosageInstruction_0_maxDosePerPeriod_denominator_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_maxDosePerPeriod_denominator_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_maxDosePerPeriod_denominator_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ dosageInstruction_0_maxDosePerPeriod_numerator_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_maxDosePerPeriod_numerator_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ dosageInstruction_0_maxDosePerPeriod_numerator_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_maxDosePerPeriod_numerator_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_maxDosePerPeriod_numerator_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ dosageInstruction_0_method_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_method_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_method_coding_0_system:
+ description: Technique for administering medication.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_patientInstruction:
+ description: Patient or consumer oriented instructions.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_route_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_route_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_route_coding_0_system:
+ description: How drug should enter body.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_sequence:
+ description: The order of the dosage instructions.
+ type:
+ - number
+ - 'null'
+ dosageInstruction_0_site_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_site_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_site_coding_0_system:
+ description: Body site to administer to.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_text:
+ description: Free text dosage instructions e.g. SIG.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_timing_event:
+ description: When the event occurs.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_timing_repeat_count:
+ description: Number of times to repeat.
+ type:
+ - number
+ - 'null'
+ dosageInstruction_0_timing_repeat_countMax:
+ description: Maximum number of times to repeat.
+ type:
+ - number
+ - 'null'
+ dosageInstruction_0_timing_repeat_dayOfWeek:
+ description: If one or more days of week is provided, then the action happens
+ only on the specified day(s).. http://hl7.org/fhir/days-of-week
+ enum:
+ - mon
+ - tue
+ - wed
+ - thu
+ - fri
+ - sat
+ - sun
+ term:
+ description: If one or more days of week is provided, then the action happens
+ only on the specified day(s).. http://hl7.org/fhir/days-of-week
+ termDef:
+ cde_id: http://hl7.org/fhir/days-of-week
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/days-of-week
+ term_url: http://hl7.org/fhir/days-of-week
+ dosageInstruction_0_timing_repeat_duration:
+ description: How long when it happens.
+ type:
+ - number
+ - 'null'
+ dosageInstruction_0_timing_repeat_durationMax:
+ description: How long when it happens (Max).
+ type:
+ - number
+ - 'null'
+ dosageInstruction_0_timing_repeat_durationUnit:
+ description: s | min | h | d | wk | mo | a - unit of time (UCUM).
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_timing_repeat_frequency:
+ description: Event occurs frequency times per period.
+ type:
+ - number
+ - 'null'
+ dosageInstruction_0_timing_repeat_frequencyMax:
+ description: Event occurs up to frequencyMax times per period.
+ type:
+ - number
+ - 'null'
+ dosageInstruction_0_timing_repeat_offset:
+ description: Minutes from event (before or after).
+ type:
+ - number
+ - 'null'
+ dosageInstruction_0_timing_repeat_period:
+ description: Event occurs frequency times per period.
+ type:
+ - number
+ - 'null'
+ dosageInstruction_0_timing_repeat_periodMax:
+ description: Upper limit of period (3-4 hours).
+ type:
+ - number
+ - 'null'
+ dosageInstruction_0_timing_repeat_periodUnit:
+ description: s | min | h | d | wk | mo | a - unit of time (UCUM).
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_timing_repeat_timeOfDay:
+ description: Time of day for action.
+ type:
+ - string
+ - 'null'
+ dosageInstruction_0_timing_repeat_when:
+ description: Code for time period of occurrence.
+ type:
+ - string
+ - 'null'
+ groupIdentifier_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ groupIdentifier_use:
+ description: Composite request this is part of.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: Composite request this is part of.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ groupIdentifier_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ id:
+ description: Logical id of this artifact.
+ type:
+ - string
+ - 'null'
+ identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ identifier_0_use:
+ description: External ids for this request.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: External ids for this request.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ instantiatesCanonical:
+ description: Instantiates FHIR protocol or definition.
+ type:
+ - string
+ - 'null'
+ instantiatesUri:
+ description: Instantiates external protocol or definition.
+ type:
+ - string
+ - 'null'
+ intent:
+ description: Whether the request is a proposal, plan, or an original order.. http://hl7.org/fhir/CodeSystem/medicationrequest-intent
+ enum:
+ - proposal
+ - plan
+ - order
+ - original-order
+ - reflex-order
+ - filler-order
+ - instance-order
+ - option
+ term:
+ description: Whether the request is a proposal, plan, or an original order..
+ http://hl7.org/fhir/CodeSystem/medicationrequest-intent
+ termDef:
+ cde_id: http://hl7.org/fhir/CodeSystem/medicationrequest-intent
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/CodeSystem/medicationrequest-intent
+ term_url: http://hl7.org/fhir/CodeSystem/medicationrequest-intent
+ language:
+ description: Language of the resource content.
+ type:
+ - string
+ - 'null'
+ medicationCodeableConcept_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ medicationCodeableConcept_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ medicationCodeableConcept_coding_0_system:
+ description: Medication to be taken.. Code defined by a terminology system.. Identity
+ of the terminology system.
+ type:
+ - string
+ - 'null'
+ performerType_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ performerType_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ performerType_coding_0_system:
+ description: Desired kind of performer of the medication administration.. Code
+ defined by a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ priority:
+ description: Indicates how quickly the Medication Request should be addressed
+ with respect to other requests.. http://hl7.org/fhir/request-priority
+ enum:
+ - routine
+ - urgent
+ - asap
+ - stat
+ term:
+ description: Indicates how quickly the Medication Request should be addressed
+ with respect to other requests.. http://hl7.org/fhir/request-priority
+ termDef:
+ cde_id: http://hl7.org/fhir/request-priority
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/request-priority
+ term_url: http://hl7.org/fhir/request-priority
+ project_id:
+ $ref: _definitions.yaml#/project_id
+ reasonCode_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ reasonCode_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ reasonCode_0_coding_0_system:
+ description: Reason or indication for ordering or not ordering the medication..
+ Code defined by a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ reportedBoolean:
+ description: Reported rather than primary record.
+ type:
+ - boolean
+ - 'null'
+ resource_type:
+ description: ''
+ type:
+ - string
+ - 'null'
+ state:
+ $ref: _definitions.yaml#/state
+ status:
+ description: A code specifying the current state of the order. Generally, this
+ will be active or completed state.. http://hl7.org/fhir/CodeSystem/medicationrequest-status
+ enum:
+ - active
+ - on-hold
+ - cancelled
+ - completed
+ - entered-in-error
+ - stopped
+ - draft
+ - unknown
+ term:
+ description: A code specifying the current state of the order. Generally, this
+ will be active or completed state.. http://hl7.org/fhir/CodeSystem/medicationrequest-status
+ termDef:
+ cde_id: http://hl7.org/fhir/CodeSystem/medicationrequest-status
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/CodeSystem/medicationrequest-status
+ term_url: http://hl7.org/fhir/CodeSystem/medicationrequest-status
+ statusReason_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ statusReason_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ statusReason_coding_0_system:
+ description: Captures the reason for the current state of the MedicationRequest..
+ Code defined by a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ submitter_id:
+ type:
+ - string
+ - 'null'
+ substitution_allowedBoolean:
+ description: Whether substitution is allowed or not.
+ type:
+ - boolean
+ substitution_allowedCodeableConcept_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ substitution_allowedCodeableConcept_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ substitution_allowedCodeableConcept_coding_0_system:
+ description: Whether substitution is allowed or not.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ substitution_reason_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ substitution_reason_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ substitution_reason_coding_0_system:
+ description: Why should (not) substitution be made.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ updated_datetime:
+ $ref: _definitions.yaml#/datetime
+required:
+- submitter_id
+- status
+- intent
+- medicationCodeableConcept
+- medicationReference
+- subject
+submittable: true
+systemProperties:
+- id
+- project_id
+- created_datetime
+- updated_datetime
+- state
+title: MedicationRequest
+type: object
+uniqueKeys:
+- - id
+- - project_id
+ - submitter_id
+validators: null
diff --git a/datadictionary/gdcdictionary/schemas/observation.yaml b/datadictionary/gdcdictionary/schemas/observation.yaml
new file mode 100644
index 00000000..249d7d43
--- /dev/null
+++ b/datadictionary/gdcdictionary/schemas/observation.yaml
@@ -0,0 +1,1365 @@
+$schema: http://json-schema.org/draft-04/schema#
+additionalProperties: false
+category: Clinical
+description: " Measurements and simple assertions.\n \n Measurements and simple\
+ \ assertions made about a patient, device or other\n subject.\n http://hl7.org/fhir/observation"
+id: observation
+links:
+- backref: observations
+ label: Observation_encounter_encounter
+ multiplicity: many_to_many
+ name: encounter_encounter
+ required: false
+ target_type: encounter
+- backref: observations
+ label: Observation_specimen_specimen
+ multiplicity: many_to_many
+ name: specimen_specimen
+ required: false
+ target_type: specimen
+- backref: observations
+ label: Observation_subject_patient
+ multiplicity: many_to_many
+ name: subject_patient
+ required: false
+ target_type: patient
+- backref: observations
+ label: Observation_subject_research_study
+ multiplicity: many_to_many
+ name: subject_research_study
+ required: false
+ target_type: research_study
+namespace: http://hl7.org/fhir
+program: '*'
+project: '*'
+properties:
+ bodySite_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ bodySite_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ bodySite_coding_0_system:
+ description: Observed body part.. Code defined by a terminology system.. Identity
+ of the terminology system.
+ type:
+ - string
+ - 'null'
+ category_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ category_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ category_0_coding_0_system:
+ description: A code that classifies the general type of observation being made..
+ Code defined by a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ code_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ code_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ code_coding_0_system:
+ description: Type of observation (code / type).. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ component_0_code_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ component_0_code_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ component_0_code_coding_0_system:
+ description: Type of component observation (code / type).. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ component_0_dataAbsentReason_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ component_0_dataAbsentReason_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ component_0_dataAbsentReason_coding_0_system:
+ description: Provides a reason why the expected value in the element Observation.component.value[x]
+ is missing.. Code defined by a terminology system.. Identity of the terminology
+ system.
+ type:
+ - string
+ - 'null'
+ component_0_interpretation_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ component_0_interpretation_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ component_0_interpretation_0_coding_0_system:
+ description: High, low, normal, etc... Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ component_0_referenceRange_0_high_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ component_0_referenceRange_0_high_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ component_0_referenceRange_0_high_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ component_0_referenceRange_0_high_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ component_0_referenceRange_0_high_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ component_0_referenceRange_0_low_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ component_0_referenceRange_0_low_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ component_0_referenceRange_0_low_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ component_0_referenceRange_0_low_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ component_0_referenceRange_0_low_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ component_0_referenceRange_0_text:
+ description: Text based reference range in an observation.
+ type:
+ - string
+ - 'null'
+ component_0_valueBoolean:
+ description: Actual component result.
+ type:
+ - boolean
+ - 'null'
+ component_0_valueCodeableConcept_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ component_0_valueCodeableConcept_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ component_0_valueCodeableConcept_coding_0_system:
+ description: Actual component result.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ component_0_valueDateTime:
+ description: Actual component result.
+ type:
+ - string
+ - 'null'
+ component_0_valueInteger:
+ description: Actual component result.
+ type:
+ - number
+ - 'null'
+ component_0_valuePeriod_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ component_0_valuePeriod_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ component_0_valueQuantity_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ component_0_valueQuantity_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ component_0_valueQuantity_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ component_0_valueQuantity_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ component_0_valueQuantity_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ component_0_valueRange_high_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ component_0_valueRange_high_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ component_0_valueRange_high_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ component_0_valueRange_high_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ component_0_valueRange_high_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ component_0_valueRange_low_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ component_0_valueRange_low_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ component_0_valueRange_low_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ component_0_valueRange_low_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ component_0_valueRange_low_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ component_0_valueRatio_denominator_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ component_0_valueRatio_denominator_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ component_0_valueRatio_denominator_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ component_0_valueRatio_denominator_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ component_0_valueRatio_denominator_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ component_0_valueRatio_numerator_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ component_0_valueRatio_numerator_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ component_0_valueRatio_numerator_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ component_0_valueRatio_numerator_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ component_0_valueRatio_numerator_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ component_0_valueSampledData_data:
+ description: Decimal values with spaces, or "E" | "U" | "L".
+ type:
+ - string
+ - 'null'
+ component_0_valueSampledData_dimensions:
+ description: Number of sample points at each time point.
+ type:
+ - number
+ component_0_valueSampledData_factor:
+ description: Multiply data by this before adding to origin.
+ type:
+ - number
+ - 'null'
+ component_0_valueSampledData_lowerLimit:
+ description: Lower limit of detection.
+ type:
+ - number
+ - 'null'
+ component_0_valueSampledData_origin_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ component_0_valueSampledData_origin_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ component_0_valueSampledData_origin_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ component_0_valueSampledData_origin_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ component_0_valueSampledData_origin_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ component_0_valueSampledData_period:
+ description: Number of milliseconds between samples.
+ type:
+ - number
+ component_0_valueSampledData_upperLimit:
+ description: Upper limit of detection.
+ type:
+ - number
+ - 'null'
+ component_0_valueString:
+ description: Actual component result.
+ type:
+ - string
+ - 'null'
+ component_0_valueTime:
+ description: Actual component result.
+ type:
+ - string
+ - 'null'
+ created_datetime:
+ $ref: _definitions.yaml#/datetime
+ dataAbsentReason_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ dataAbsentReason_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ dataAbsentReason_coding_0_system:
+ description: Provides a reason why the expected value in the element Observation.value[x]
+ is missing.. Code defined by a terminology system.. Identity of the terminology
+ system.
+ type:
+ - string
+ - 'null'
+ effectiveDateTime:
+ description: Clinically relevant time/time-period for observation.
+ type:
+ - string
+ - 'null'
+ effectiveInstant:
+ description: Clinically relevant time/time-period for observation.
+ type:
+ - string
+ - 'null'
+ effectivePeriod_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ effectivePeriod_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ effectiveTiming_code_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ effectiveTiming_code_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ effectiveTiming_code_coding_0_system:
+ description: BID | TID | QID | AM | PM | QD | QOD | +.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ effectiveTiming_event:
+ description: When the event occurs.
+ type:
+ - string
+ - 'null'
+ effectiveTiming_repeat_boundsDuration_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ effectiveTiming_repeat_boundsDuration_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.
+ type:
+ - string
+ - 'null'
+ effectiveTiming_repeat_boundsDuration_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ effectiveTiming_repeat_boundsDuration_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ effectiveTiming_repeat_boundsDuration_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ effectiveTiming_repeat_boundsPeriod_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ effectiveTiming_repeat_boundsPeriod_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ effectiveTiming_repeat_count:
+ description: Number of times to repeat.
+ type:
+ - number
+ - 'null'
+ effectiveTiming_repeat_countMax:
+ description: Maximum number of times to repeat.
+ type:
+ - number
+ - 'null'
+ effectiveTiming_repeat_dayOfWeek:
+ description: If one or more days of week is provided, then the action happens
+ only on the specified day(s).. http://hl7.org/fhir/days-of-week
+ enum:
+ - mon
+ - tue
+ - wed
+ - thu
+ - fri
+ - sat
+ - sun
+ term:
+ description: If one or more days of week is provided, then the action happens
+ only on the specified day(s).. http://hl7.org/fhir/days-of-week
+ termDef:
+ cde_id: http://hl7.org/fhir/days-of-week
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/days-of-week
+ term_url: http://hl7.org/fhir/days-of-week
+ effectiveTiming_repeat_duration:
+ description: How long when it happens.
+ type:
+ - number
+ - 'null'
+ effectiveTiming_repeat_durationMax:
+ description: How long when it happens (Max).
+ type:
+ - number
+ - 'null'
+ effectiveTiming_repeat_durationUnit:
+ description: s | min | h | d | wk | mo | a - unit of time (UCUM).
+ type:
+ - string
+ - 'null'
+ effectiveTiming_repeat_frequency:
+ description: Event occurs frequency times per period.
+ type:
+ - number
+ - 'null'
+ effectiveTiming_repeat_frequencyMax:
+ description: Event occurs up to frequencyMax times per period.
+ type:
+ - number
+ - 'null'
+ effectiveTiming_repeat_offset:
+ description: Minutes from event (before or after).
+ type:
+ - number
+ - 'null'
+ effectiveTiming_repeat_period:
+ description: Event occurs frequency times per period.
+ type:
+ - number
+ - 'null'
+ effectiveTiming_repeat_periodMax:
+ description: Upper limit of period (3-4 hours).
+ type:
+ - number
+ - 'null'
+ effectiveTiming_repeat_periodUnit:
+ description: s | min | h | d | wk | mo | a - unit of time (UCUM).
+ type:
+ - string
+ - 'null'
+ effectiveTiming_repeat_timeOfDay:
+ description: Time of day for action.
+ type:
+ - string
+ - 'null'
+ effectiveTiming_repeat_when:
+ description: Code for time period of occurrence.
+ type:
+ - string
+ - 'null'
+ id:
+ description: Logical id of this artifact.
+ type:
+ - string
+ - 'null'
+ identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ identifier_0_use:
+ description: Business Identifier for observation.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: Business Identifier for observation.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ interpretation_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ interpretation_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ interpretation_0_coding_0_system:
+ description: High, low, normal, etc... Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ issued:
+ description: Date/Time this version was made available.
+ type:
+ - string
+ - 'null'
+ language:
+ description: Language of the resource content.
+ type:
+ - string
+ - 'null'
+ method_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ method_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ method_coding_0_system:
+ description: How it was done.. Code defined by a terminology system.. Identity
+ of the terminology system.
+ type:
+ - string
+ - 'null'
+ project_id:
+ $ref: _definitions.yaml#/project_id
+ referenceRange_0_age_high_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ referenceRange_0_age_high_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ referenceRange_0_age_high_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ referenceRange_0_age_high_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ referenceRange_0_age_high_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ referenceRange_0_age_low_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ referenceRange_0_age_low_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ referenceRange_0_age_low_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ referenceRange_0_age_low_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ referenceRange_0_age_low_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ referenceRange_0_appliesTo_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ referenceRange_0_appliesTo_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ referenceRange_0_appliesTo_0_coding_0_system:
+ description: Reference range population.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ referenceRange_0_high_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ referenceRange_0_high_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ referenceRange_0_high_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ referenceRange_0_high_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ referenceRange_0_high_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ referenceRange_0_low_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ referenceRange_0_low_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ referenceRange_0_low_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ referenceRange_0_low_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ referenceRange_0_low_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ referenceRange_0_text:
+ description: Text based reference range in an observation.
+ type:
+ - string
+ - 'null'
+ referenceRange_0_type_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ referenceRange_0_type_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ referenceRange_0_type_coding_0_system:
+ description: Codes to indicate the what part of the targeted reference population
+ it applies to. For example, the normal or therapeutic range.. Code defined by
+ a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ resource_type:
+ description: ''
+ type:
+ - string
+ - 'null'
+ state:
+ $ref: _definitions.yaml#/state
+ status:
+ description: The status of the result value.. http://hl7.org/fhir/observation-status
+ enum:
+ - registered
+ - preliminary
+ - final
+ - amended
+ - corrected
+ - cancelled
+ - entered-in-error
+ - unknown
+ term:
+ description: The status of the result value.. http://hl7.org/fhir/observation-status
+ termDef:
+ cde_id: http://hl7.org/fhir/observation-status
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/observation-status
+ term_url: http://hl7.org/fhir/observation-status
+ submitter_id:
+ type:
+ - string
+ - 'null'
+ updated_datetime:
+ $ref: _definitions.yaml#/datetime
+ valueBoolean:
+ description: Actual result.
+ type:
+ - boolean
+ - 'null'
+ valueCodeableConcept_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ valueCodeableConcept_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ valueCodeableConcept_coding_0_system:
+ description: Actual result.. Code defined by a terminology system.. Identity of
+ the terminology system.
+ type:
+ - string
+ - 'null'
+ valueDateTime:
+ description: Actual result.
+ type:
+ - string
+ - 'null'
+ valueInteger:
+ description: Actual result.
+ type:
+ - number
+ - 'null'
+ valuePeriod_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ valuePeriod_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ valueQuantity_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ valueQuantity_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ valueQuantity_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ valueQuantity_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ valueQuantity_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ valueRange_high_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ valueRange_high_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ valueRange_high_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ valueRange_high_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ valueRange_high_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ valueRange_low_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ valueRange_low_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ valueRange_low_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ valueRange_low_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ valueRange_low_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ valueRatio_denominator_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ valueRatio_denominator_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ valueRatio_denominator_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ valueRatio_denominator_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ valueRatio_denominator_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ valueRatio_numerator_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ valueRatio_numerator_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ valueRatio_numerator_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ valueRatio_numerator_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ valueRatio_numerator_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ valueSampledData_data:
+ description: Decimal values with spaces, or "E" | "U" | "L".
+ type:
+ - string
+ - 'null'
+ valueSampledData_dimensions:
+ description: Number of sample points at each time point.
+ type:
+ - number
+ valueSampledData_factor:
+ description: Multiply data by this before adding to origin.
+ type:
+ - number
+ - 'null'
+ valueSampledData_lowerLimit:
+ description: Lower limit of detection.
+ type:
+ - number
+ - 'null'
+ valueSampledData_origin_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ valueSampledData_origin_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ valueSampledData_origin_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ valueSampledData_origin_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ valueSampledData_origin_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ valueSampledData_period:
+ description: Number of milliseconds between samples.
+ type:
+ - number
+ valueSampledData_upperLimit:
+ description: Upper limit of detection.
+ type:
+ - number
+ - 'null'
+ valueString:
+ description: Actual result.
+ type:
+ - string
+ - 'null'
+ valueTime:
+ description: Actual result.
+ type:
+ - string
+ - 'null'
+required:
+- submitter_id
+- status
+- code
+submittable: true
+systemProperties:
+- id
+- project_id
+- created_datetime
+- updated_datetime
+- state
+title: Observation
+type: object
+uniqueKeys:
+- - id
+- - project_id
+ - submitter_id
+validators: null
diff --git a/datadictionary/gdcdictionary/schemas/organization.yaml b/datadictionary/gdcdictionary/schemas/organization.yaml
new file mode 100644
index 00000000..c0194bdc
--- /dev/null
+++ b/datadictionary/gdcdictionary/schemas/organization.yaml
@@ -0,0 +1,134 @@
+$schema: http://json-schema.org/draft-04/schema#
+additionalProperties: false
+category: Administrative
+description: " A grouping of people or organizations with a common purpose.\n \n\
+ \ A formally or informally recognized grouping of people or organizations\n \
+ \ formed for the purpose of achieving some form of collective action.\n Includes\
+ \ companies, institutions, corporations, departments, community\n groups, healthcare\
+ \ practice groups, payer/insurer, etc.\n http://hl7.org/fhir/organization"
+id: organization
+links: []
+namespace: http://hl7.org/fhir
+program: '*'
+project: '*'
+properties:
+ active:
+ description: Whether the organization's record is still in active use.
+ type:
+ - boolean
+ - 'null'
+ alias:
+ description: A list of alternate names that the organization is known as, or was
+ known as in the past.
+ type:
+ - string
+ - 'null'
+ contact_0_purpose_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ contact_0_purpose_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ contact_0_purpose_coding_0_system:
+ description: Indicates a purpose for which the contact can be reached.. Code defined
+ by a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ created_datetime:
+ $ref: _definitions.yaml#/datetime
+ id:
+ description: Logical id of this artifact.
+ type:
+ - string
+ - 'null'
+ identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ identifier_0_use:
+ description: Identifies this organization across multiple systems.. The purpose
+ of this identifier.. http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: Identifies this organization across multiple systems.. The purpose
+ of this identifier.. http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ language:
+ description: Language of the resource content.
+ type:
+ - string
+ - 'null'
+ name:
+ description: Name used for the organization.
+ type:
+ - string
+ - 'null'
+ project_id:
+ $ref: _definitions.yaml#/project_id
+ resource_type:
+ description: ''
+ type:
+ - string
+ - 'null'
+ state:
+ $ref: _definitions.yaml#/state
+ submitter_id:
+ type:
+ - string
+ - 'null'
+ type_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ type_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ type_0_coding_0_system:
+ description: The kind(s) of organization that this is.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ updated_datetime:
+ $ref: _definitions.yaml#/datetime
+required:
+- submitter_id
+submittable: true
+systemProperties:
+- id
+- project_id
+- created_datetime
+- updated_datetime
+- state
+title: Organization
+type: object
+uniqueKeys:
+- - id
+- - project_id
+ - submitter_id
+validators: null
diff --git a/datadictionary/gdcdictionary/schemas/patient.yaml b/datadictionary/gdcdictionary/schemas/patient.yaml
new file mode 100644
index 00000000..dca552c9
--- /dev/null
+++ b/datadictionary/gdcdictionary/schemas/patient.yaml
@@ -0,0 +1,306 @@
+$schema: http://json-schema.org/draft-04/schema#
+additionalProperties: false
+category: Administrative
+description: " Information about an individual or animal receiving health care services.\n\
+ \ \n Demographics and other administrative information about an individual\
+ \ or\n animal receiving care or other health-related services.\n http://hl7.org/fhir/patient"
+id: patient
+links:
+- backref: patients
+ label: Patient_generalPractitioner_practitioner
+ multiplicity: many_to_many
+ name: generalPractitioner_practitioner
+ required: false
+ target_type: practitioner
+- backref: patients
+ label: Patient_managingOrganization_organization
+ multiplicity: many_to_many
+ name: managingOrganization_organization
+ required: false
+ target_type: organization
+namespace: http://hl7.org/fhir
+program: '*'
+project: '*'
+properties:
+ active:
+ description: Whether this patient's record is in active use.
+ type:
+ - boolean
+ - 'null'
+ birthDate:
+ description: The date of birth for the individual.
+ type:
+ - string
+ - 'null'
+ communication_0_language_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ communication_0_language_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ communication_0_language_coding_0_system:
+ description: The language which can be used to communicate with the patient about
+ his or her health.. Code defined by a terminology system.. Identity of the terminology
+ system.
+ type:
+ - string
+ - 'null'
+ communication_0_preferred:
+ description: Language preference indicator.
+ type:
+ - boolean
+ - 'null'
+ contact_0_gender:
+ description: Administrative Gender - the gender that the contact person is considered
+ to have for administration and record keeping purposes.. http://hl7.org/fhir/administrative-gender
+ enum:
+ - male
+ - female
+ - other
+ - unknown
+ term:
+ description: Administrative Gender - the gender that the contact person is considered
+ to have for administration and record keeping purposes.. http://hl7.org/fhir/administrative-gender
+ termDef:
+ cde_id: http://hl7.org/fhir/administrative-gender
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/administrative-gender
+ term_url: http://hl7.org/fhir/administrative-gender
+ contact_0_organization_reference:
+ description: Organization that is associated with the contact.. Literal reference,
+ Relative, internal or absolute URL.
+ type:
+ - string
+ - 'null'
+ contact_0_organization_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ contact_0_period_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ contact_0_period_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ contact_0_relationship_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ contact_0_relationship_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ contact_0_relationship_0_coding_0_system:
+ description: The kind of relationship.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ created_datetime:
+ $ref: _definitions.yaml#/datetime
+ deceasedBoolean:
+ description: Indicates if the individual is deceased or not.
+ type:
+ - boolean
+ - 'null'
+ deceasedDateTime:
+ description: Indicates if the individual is deceased or not.
+ type:
+ - string
+ - 'null'
+ gender:
+ description: Administrative Gender - the gender that the patient is considered
+ to have for administration and record keeping purposes.. http://hl7.org/fhir/administrative-gender
+ enum:
+ - male
+ - female
+ - other
+ - unknown
+ term:
+ description: Administrative Gender - the gender that the patient is considered
+ to have for administration and record keeping purposes.. http://hl7.org/fhir/administrative-gender
+ termDef:
+ cde_id: http://hl7.org/fhir/administrative-gender
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/administrative-gender
+ term_url: http://hl7.org/fhir/administrative-gender
+ id:
+ description: Logical id of this artifact.
+ type:
+ - string
+ - 'null'
+ identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ identifier_0_use:
+ description: An identifier for this patient.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: An identifier for this patient.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ language:
+ description: Language of the resource content.
+ type:
+ - string
+ - 'null'
+ link_0_other_reference:
+ description: The other patient or related person resource that the link refers
+ to.. Literal reference, Relative, internal or absolute URL.
+ type:
+ - string
+ - 'null'
+ link_0_other_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ link_0_type:
+ description: The type of link between this patient resource and another patient
+ resource.. http://hl7.org/fhir/link-type
+ enum:
+ - replaced-by
+ - replaces
+ - refer
+ - seealso
+ term:
+ description: The type of link between this patient resource and another patient
+ resource.. http://hl7.org/fhir/link-type
+ termDef:
+ cde_id: http://hl7.org/fhir/link-type
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/link-type
+ term_url: http://hl7.org/fhir/link-type
+ maritalStatus_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ maritalStatus_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ maritalStatus_coding_0_system:
+ description: Marital (civil) status of a patient.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ multipleBirthBoolean:
+ description: Whether patient is part of a multiple birth.
+ type:
+ - boolean
+ - 'null'
+ multipleBirthInteger:
+ description: Whether patient is part of a multiple birth.
+ type:
+ - number
+ - 'null'
+ photo_0_contentType:
+ description: Mime type of the content, with charset etc..
+ type:
+ - string
+ - 'null'
+ photo_0_creation:
+ description: Date attachment was first created.
+ type:
+ - string
+ - 'null'
+ photo_0_data:
+ description: Data inline, base64ed.
+ type:
+ - string
+ - 'null'
+ photo_0_hash:
+ description: Hash of the data (sha-1, base64ed).
+ type:
+ - string
+ - 'null'
+ photo_0_language:
+ description: Human language of the content (BCP-47).
+ type:
+ - string
+ - 'null'
+ photo_0_size:
+ description: Number of bytes of content (if url provided).
+ type:
+ - number
+ - 'null'
+ photo_0_title:
+ description: Label to display in place of the data.
+ type:
+ - string
+ - 'null'
+ photo_0_url:
+ description: Uri where the data can be found.
+ type:
+ - string
+ - 'null'
+ project_id:
+ $ref: _definitions.yaml#/project_id
+ resource_type:
+ description: ''
+ type:
+ - string
+ - 'null'
+ state:
+ $ref: _definitions.yaml#/state
+ submitter_id:
+ type:
+ - string
+ - 'null'
+ updated_datetime:
+ $ref: _definitions.yaml#/datetime
+required:
+- submitter_id
+submittable: true
+systemProperties:
+- id
+- project_id
+- created_datetime
+- updated_datetime
+- state
+title: Patient
+type: object
+uniqueKeys:
+- - id
+- - project_id
+ - submitter_id
+validators: null
diff --git a/datadictionary/gdcdictionary/schemas/practitioner.yaml b/datadictionary/gdcdictionary/schemas/practitioner.yaml
new file mode 100644
index 00000000..ecc30f23
--- /dev/null
+++ b/datadictionary/gdcdictionary/schemas/practitioner.yaml
@@ -0,0 +1,234 @@
+$schema: http://json-schema.org/draft-04/schema#
+additionalProperties: false
+category: Administrative
+description: " A person with a formal responsibility in the provisioning of healthcare\
+ \ or\n related services.\n \n A person who is directly or indirectly involved\
+ \ in the provisioning of\n healthcare.\n http://hl7.org/fhir/practitioner"
+id: practitioner
+links: []
+namespace: http://hl7.org/fhir
+program: '*'
+project: '*'
+properties:
+ active:
+ description: Whether this practitioner's record is in active use.
+ type:
+ - boolean
+ - 'null'
+ birthDate:
+ description: The date on which the practitioner was born.
+ type:
+ - string
+ - 'null'
+ communication_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ communication_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ communication_0_coding_0_system:
+ description: A language the practitioner can use in patient communication.. Code
+ defined by a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ created_datetime:
+ $ref: _definitions.yaml#/datetime
+ gender:
+ description: Administrative Gender - the gender that the person is considered
+ to have for administration and record keeping purposes.. http://hl7.org/fhir/administrative-gender
+ enum:
+ - male
+ - female
+ - other
+ - unknown
+ term:
+ description: Administrative Gender - the gender that the person is considered
+ to have for administration and record keeping purposes.. http://hl7.org/fhir/administrative-gender
+ termDef:
+ cde_id: http://hl7.org/fhir/administrative-gender
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/administrative-gender
+ term_url: http://hl7.org/fhir/administrative-gender
+ id:
+ description: Logical id of this artifact.
+ type:
+ - string
+ - 'null'
+ identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ identifier_0_use:
+ description: An identifier for the person as this agent.. The purpose of this
+ identifier.. http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: An identifier for the person as this agent.. The purpose of this
+ identifier.. http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ language:
+ description: Language of the resource content.
+ type:
+ - string
+ - 'null'
+ photo_0_contentType:
+ description: Mime type of the content, with charset etc..
+ type:
+ - string
+ - 'null'
+ photo_0_creation:
+ description: Date attachment was first created.
+ type:
+ - string
+ - 'null'
+ photo_0_data:
+ description: Data inline, base64ed.
+ type:
+ - string
+ - 'null'
+ photo_0_hash:
+ description: Hash of the data (sha-1, base64ed).
+ type:
+ - string
+ - 'null'
+ photo_0_language:
+ description: Human language of the content (BCP-47).
+ type:
+ - string
+ - 'null'
+ photo_0_size:
+ description: Number of bytes of content (if url provided).
+ type:
+ - number
+ - 'null'
+ photo_0_title:
+ description: Label to display in place of the data.
+ type:
+ - string
+ - 'null'
+ photo_0_url:
+ description: Uri where the data can be found.
+ type:
+ - string
+ - 'null'
+ project_id:
+ $ref: _definitions.yaml#/project_id
+ qualification_0_code_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ qualification_0_code_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ qualification_0_code_coding_0_system:
+ description: Coded representation of the qualification.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ qualification_0_identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ qualification_0_identifier_0_use:
+ description: An identifier for this qualification for the practitioner.. The purpose
+ of this identifier.. http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: An identifier for this qualification for the practitioner.. The
+ purpose of this identifier.. http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ qualification_0_identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ qualification_0_issuer_reference:
+ description: Organization that regulates and issues the qualification.. Literal
+ reference, Relative, internal or absolute URL.
+ type:
+ - string
+ - 'null'
+ qualification_0_issuer_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ qualification_0_period_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ qualification_0_period_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ resource_type:
+ description: ''
+ type:
+ - string
+ - 'null'
+ state:
+ $ref: _definitions.yaml#/state
+ submitter_id:
+ type:
+ - string
+ - 'null'
+ updated_datetime:
+ $ref: _definitions.yaml#/datetime
+required:
+- submitter_id
+submittable: true
+systemProperties:
+- id
+- project_id
+- created_datetime
+- updated_datetime
+- state
+title: Practitioner
+type: object
+uniqueKeys:
+- - id
+- - project_id
+ - submitter_id
+validators: null
diff --git a/datadictionary/gdcdictionary/schemas/practitioner_role.yaml b/datadictionary/gdcdictionary/schemas/practitioner_role.yaml
new file mode 100644
index 00000000..7723c7f4
--- /dev/null
+++ b/datadictionary/gdcdictionary/schemas/practitioner_role.yaml
@@ -0,0 +1,198 @@
+$schema: http://json-schema.org/draft-04/schema#
+additionalProperties: false
+category: Administrative
+description: " Roles/organizations the practitioner is associated with.\n \n \
+ \ A specific set of Roles/Locations/specialties/services that a practitioner\n\
+ \ may perform at an organization for a period of time.\n http://hl7.org/fhir/practitionerrole"
+id: practitioner_role
+links:
+- backref: practitioner_roles
+ label: PractitionerRole_organization_organization
+ multiplicity: many_to_many
+ name: organization_organization
+ required: false
+ target_type: organization
+- backref: practitioner_roles
+ label: PractitionerRole_practitioner_practitioner
+ multiplicity: many_to_many
+ name: practitioner_practitioner
+ required: false
+ target_type: practitioner
+namespace: http://hl7.org/fhir
+program: '*'
+project: '*'
+properties:
+ active:
+ description: Whether this practitioner role record is in active use.
+ type:
+ - boolean
+ - 'null'
+ availabilityExceptions:
+ description: Description of availability exceptions.
+ type:
+ - string
+ - 'null'
+ availableTime_0_allDay:
+ description: Always available? e.g. 24 hour service.
+ type:
+ - boolean
+ - 'null'
+ availableTime_0_availableEndTime:
+ description: Closing time of day (ignored if allDay = true).
+ type:
+ - string
+ - 'null'
+ availableTime_0_availableStartTime:
+ description: Opening time of day (ignored if allDay = true).
+ type:
+ - string
+ - 'null'
+ availableTime_0_daysOfWeek:
+ description: Indicates which days of the week are available between the start
+ and end Times.. http://hl7.org/fhir/days-of-week
+ enum:
+ - mon
+ - tue
+ - wed
+ - thu
+ - fri
+ - sat
+ - sun
+ term:
+ description: Indicates which days of the week are available between the start
+ and end Times.. http://hl7.org/fhir/days-of-week
+ termDef:
+ cde_id: http://hl7.org/fhir/days-of-week
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/days-of-week
+ term_url: http://hl7.org/fhir/days-of-week
+ code_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ code_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ code_0_coding_0_system:
+ description: Roles which this practitioner may perform.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ created_datetime:
+ $ref: _definitions.yaml#/datetime
+ id:
+ description: Logical id of this artifact.
+ type:
+ - string
+ - 'null'
+ identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ identifier_0_use:
+ description: Business Identifiers that are specific to a role/location.. The purpose
+ of this identifier.. http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: Business Identifiers that are specific to a role/location.. The
+ purpose of this identifier.. http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ language:
+ description: Language of the resource content.
+ type:
+ - string
+ - 'null'
+ notAvailable_0_description:
+ description: Reason presented to the user explaining why time not available.
+ type:
+ - string
+ notAvailable_0_during_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ notAvailable_0_during_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ period_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ period_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ project_id:
+ $ref: _definitions.yaml#/project_id
+ resource_type:
+ description: ''
+ type:
+ - string
+ - 'null'
+ specialty_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ specialty_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ specialty_0_coding_0_system:
+ description: Specific specialty of the practitioner.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ state:
+ $ref: _definitions.yaml#/state
+ submitter_id:
+ type:
+ - string
+ - 'null'
+ updated_datetime:
+ $ref: _definitions.yaml#/datetime
+required:
+- submitter_id
+submittable: true
+systemProperties:
+- id
+- project_id
+- created_datetime
+- updated_datetime
+- state
+title: PractitionerRole
+type: object
+uniqueKeys:
+- - id
+- - project_id
+ - submitter_id
+validators: null
diff --git a/datadictionary/gdcdictionary/schemas/procedure.yaml b/datadictionary/gdcdictionary/schemas/procedure.yaml
new file mode 100644
index 00000000..5b7177e8
--- /dev/null
+++ b/datadictionary/gdcdictionary/schemas/procedure.yaml
@@ -0,0 +1,440 @@
+$schema: http://json-schema.org/draft-04/schema#
+additionalProperties: false
+category: Clinical
+description: " An action that is being or was performed on a patient.\n \n An\
+ \ action that is or was performed on or for a patient. This can be a\n physical\
+ \ intervention like an operation, or less invasive like long term\n services,\
+ \ counseling, or hypnotherapy.\n http://hl7.org/fhir/procedure"
+id: procedure
+links:
+- backref: procedures
+ label: Procedure_encounter_encounter
+ multiplicity: many_to_many
+ name: encounter_encounter
+ required: false
+ target_type: encounter
+- backref: procedures
+ label: Procedure_location_location
+ multiplicity: many_to_many
+ name: location_location
+ required: false
+ target_type: location
+namespace: http://hl7.org/fhir
+program: '*'
+project: '*'
+properties:
+ bodySite_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ bodySite_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ bodySite_0_coding_0_system:
+ description: Target body sites.. Code defined by a terminology system.. Identity
+ of the terminology system.
+ type:
+ - string
+ - 'null'
+ category_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ category_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ category_coding_0_system:
+ description: Classification of the procedure.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ code_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ code_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ code_coding_0_system:
+ description: Identification of the procedure.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ complication_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ complication_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ complication_0_coding_0_system:
+ description: Complication following the procedure.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ created_datetime:
+ $ref: _definitions.yaml#/datetime
+ focalDevice_0_action_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ focalDevice_0_action_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ focalDevice_0_action_coding_0_system:
+ description: Kind of change to device.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ focalDevice_0_manipulated_reference:
+ description: Device that was changed.. Literal reference, Relative, internal or
+ absolute URL.
+ type:
+ - string
+ - 'null'
+ focalDevice_0_manipulated_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ followUp_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ followUp_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ followUp_0_coding_0_system:
+ description: Instructions for follow up.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ id:
+ description: Logical id of this artifact.
+ type:
+ - string
+ - 'null'
+ identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ identifier_0_use:
+ description: External Identifiers for this procedure.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: External Identifiers for this procedure.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ instantiatesCanonical:
+ description: Instantiates FHIR protocol or definition.
+ type:
+ - string
+ - 'null'
+ instantiatesUri:
+ description: Instantiates external protocol or definition.
+ type:
+ - string
+ - 'null'
+ language:
+ description: Language of the resource content.
+ type:
+ - string
+ - 'null'
+ outcome_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ outcome_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ outcome_coding_0_system:
+ description: The result of procedure.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ performedDateTime:
+ description: When the procedure was performed.
+ type:
+ - string
+ - 'null'
+ performedPeriod_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ performedPeriod_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ performedRange_high_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ performedRange_high_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ performedRange_high_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ performedRange_high_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ performedRange_high_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ performedRange_low_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ performedRange_low_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ performedRange_low_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ performedRange_low_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ performedRange_low_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ performedString:
+ description: When the procedure was performed.
+ type:
+ - string
+ - 'null'
+ performer_0_actor_reference:
+ description: The reference to the practitioner.. Literal reference, Relative,
+ internal or absolute URL.
+ type:
+ - string
+ - 'null'
+ performer_0_actor_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ performer_0_function_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ performer_0_function_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ performer_0_function_coding_0_system:
+ description: Type of performance.. Code defined by a terminology system.. Identity
+ of the terminology system.
+ type:
+ - string
+ - 'null'
+ performer_0_onBehalfOf_reference:
+ description: Organization the device or practitioner was acting for.. Literal
+ reference, Relative, internal or absolute URL.
+ type:
+ - string
+ - 'null'
+ performer_0_onBehalfOf_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ project_id:
+ $ref: _definitions.yaml#/project_id
+ reasonCode_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ reasonCode_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ reasonCode_0_coding_0_system:
+ description: Coded reason procedure performed.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ resource_type:
+ description: ''
+ type:
+ - string
+ - 'null'
+ state:
+ $ref: _definitions.yaml#/state
+ status:
+ description: A code specifying the state of the procedure. Generally, this will
+ be the in-progress or completed state.. http://hl7.org/fhir/event-status
+ enum:
+ - preparation
+ - in-progress
+ - not-done
+ - on-hold
+ - stopped
+ - completed
+ - entered-in-error
+ - unknown
+ term:
+ description: A code specifying the state of the procedure. Generally, this will
+ be the in-progress or completed state.. http://hl7.org/fhir/event-status
+ termDef:
+ cde_id: http://hl7.org/fhir/event-status
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/event-status
+ term_url: http://hl7.org/fhir/event-status
+ statusReason_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ statusReason_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ statusReason_coding_0_system:
+ description: Reason for current status.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ submitter_id:
+ type:
+ - string
+ - 'null'
+ updated_datetime:
+ $ref: _definitions.yaml#/datetime
+ usedCode_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ usedCode_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ usedCode_0_coding_0_system:
+ description: Coded items used during the procedure.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+required:
+- submitter_id
+- status
+- subject
+submittable: true
+systemProperties:
+- id
+- project_id
+- created_datetime
+- updated_datetime
+- state
+title: Procedure
+type: object
+uniqueKeys:
+- - id
+- - project_id
+ - submitter_id
+validators: null
diff --git a/datadictionary/gdcdictionary/schemas/publication.yaml b/datadictionary/gdcdictionary/schemas/publication.yaml
deleted file mode 100644
index 2feb6a1a..00000000
--- a/datadictionary/gdcdictionary/schemas/publication.yaml
+++ /dev/null
@@ -1,62 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "publication"
-title: Publication
-type: object
-namespace: http://gdc.nci.nih.gov
-category: administrative
-program: '*'
-project: '*'
-description: "Publication for a project."
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - state
- - created_datetime
- - updated_datetime
-
-links:
- - name: projects
- backref: publications
- label: refers_to
- target_type: project
- multiplicity: many_to_many
- required: true
-
-required:
- - submitter_id
- - type
- - projects
-
-uniqueKeys:
- - [ id ]
- - [ project_id, submitter_id ]
-
-properties:
- type:
- enum: [ "publication" ]
- id:
- $ref: "_definitions.yaml#/UUID"
- systemAlias: node_id
- state:
- $ref: "_definitions.yaml#/state"
- submitter_id:
- type:
- - string
- - "null"
- pmid:
- type: string
- doi:
- type: string
- projects:
- $ref: "_definitions.yaml#/to_many_project"
- project_id:
- type: string
- created_datetime:
- $ref: "_definitions.yaml#/datetime"
- updated_datetime:
- $ref: "_definitions.yaml#/datetime"
diff --git a/datadictionary/gdcdictionary/schemas/read_group.yaml b/datadictionary/gdcdictionary/schemas/read_group.yaml
deleted file mode 100644
index d1a8bdd4..00000000
--- a/datadictionary/gdcdictionary/schemas/read_group.yaml
+++ /dev/null
@@ -1,225 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "read_group"
-title: Read Group
-type: object
-description: "Sequencing reads from one lane of an NGS experiment."
-namespace: http://gdc.nci.nih.gov
-category: biospecimen
-project: '*'
-program: '*'
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - created_datetime
- - updated_datetime
- - state
-
-links:
- - name: aliquots
- label: derived_from
- target_type: aliquot
- multiplicity: many_to_one
- required: true
- backref: read_groups
-
-required:
- - type
- - submitter_id
- - aliquots
-
-uniqueKeys:
- - [ id ]
- - [ project_id, submitter_id ]
-
-properties:
- id:
- $ref: "_definitions.yaml#/UUID"
- project_id:
- $ref: "_definitions.yaml#/project_id"
- submitter_id:
- type: string
- state:
- $ref: "_definitions.yaml#/state"
- type:
- enum: [ "read_group" ]
- experiment_name:
- term:
- $ref: "_terms.yaml#/experiment_name"
- type: string
- sequencing_center:
- term:
- $ref: "_terms.yaml#/sequencing_center"
- type: string
- sequencing_date:
- $ref: "_definitions.yaml#/datetime"
- platform:
- term:
- $ref: "_terms.yaml#/platform"
- enum:
- - Illumina
- - SOLiD
- - LS454
- - Ion Torrent
- - Complete Genomics
- - PacBio
- - Other
- instrument_model:
- terms:
- $ref: "_terms.yaml#/instrument_model"
- enum:
- - 454 GS FLX Titanium
- - AB SOLiD 4
- - AB SOLiD 2
- - AB SOLiD 3
- - Complete Genomics
- - Illumina HiSeq X Ten
- - Illumina HiSeq X Five
- - Illumina Genome Analyzer II
- - Illumina Genome Analyzer IIx
- - Illumina HiSeq 2000
- - Illumina HiSeq 2500
- - Illumina HiSeq 4000
- - Illumina MiSeq
- - Illumina NextSeq
- - Ion Torrent PGM
- - Ion Torrent Proton
- - PacBio RS
- - Ion S5 XL System, Ion 530 Chip
- - Other
- library_strategy:
- term:
- $ref: "_terms.yaml#/library_strategy"
- enum:
- - WGS
- - WXS
- - RNA-Seq
- - ChIP-Seq
- - miRNA-Seq
- - Bisulfite-Seq
- - Validation
- - Amplicon
- - Other
- RIN:
- term:
- $ref: "_terms.yaml#/RIN"
- type: number
- flow_cell_barcode:
- term:
- $ref: "_terms.yaml#/flow_cell_barcode"
- type: string
- includes_spike_ins:
- term:
- $ref: "_terms.yaml#/includes_spike_ins"
- type: boolean
- spike_ins_fasta:
- term:
- $ref: "_terms.yaml#/spike_ins_fasta"
- type: string
- spike_ins_concentration:
- term:
- $ref: "_terms.yaml#/spike_ins_concentration"
- type: string
- library_selection:
- term:
- $ref: "_terms.yaml#/library_selection"
- enum:
- - Hybrid_Selection
- - PCR
- - Affinity_Enrichment
- - Poly-T_Enrichment
- - RNA_Depletion
- - Other
- library_preparation_kit_name:
- term:
- $ref: "_terms.yaml#/library_preparation_kit_name"
- type: string
- library_preparation_kit_vendor:
- term:
- $ref: "_terms.yaml#/library_preparation_kit_vendor"
- type: string
- library_preparation_kit_catalog_number:
- term:
- $ref: "_terms.yaml#/library_preparation_kit_catalog_number"
- type: string
- library_preparation_kit_version:
- term:
- $ref: "_terms.yaml#/library_preparation_kit_version"
- type: string
- library_name:
- term:
- $ref: "_terms.yaml#/library_name"
- type: string
- target_capture_kit_name: # conditionally required for WXS etc
- term:
- $ref: "_terms.yaml#/target_capture_kit_name"
- type: string
- target_capture_kit_vendor:
- term:
- $ref: "_terms.yaml#/target_capture_kit_vendor"
- type: string
- target_capture_kit_catalog_number:
- term:
- $ref: "_terms.yaml#/target_capture_kit_catalog_number"
- type: string
- target_capture_kit_version:
- term:
- $ref: "_terms.yaml#/target_capture_kit_version"
- type: string
- target_capture_kit_target_region:
- term:
- $ref: "_terms.yaml#/target_capture_kit_target_region"
- type: string
- size_selection_range:
- term:
- $ref: "_terms.yaml#/size_selection_range"
- type: string
- adapter_name:
- term:
- $ref: "_terms.yaml#/adapter_name"
- type: string
- adapter_sequence:
- term:
- $ref: "_terms.yaml#/adapter_sequence"
- type: string
- to_trim_adapter_sequence:
- term:
- $ref: "_terms.yaml#/to_trim_adapter_sequence"
- type: boolean
- library_strand:
- term:
- $ref: "_terms.yaml#/library_strand"
- enum:
- - Unstranded
- - First_Stranded
- - Second_Stranded
- base_caller_name:
- term:
- $ref: "_terms.yaml#/base_caller_name"
- type: string
- base_caller_version:
- term:
- $ref: "_terms.yaml#/base_caller_version"
- type: string
- is_paired_end:
- term:
- $ref: "_terms.yaml#/is_paired_end"
- type: boolean
- read_length:
- type: integer
- read_group_name: # it may be good to assign UUID to read group
- description: "Read Group Name"
- type: string
- barcoding_applied:
- description: "True/False: was barcoding applied?"
- type: boolean
- aliquots:
- $ref: "_definitions.yaml#/to_one"
- created_datetime:
- $ref: "_definitions.yaml#/datetime"
- updated_datetime:
- $ref: "_definitions.yaml#/datetime"
diff --git a/datadictionary/gdcdictionary/schemas/read_group_qc.yaml b/datadictionary/gdcdictionary/schemas/read_group_qc.yaml
deleted file mode 100644
index 9b235528..00000000
--- a/datadictionary/gdcdictionary/schemas/read_group_qc.yaml
+++ /dev/null
@@ -1,133 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "read_group_qc"
-title: Read Group QC
-type: object
-namespace: http://gdc.nci.nih.gov
-category: notation
-project: '*'
-program: '*'
-description: "GDC QC run metadata."
-additionalProperties: false
-submittable: false
-validators: null
-
-systemProperties:
- - id
- - project_id
- - created_datetime
- - updated_datetime
- - state
-
-links:
- - exclusive: true
- required: true
- subgroup:
- - name: submitted_aligned_reads_files
- backref: read_group_qcs
- label: data_from
- target_type: submitted_aligned_reads
- multiplicity: one_to_one
- required: false
- - name: submitted_unaligned_reads_files
- backref: read_group_qcs
- label: data_from
- target_type: submitted_unaligned_reads
- multiplicity: one_to_many
- required: false
- - name: read_groups
- label: generated_from
- target_type: read_group
- multiplicity: many_to_one
- required: true
- backref: read_group_qcs
-
-required:
- - submitter_id
- - workflow_link
- - type
- - percent_gc_content
- - encoding
- - total_sequences
- - basic_statistics
- - per_base_sequence_quality
- - per_tile_sequence_quality
- - per_sequence_quality_score
- - per_base_sequence_content
- - per_sequence_gc_content
- - per_base_n_content
- - sequence_length_distribution
- - sequence_duplication_levels
- - overrepresented_sequences
- - adapter_content
- - kmer_content
- - read_groups
-
-uniqueKeys:
- - [ id ]
- - [ project_id, submitter_id ]
-
-properties:
- $ref: "_definitions.yaml#/workflow_properties"
- type:
- enum: [ "read_group_qc" ]
- workflow_type:
- term:
- $ref: "_terms.yaml#/workflow_type"
- enum: [ "Read Group Quality Control" ]
- fastq_name:
- term:
- $ref: "_terms.yaml#/file_name"
- type: string
- percent_aligned:
- description: "The percent of reads with at least one reported alignment."
- type: integer
- minimum: 0
- maximum: 100
- percent_gc_content:
- term:
- $ref: "_terms.yaml#/percent_gc_content"
- type: integer
- minimum: 0
- maximum: 100
- encoding:
- term:
- $ref: "_terms.yaml#/encoding"
- type: string
- total_aligned_reads:
- description: "The total number of reads with at least one reported alignment."
- type: integer
- total_sequences:
- term:
- $ref: "_terms.yaml#/total_sequences"
- type: integer
- basic_statistics:
- $ref: "_definitions.yaml#/qc_metrics_state"
- per_base_sequence_quality:
- $ref: "_definitions.yaml#/qc_metrics_state"
- per_tile_sequence_quality:
- $ref: "_definitions.yaml#/qc_metrics_state"
- per_sequence_quality_score:
- $ref: "_definitions.yaml#/qc_metrics_state"
- per_base_sequence_content:
- $ref: "_definitions.yaml#/qc_metrics_state"
- per_sequence_gc_content:
- $ref: "_definitions.yaml#/qc_metrics_state"
- per_base_n_content:
- $ref: "_definitions.yaml#/qc_metrics_state"
- sequence_length_distribution:
- $ref: "_definitions.yaml#/qc_metrics_state"
- sequence_duplication_levels:
- $ref: "_definitions.yaml#/qc_metrics_state"
- overrepresented_sequences:
- $ref: "_definitions.yaml#/qc_metrics_state"
- adapter_content:
- $ref: "_definitions.yaml#/qc_metrics_state"
- kmer_content:
- $ref: "_definitions.yaml#/qc_metrics_state"
- submitted_aligned_reads_files:
- $ref: "_definitions.yaml#/to_one"
- submitted_unaligned_reads_files:
- $ref: "_definitions.yaml#/to_many"
- read_groups:
- $ref: "_definitions.yaml#/to_one"
diff --git a/datadictionary/gdcdictionary/schemas/research_study.yaml b/datadictionary/gdcdictionary/schemas/research_study.yaml
new file mode 100644
index 00000000..d85b7713
--- /dev/null
+++ b/datadictionary/gdcdictionary/schemas/research_study.yaml
@@ -0,0 +1,405 @@
+$schema: http://json-schema.org/draft-04/schema#
+additionalProperties: false
+category: Administrative
+description: " Investigation to increase healthcare-related patient-independent knowledge.\n\
+ \ \n A process where a researcher or organization plans and then executes\
+ \ a\n series of steps intended to increase the field of healthcare-related\n\
+ \ knowledge. This includes studies of safety, efficacy, comparative\n effectiveness\
+ \ and other information about medications, devices, therapies\n and other interventional\
+ \ and investigative techniques. A ResearchStudy\n involves the gathering of\
+ \ information about human or animal subjects.\n http://hl7.org/fhir/researchstudy"
+id: research_study
+links:
+- backref: research_studies
+ label: ResearchStudy_project_project
+ multiplicity: many_to_many
+ name: project_project
+ required: true
+ target_type: project
+namespace: http://hl7.org/fhir
+program: '*'
+project: '*'
+properties:
+ arm_0_description:
+ description: Short explanation of study path.
+ type:
+ - string
+ - 'null'
+ arm_0_name:
+ description: Label for study arm.
+ type:
+ - string
+ arm_0_type_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ arm_0_type_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ arm_0_type_coding_0_system:
+ description: Categorization of study arm.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ category_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ category_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ category_0_coding_0_system:
+ description: Classifications for the study.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ condition_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ condition_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ condition_0_coding_0_system:
+ description: Condition being studied.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ contact_0_name:
+ description: Name of an individual to contact.
+ type:
+ - string
+ - 'null'
+ created_datetime:
+ $ref: _definitions.yaml#/datetime
+ description:
+ description: What this is study doing.
+ type:
+ - string
+ - 'null'
+ focus_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ focus_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ focus_0_coding_0_system:
+ description: Drugs, devices, etc. under study.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ id:
+ description: Logical id of this artifact.
+ type:
+ - string
+ - 'null'
+ identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ identifier_0_use:
+ description: Business Identifier for study.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: Business Identifier for study.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ keyword_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ keyword_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ keyword_0_coding_0_system:
+ description: Used to search for the study.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ language:
+ description: Language of the resource content.
+ type:
+ - string
+ - 'null'
+ location_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ location_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ location_0_coding_0_system:
+ description: Geographic region(s) for study.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ objective_0_name:
+ description: Label for the objective.
+ type:
+ - string
+ - 'null'
+ objective_0_type_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ objective_0_type_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ objective_0_type_coding_0_system:
+ description: The kind of study objective.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ period_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ period_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ phase_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ phase_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ phase_coding_0_system:
+ description: The stage in the progression of a therapy from initial experimental
+ use in humans in clinical trials to post-market evaluation.. Code defined by
+ a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ primaryPurposeType_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ primaryPurposeType_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ primaryPurposeType_coding_0_system:
+ description: The type of study based upon the intent of the study's activities.
+ A classification of the intent of the study.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ project_id:
+ $ref: _definitions.yaml#/project_id
+ reasonStopped_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ reasonStopped_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ reasonStopped_coding_0_system:
+ description: A description and/or code explaining the premature termination of
+ the study.. Code defined by a terminology system.. Identity of the terminology
+ system.
+ type:
+ - string
+ - 'null'
+ relatedArtifact_0_citation:
+ description: Bibliographic citation for the artifact.
+ type:
+ - string
+ - 'null'
+ relatedArtifact_0_display:
+ description: Brief description of the related artifact.
+ type:
+ - string
+ - 'null'
+ relatedArtifact_0_document_contentType:
+ description: Mime type of the content, with charset etc..
+ type:
+ - string
+ - 'null'
+ relatedArtifact_0_document_creation:
+ description: Date attachment was first created.
+ type:
+ - string
+ - 'null'
+ relatedArtifact_0_document_data:
+ description: Data inline, base64ed.
+ type:
+ - string
+ - 'null'
+ relatedArtifact_0_document_hash:
+ description: Hash of the data (sha-1, base64ed).
+ type:
+ - string
+ - 'null'
+ relatedArtifact_0_document_language:
+ description: Human language of the content (BCP-47).
+ type:
+ - string
+ - 'null'
+ relatedArtifact_0_document_size:
+ description: Number of bytes of content (if url provided).
+ type:
+ - number
+ - 'null'
+ relatedArtifact_0_document_title:
+ description: Label to display in place of the data.
+ type:
+ - string
+ - 'null'
+ relatedArtifact_0_document_url:
+ description: Uri where the data can be found.
+ type:
+ - string
+ - 'null'
+ relatedArtifact_0_label:
+ description: Short label.
+ type:
+ - string
+ - 'null'
+ relatedArtifact_0_resource:
+ description: What resource is being referenced.
+ type:
+ - string
+ - 'null'
+ relatedArtifact_0_type:
+ description: The type of relationship to the related artifact.. http://hl7.org/fhir/related-artifact-type
+ enum:
+ - documentation
+ - justification
+ - citation
+ - predecessor
+ - successor
+ - derived-from
+ - depends-on
+ - composed-of
+ term:
+ description: The type of relationship to the related artifact.. http://hl7.org/fhir/related-artifact-type
+ termDef:
+ cde_id: http://hl7.org/fhir/related-artifact-type
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/related-artifact-type
+ term_url: http://hl7.org/fhir/related-artifact-type
+ relatedArtifact_0_url:
+ description: Where the artifact can be accessed.
+ type:
+ - string
+ - 'null'
+ resource_type:
+ description: ''
+ type:
+ - string
+ - 'null'
+ state:
+ $ref: _definitions.yaml#/state
+ status:
+ description: The current state of the study.. http://hl7.org/fhir/research-study-status
+ enum:
+ - active
+ - administratively-completed
+ - approved
+ - closed-to-accrual
+ - closed-to-accrual-and-intervention
+ - completed
+ - disapproved
+ - in-review
+ - temporarily-closed-to-accrual
+ - temporarily-closed-to-accrual-and-intervention
+ - withdrawn
+ term:
+ description: The current state of the study.. http://hl7.org/fhir/research-study-status
+ termDef:
+ cde_id: http://hl7.org/fhir/research-study-status
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/research-study-status
+ term_url: http://hl7.org/fhir/research-study-status
+ submitter_id:
+ type:
+ - string
+ - 'null'
+ title:
+ description: Name for this study.
+ type:
+ - string
+ - 'null'
+ updated_datetime:
+ $ref: _definitions.yaml#/datetime
+required:
+- submitter_id
+- status
+submittable: true
+systemProperties:
+- id
+- project_id
+- created_datetime
+- updated_datetime
+- state
+title: ResearchStudy
+type: object
+uniqueKeys:
+- - id
+- - project_id
+ - submitter_id
+validators: null
diff --git a/datadictionary/gdcdictionary/schemas/research_subject.yaml b/datadictionary/gdcdictionary/schemas/research_subject.yaml
new file mode 100644
index 00000000..a434643a
--- /dev/null
+++ b/datadictionary/gdcdictionary/schemas/research_subject.yaml
@@ -0,0 +1,144 @@
+$schema: http://json-schema.org/draft-04/schema#
+additionalProperties: false
+category: Administrative
+description: " Physical entity which is the primary unit of interest in the study.\n\
+ \ \n A physical entity which is the primary unit of operational and/or\n \
+ \ administrative interest in a study.\n http://hl7.org/fhir/researchsubject"
+id: research_subject
+links:
+- backref: research_subjects
+ label: ResearchSubject_individual_patient
+ multiplicity: many_to_many
+ name: individual_patient
+ required: false
+ target_type: patient
+- backref: research_subjects
+ label: ResearchSubject_study_research_study
+ multiplicity: many_to_many
+ name: study_research_study
+ required: false
+ target_type: research_study
+namespace: http://hl7.org/fhir
+program: '*'
+project: '*'
+properties:
+ actualArm:
+ description: What path was followed.
+ type:
+ - string
+ - 'null'
+ assignedArm:
+ description: What path should be followed.
+ type:
+ - string
+ - 'null'
+ created_datetime:
+ $ref: _definitions.yaml#/datetime
+ id:
+ description: Logical id of this artifact.
+ type:
+ - string
+ - 'null'
+ identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ identifier_0_use:
+ description: Business Identifier for research subject in a study.. The purpose
+ of this identifier.. http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: Business Identifier for research subject in a study.. The purpose
+ of this identifier.. http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ language:
+ description: Language of the resource content.
+ type:
+ - string
+ - 'null'
+ period_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ period_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ project_id:
+ $ref: _definitions.yaml#/project_id
+ resource_type:
+ description: ''
+ type:
+ - string
+ - 'null'
+ state:
+ $ref: _definitions.yaml#/state
+ status:
+ description: The current state of the subject.. http://hl7.org/fhir/research-subject-status
+ enum:
+ - candidate
+ - eligible
+ - follow-up
+ - ineligible
+ - not-registered
+ - off-study
+ - on-study
+ - on-study-intervention
+ - on-study-observation
+ - pending-on-study
+ - potential-candidate
+ - screening
+ - withdrawn
+ term:
+ description: The current state of the subject.. http://hl7.org/fhir/research-subject-status
+ termDef:
+ cde_id: http://hl7.org/fhir/research-subject-status
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/research-subject-status
+ term_url: http://hl7.org/fhir/research-subject-status
+ submitter_id:
+ type:
+ - string
+ - 'null'
+ updated_datetime:
+ $ref: _definitions.yaml#/datetime
+required:
+- submitter_id
+- status
+- study
+- individual
+submittable: true
+systemProperties:
+- id
+- project_id
+- created_datetime
+- updated_datetime
+- state
+title: ResearchSubject
+type: object
+uniqueKeys:
+- - id
+- - project_id
+ - submitter_id
+validators: null
diff --git a/datadictionary/gdcdictionary/schemas/sample.yaml b/datadictionary/gdcdictionary/schemas/sample.yaml
deleted file mode 100644
index 7680e5cb..00000000
--- a/datadictionary/gdcdictionary/schemas/sample.yaml
+++ /dev/null
@@ -1,686 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "sample"
-title: Sample
-type: object
-namespace: http://gdc.nci.nih.gov
-category: biospecimen
-program: '*'
-project: '*'
-description: >
- Any material sample taken from a biological entity for testing, diagnostic, propagation, treatment
- or research purposes, including a sample obtained from a living organism or taken from the
- biological object after halting of all its life functions. Biospecimen can contain one or more
- components including but not limited to cellular molecules, cells, tissues, organs, body fluids,
- embryos, and body excretory products.
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - state
- - created_datetime
- - updated_datetime
-
-required:
- - submitter_id
- - type
- - cases
-
-uniqueKeys:
- - [id]
- - [project_id, submitter_id]
-
-links:
- - name: cases
- backref: samples
- label: derived_from
- target_type: case
- multiplicity: many_to_one #not sure
- required: true
- - name: diagnoses
- backref: samples
- label: related_to
- target_type: diagnosis
- multiplicity: many_to_one
- required: false
-
-# Sample properties: keep TCGA-specific fields
-properties:
- type:
- type: string
- id:
- $ref: "_definitions.yaml#/UUID"
- systemAlias: node_id
- state:
- $ref: "_definitions.yaml#/state"
- submitter_id:
- type:
- - string
- - "null"
- description: >
- The legacy barcode used before prior to the use UUIDs, varies by project.
- For TCGA this is bcrsamplebarcode.
- biospecimen_anatomic_site:
- term:
- $ref: "_terms.yaml#/biospecimen_anatomic_site"
- enum:
- - Abdomen
- - Abdominal Wall
- - Acetabulum
- - Adenoid
- - Adipose
- - Adrenal
- - Alveolar Ridge
- - Amniotic Fluid
- - Ampulla Of Vater
- - Anal Sphincter
- - Ankle
- - Anorectum
- - Antecubital Fossa
- - Antrum
- - Anus
- - Aorta
- - Aortic Body
- - Appendix
- - Aqueous Fluid
- - Arm
- - Artery
- - Ascending Colon
- - Ascending Colon Hepatic Flexure
- - Auditory Canal
- - Autonomic Nervous System
- - Axilla
- - Back
- - Bile Duct
- - Bladder
- - Blood
- - Blood Vessel
- - Bone
- - Bone Marrow
- - Bowel
- - Brain
- - Brain Stem
- - Breast
- - Broad Ligament
- - Bronchiole
- - Bronchus
- - Brow
- - Buccal Cavity
- - Buccal Mucosa
- - Buttock
- - Calf
- - Capillary
- - Cardia
- - Carina
- - Carotid Artery
- - Carotid Body
- - Cartilage
- - Cecum
- - Cell-Line
- - Central Nervous System
- - Cerebellum
- - Cerebral Cortex
- - Cerebrospinal Fluid
- - Cerebrum
- - Cervical Spine
- - Cervix
- - Chest
- - Chest Wall
- - Chin
- - Clavicle
- - Clitoris
- - Colon
- - Colon - Mucosa Only
- - Common Duct
- - Conjunctiva
- - Connective Tissue
- - Dermal
- - Descending Colon
- - Diaphragm
- - Duodenum
- - Ear
- - Ear Canal
- - Ear, Pinna (External)
- - Effusion
- - Elbow
- - Endocrine Gland
- - Epididymis
- - Epidural Space
- - Esophagogastric Junction
- - Esophagus
- - Esophagus - Mucosa Only
- - Eye
- - Fallopian Tube
- - Femoral Artery
- - Femoral Vein
- - Femur
- - Fibroblasts
- - Fibula
- - Finger
- - Floor Of Mouth
- - Fluid
- - Foot
- - Forearm
- - Forehead
- - Foreskin
- - Frontal Cortex
- - Frontal Lobe
- - Fundus Of Stomach
- - Gallbladder
- - Ganglia
- - Gastroesophageal Junction
- - Gastrointestinal Tract
- - Groin
- - Gum
- - Hand
- - Hard Palate
- - Head & Neck
- - Head - Face Or Neck, Nos
- - Heart
- - Hepatic
- - Hepatic Duct
- - Hepatic Vein
- - Hip
- - Hippocampus
- - Humerus
- - Hypopharynx
- - Ileum
- - Ilium
- - Index Finger
- - Ischium
- - Islet Cells
- - Jaw
- - Jejunum
- - Joint
- - Kidney
- - Knee
- - Lacrimal Gland
- - Large Bowel
- - Laryngopharynx
- - Larynx
- - Leg
- - Leptomeninges
- - Ligament
- - Lip
- - Liver
- - Lumbar Spine
- - Lung
- - Lymph Node
- - Lymph Node(s) Axilla
- - Lymph Node(s) Cervical
- - Lymph Node(s) Distant
- - Lymph Node(s) Epitrochlear
- - Lymph Node(s) Femoral
- - Lymph Node(s) Hilar
- - Lymph Node(s) Iliac-Common
- - Lymph Node(s) Iliac-External
- - Lymph Node(s) Inguinal
- - Lymph Node(s) Internal Mammary
- - Lymph Node(s) Mammary
- - Lymph Node(s) Mesenteric
- - Lymph Node(s) Occipital
- - Lymph Node(s) Paraaortic
- - Lymph Node(s) Parotid
- - Lymph Node(s) Pelvic
- - Lymph Node(s) Popliteal
- - Lymph Node(s) Regional
- - Lymph Node(s) Retroperitoneal
- - Lymph Node(s) Scalene
- - Lymph Node(s) Splenic
- - Lymph Node(s) Subclavicular
- - Lymph Node(s) Submandibular
- - Lymph Node(s) Supraclavicular
- - Lymph Nodes(s) Mediastinal
- - Mandible
- - Maxilla
- - Mediastinal Soft Tissue
- - Mediastinum
- - Mesentery
- - Mesothelium
- - Middle Finger
- - Mitochondria
- - Muscle
- - Nails
- - Nasal Cavity
- - Nasal Soft Tissue
- - Nasopharynx
- - Neck
- - Nerve
- - Nerve(s) Cranial
- - Occipital Cortex
- - Ocular Orbits
- - Omentum
- - Oral Cavity
- - Oral Cavity - Mucosa Only
- - Oropharynx
- - Other
- - Ovary
- - Palate
- - Pancreas
- - Paraspinal Ganglion
- - Parathyroid
- - Parotid Gland
- - Patella
- - Pelvis
- - Penis
- - Pericardium
- - Periorbital Soft Tissue
- - Peritoneal Cavity
- - Peritoneum
- - Pharynx
- - Pineal
- - Pineal Gland
- - Pituitary Gland
- - Placenta
- - Pleura
- - Popliteal Fossa
- - Prostate
- - Pylorus
- - Rectosigmoid Junction
- - Rectum
- - Retina
- - Retro-Orbital Region
- - Retroperitoneum
- - Rib
- - Ring Finger
- - Round Ligament
- - Sacrum
- - Salivary Gland
- - Scalp
- - Scapula
- - Sciatic Nerve
- - Scrotum
- - Seminal Vesicle
- - Shoulder
- - Sigmoid Colon
- - Sinus
- - Sinus(es), Maxillary
- - Skeletal Muscle
- - Skin
- - Skull
- - Small Bowel
- - Small Bowel - Mucosa Only
- - Small Finger
- - Soft Tissue
- - Spinal Column
- - Spinal Cord
- - Spleen
- - Splenic Flexure
- - Sternum
- - Stomach
- - Stomach - Mucosa Only
- - Subcutaneous Tissue
- - Synovium
- - Temporal Cortex
- - Tendon
- - Testis
- - Thigh
- - Thoracic Spine
- - Thorax
- - Throat
- - Thumb
- - Thymus
- - Thyroid
- - Tibia
- - Tongue
- - Tonsil
- - Tonsil (Pharyngeal)
- - Trachea / Major Bronchi
- - Transverse Colon
- - Trunk
- - Umbilical Cord
- - Ureter
- - Urethra
- - Urinary Tract
- - Uterus
- - Uvula
- - Vagina
- - Vas Deferens
- - Vein
- - Venous
- - Vertebra
- - Vulva
- - White Blood Cells
- - Wrist
- - Unknown
- - Not Reported
- - Not Allowed To Collect
- composition:
- term:
- $ref: "_terms.yaml#/composition"
- enum:
- - Buccal Cells
- - Buffy Coat
- - Bone Marrow Components
- - Bone Marrow Components NOS
- - Control Analyte
- - Cell
- - Circulating Tumor Cell (CTC)
- - Derived Cell Line
- - EBV Immortalized
- - Fibroblasts from Bone Marrow Normal
- - Granulocytes
- - Human Original Cells
- - Lymphocytes
- - Mononuclear Cells from Bone Marrow Normal
- - Peripheral Blood Components NOS
- - Peripheral Blood Nucleated Cells
- - Pleural Effusion
- - Plasma
- - Peripheral Whole Blood
- - Serum
- - Saliva
- - Sputum
- - Solid Tissue
- - Whole Bone Marrow
- - Unknown
- - Not Reported
- - Not Allowed To Collect
- current_weight:
- term:
- $ref: "_terms.yaml#/current_weight"
- type: number
- days_to_collection:
- term:
- $ref: "_terms.yaml#/days_to_collection"
- type: integer
- days_to_sample_procurement:
- term:
- $ref: "_terms.yaml#/days_to_sample_procurement"
- type: integer
- diagnosis_pathologically_confirmed:
- term:
- ref: "_terms.yaml#/diagnosis_pathologically_confirmed"
- enum:
- - "Yes"
- - "No"
- - Unknown
- freezing_method:
- term:
- $ref: "_terms.yaml#/freezing_method"
- type: string
- initial_weight:
- term:
- $ref: "_terms.yaml#/initial_weight"
- type: number
- intermediate_dimension:
- terms:
- $ref: "_terms.yaml#/intermediate_dimension"
- type: string
- is_ffpe:
- term:
- $ref: "_terms.yaml#/is_ffpe"
- type: boolean
- longest_dimension:
- terms:
- $ref: "_terms.yaml#/longest_dimension"
- type: string
- method_of_sample_procurement:
- term:
- $ref: "_terms.yaml#/method_of_sample_procurement"
- enum:
- - Abdomino-perineal Resection of Rectum
- - Anterior Resection of Rectum
- - Aspirate
- - Biopsy
- - Blood Draw
- - Bone Marrow Aspirate
- - Core Biopsy
- - Cystectomy
- - Endo Rectal Tumor Resection
- - Endoscopic Biopsy
- - Endoscopic Mucosal Resection (EMR)
- - Enucleation
- - Excisional Biopsy
- - Fine Needle Aspiration
- - Full Hysterectomy
- - Gross Total Resection
- - Hand Assisted Laparoscopic Radical Nephrectomy
- - Hysterectomy NOS
- - Incisional Biopsy
- - Indeterminant
- - Laparoscopic Biopsy
- - Laparoscopic Partial Nephrectomy
- - Laparoscopic Radical Nephrectomy
- - Laparoscopic Radical Prostatectomy with Robotics
- - Laparoscopic Radical Prostatectomy without Robotics
- - Left Hemicolectomy
- - Lobectomy
- - Local Resection (Exoresection; wall resection)
- - Lumpectomy
- - Modified Radical Mastectomy
- - Needle Biopsy
- - Open Craniotomy
- - Open Partial Nephrectomy
- - Open Radical Nephrectomy
- - Open Radical Prostatectomy
- - Orchiectomy
- - Other
- - Other Surgical Resection
- - Pan-Procto Colectomy
- - Pneumonectomy
- - Right Hemicolectomy
- - Sigmoid Colectomy
- - Simple Mastectomy
- - Subtotal Resection
- - Surgical Resection
- - Thoracoscopic Biopsy
- - Total Colectomy
- - Total Mastectomy
- - Transplant
- - Transurethral resection (TURBT)
- - Transverse Colectomy
- - Tumor Resection
- - Wedge Resection
- - Unknown
- - Not Reported
- - Not Allowed To Collect
- oct_embedded:
- term:
- $ref: "_terms.yaml#/oct_embedded"
- type: string
- preservation_method:
- term:
- $ref: "_terms.yaml#/preservation_method"
- enum:
- - Cryopreserved
- - FFPE
- - Fresh
- - OCT
- - Snap Frozen
- - Frozen
- - Unknown
- - Not Reported
- - Not Allowed To Collect
- sample_type:
- description: "Characterization of the sample as either clinical or contrived."
- enum:
- - Additional Metastatic
- - Additional - New Primary
- - Blood Derived Cancer - Bone Marrow, Post-treatment
- - Blood Derived Cancer - Peripheral Blood, Post-treatment
- - Blood Derived Normal
- - Bone Marrow Normal
- - Buccal Cell Normal
- - Cell Line Derived Xenograft Tissue
- - Cell Lines
- - cfDNA
- - Circulating Tumor Cell (CTC)
- - Control Analyte
- - Clinical
- - Contrived
- - ctDNA
- - DNA
- - EBV Immortalized Normal
- - FFPE Recurrent
- - FFPE Scrolls
- - Fibroblasts from Bone Marrow Normal
- - GenomePlex (Rubicon) Amplified DNA
- - Granulocytes
- - Human Tumor Original Cells
- - Metastatic
- - Mononuclear Cells from Bone Marrow Normal
- - Primary Blood Derived Cancer - Peripheral Blood
- - Recurrent Blood Derived Cancer - Peripheral Blood
- - Pleural Effusion
- - Primary Blood Derived Cancer - Bone Marrow
- - Primary Tumor
- - Primary Xenograft Tissue
- - Post neo-adjuvant therapy
- - Recurrent Blood Derived Cancer - Bone Marrow
- - Recurrent Tumor
- - Repli-G (Qiagen) DNA
- - Repli-G X (Qiagen) DNA
- - RNA
- - Slides
- - Solid Tissue Normal
- - Total RNA
- - Xenograft Tissue
- - Unknown
- - Not Reported
- - Not Allowed To Collect
- sample_type_id:
- term:
- $ref: "_terms.yaml#/sample_type_id"
- enum:
- - '01'
- - '02'
- - '03'
- - '04'
- - '05'
- - '06'
- - '07'
- - '08'
- - '09'
- - '10'
- - '11'
- - '12'
- - '13'
- - '14'
- - '15'
- - '16'
- - '20'
- - '40'
- - '41'
- - '42'
- - '50'
- - '60'
- - '61'
- - '99'
- sample_volume:
- description: "The volume of the sample in mL."
- type: number
- shortest_dimension:
- term:
- $ref: "_terms.yaml#/shortest_dimension"
- type: string
- time_between_clamping_and_freezing:
- term:
- $ref: "_terms.yaml#/time_between_clamping_and_freezing"
- type: string
- time_between_excision_and_freezing:
- term:
- $ref: "_terms.yaml#/time_between_excision_and_freezing"
- type: string
- tissue_type:
- term:
- $ref: "_terms.yaml#/tissue_type"
- enum:
- - Tumor
- - Normal
- - Abnormal
- - Peritumoral
- - Contrived
- - Unknown
- - Not Reported
- - Not Allowed To Collect
- tumor_code:
- term:
- $ref: "_terms.yaml#/tumor_code"
- enum:
- - Non cancerous tissue
- - Diffuse Large B-Cell Lymphoma (DLBCL)
- - Lung Cancer (all types)
- - Lung Adenocarcinoma
- - Non-small Cell Lung Carcinoma (NSCLC)
- - Colon Cancer (all types)
- - Breast Cancer (all types)
- - Cervical Cancer (all types)
- - Anal Cancer (all types)
- - Acute lymphoblastic leukemia (ALL)
- - Acute myeloid leukemia (AML)
- - Induction Failure AML (AML-IF)
- - Neuroblastoma (NBL)
- - Osteosarcoma (OS)
- - Ewing sarcoma
- - Wilms tumor (WT)
- - Clear cell sarcoma of the kidney (CCSK)
- - Rhabdoid tumor (kidney) (RT)
- - CNS, ependymoma
- - CNS, glioblastoma (GBM)
- - CNS, rhabdoid tumor
- - CNS, low grade glioma (LGG)
- - CNS, medulloblastoma
- - CNS, other
- - NHL, anaplastic large cell lymphoma
- - NHL, Burkitt lymphoma (BL)
- - Rhabdomyosarcoma
- - Soft tissue sarcoma, non-rhabdomyosarcoma
- - Castration-Resistant Prostate Cancer (CRPC)
- - Prostate Cancer
- - Hepatocellular Carcinoma (HCC)
- tumor_code_id:
- term:
- $ref: "_terms.yaml#/tumor_code_id"
- enum:
- - "00"
- - "01"
- - "02"
- - "03"
- - "04"
- - "10"
- - "20"
- - "21"
- - "30"
- - "40"
- - "41"
- - "50"
- - "51"
- - "52"
- - "60"
- - "61"
- - "62"
- - "63"
- - "64"
- - "65"
- - "70"
- - "71"
- - "80"
- - "81"
- tumor_descriptor:
- term:
- $ref: "_terms.yaml#/tumor_descriptor"
- enum:
- - Metastatic
- - Not Applicable
- - Primary
- - Recurrence
- - Xenograft
- - NOS
- - Unknown
- - Not Reported
- - Not Allowed To Collect
- description: "A description of the tumor from which the sample was derived." # TOREVIEW
- cases:
- $ref: "_definitions.yaml#/to_one"
- diagnoses:
- $ref: "_definitions.yaml#/to_one"
- project_id:
- type: string
- created_datetime:
- $ref: "_definitions.yaml#/datetime"
- updated_datetime:
- $ref: "_definitions.yaml#/datetime"
diff --git a/datadictionary/gdcdictionary/schemas/slide.yaml b/datadictionary/gdcdictionary/schemas/slide.yaml
deleted file mode 100644
index 2812c2d0..00000000
--- a/datadictionary/gdcdictionary/schemas/slide.yaml
+++ /dev/null
@@ -1,138 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "slide"
-title: Slide
-type: object
-namespace: http://gdc.nci.nih.gov
-category: biospecimen
-program: '*'
-project: '*'
-description: >
- A digital image, microscopic or otherwise, of any sample, portion, or sub-part thereof. (GDC)
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - state
- - created_datetime
- - updated_datetime
-
-links:
- - name: samples
- backref: slides
- label: derived_from
- target_type: sample
- multiplicity: many_to_many
- required: true
-
-required:
- - submitter_id
- - type
- - samples
-
-uniqueKeys:
- - [id]
- - [project_id, submitter_id]
-
-# slide properties
-properties:
- type:
- type: string
- id:
- $ref: "_definitions.yaml#/UUID"
- systemAlias: node_id
- state:
- $ref: "_definitions.yaml#/state"
- submitter_id:
- type:
- - string
- - "null"
- apoptotic_concentration:
- description: "The concentration, in cells/mL, of apoptotic cells in the slide blood."
- type: number
- ctc_concentration:
- description: "The concentration, in cells/mL, of traditional CTC cells (intact and enlarged cell and nucleus, cytokeratin positive, and CD45 negative) in the slide blood."
- type: number
- ctc_low_concentration:
- description: "The concentration, in cells/mL, of CTC-low cells (those with low cytokeratin levels compared to traditional CTCs) in the slide blood."
- type: number
- ctc_small_concentration:
- description: "The concentration, in cells/mL, of CTC-small cells (those with a small nuclear and cellular size relative to traditional CTCs) in the slide blood."
- type: number
- section_location:
- term:
- $ref: "_terms.yaml#/section_location"
- type: string
- methanol_added:
- description: "True/False indicator for if methanol was used in the slide preparation process."
- type: boolean
- number_proliferating_cells:
- term:
- $ref: "_terms.yaml#/number_proliferating_cells"
- type: integer
- number_nucleated_cells:
- description: "The total number of nucleated cells identified on the slide."
- type: integer
- percent_tumor_cells:
- term:
- $ref: "_terms.yaml#/percent_tumor_cells"
- type: number
- percent_tumor_nuclei:
- term:
- $ref: "_terms.yaml#/percent_tumor_nuclei"
- type: number
- percent_normal_cells:
- term:
- $ref: "_terms.yaml#/percent_normal_cells"
- type: number
- percent_necrosis:
- term:
- $ref: "_terms.yaml#/percent_necrosis"
- type: number
- percent_stromal_cells:
- term:
- $ref: "_terms.yaml#/percent_stromal_cells"
- type: number
- percent_inflam_infiltration:
- term:
- $ref: "_terms.yaml#/percent_inflam_infiltration"
- type: number
- percent_lymphocyte_infiltration:
- term:
- $ref: "_terms.yaml#/percent_lymphocyte_infiltration"
- type: number
- percent_monocyte_infiltration:
- term:
- $ref: "_terms.yaml#/percent_monocyte_infiltration"
- type: number
- percent_granulocyte_infiltration:
- term:
- $ref: "_terms.yaml#/percent_granulocyte_infiltration"
- type: number
- percent_neutrophil_infiltration:
- term:
- $ref: "_terms.yaml#/percent_neutrophil_infiltration"
- type: number
- percent_eosinophil_infiltration:
- term:
- $ref: "_terms.yaml#/percent_eosinophil_infiltration"
- type: number
- run_datetime:
- $ref: "_definitions.yaml#/datetime"
- run_name:
- description: "Name, number, or other identifier given to this slide's run."
- type: string
- slide_identifier:
- description: "Unique identifier given to the this slide."
- type: string
- samples:
- $ref: "_definitions.yaml#/to_many"
- project_id:
- $ref: "_definitions.yaml#/project_id"
- created_datetime:
- $ref: "_definitions.yaml#/datetime"
- updated_datetime:
- $ref: "_definitions.yaml#/datetime"
diff --git a/datadictionary/gdcdictionary/schemas/slide_count.yaml b/datadictionary/gdcdictionary/schemas/slide_count.yaml
deleted file mode 100644
index 30002260..00000000
--- a/datadictionary/gdcdictionary/schemas/slide_count.yaml
+++ /dev/null
@@ -1,100 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "slide_count"
-title: Slide Count
-type: object
-namespace: http://gdc.nci.nih.gov
-category: notation
-program: '*'
-project: '*'
-description: >
- Information pertaining to processed results obtained from slides; often in the form of counts.
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - created_datetime
- - updated_datetime
- - state
-
-links:
- - name: slides
- backref: slide_counts
- label: data_from
- target_type: slide
- multiplicity: many_to_many
- required: true
-
-required:
- - submitter_id
- - type
- - slides
-
-uniqueKeys:
- - [ id ]
- - [ project_id, submitter_id ]
-
-properties:
- type:
- enum: [ "slide_count" ]
- id:
- $ref: "_definitions.yaml#/UUID"
- systemAlias: node_id
- state:
- $ref: "_definitions.yaml#/state"
- submitter_id:
- type:
- - string
- - "null"
- cell_type:
- description: "The type of cell being counted or measured."
- type: string
- cell_identifier:
- description: "An alternative identifier for a given cell type."
- type: string
- cell_count:
- description: "Raw count of a particular cell type."
- type: integer
- ck_signal:
- description: "Numeric quantification of the CK signal."
- type: number
- biomarker_signal:
- description: "Numeric quantification of the biomarker signal."
- type: number
- er_localization:
- description: "Cellular localization of the endoplasmic reticulum as determined by staining."
- enum:
- - Nuclear
- - Cytoplasmic
- - Both
- - None
- - Not Determined
- frame_identifier:
- description: "Name, number, or other identifier given to the frame of the slide from which this image was taken."
- type: string
- relative_nuclear_size:
- description: "The ratio of the single cell's nucleus size to the average of the surrounding cells."
- type: number
- relative_nuclear_intensity:
- description: "The ratio of the single cell's nuclear staining intensity to the average of the surrounding cells."
- type: number
- relative_cytokeratin_intensity:
- description: "The ratio of the single cell's cytokeratin staining intensity to the average of the surrounding cells."
- type: number
- relative_er_intensity:
- description: "The ratio of the single cell's endoplasmic reticulum staining intensity to the average of the surrounding cells."
- type: number
- run_name:
- description: "The name or identifier given to the run that was used to generate this slide count."
- type: string
- slides:
- $ref: "_definitions.yaml#/to_many"
- project_id:
- type: string
- created_datetime:
- $ref: "_definitions.yaml#/datetime"
- updated_datetime:
- $ref: "_definitions.yaml#/datetime"
diff --git a/datadictionary/gdcdictionary/schemas/slide_image.yaml b/datadictionary/gdcdictionary/schemas/slide_image.yaml
deleted file mode 100644
index 50681f71..00000000
--- a/datadictionary/gdcdictionary/schemas/slide_image.yaml
+++ /dev/null
@@ -1,109 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "slide_image"
-title: Slide Image
-type: object
-namespace: http://gdc.nci.nih.gov
-category: data_file
-program: '*'
-project: '*'
-description: >
- Data file containing image of a slide.
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - created_datetime
- - updated_datetime
- - state
- - file_state
- - error_type
-
-links:
- - exclusive: false
- required: true
- subgroup:
- - name: slides
- backref: slide_images
- label: data_from
- target_type: slide
- multiplicity: many_to_one
- required: false
- - name: core_metadata_collections
- backref: slide_images
- label: data_from
- target_type: core_metadata_collection
- multiplicity: many_to_many
- required: false
-
-required:
- - submitter_id
- - type
- - file_name
- - file_size
- - md5sum
- - data_category
- - data_type
- - data_format
-
-uniqueKeys:
- - [ id ]
- - [ project_id, submitter_id ]
-
-properties:
- $ref: "_definitions.yaml#/data_file_properties"
- type:
- enum: [ "slide_image" ]
- data_category:
- term:
- $ref: "_terms.yaml#/data_category"
- enum:
- - Biospecimen
- - Slide Image
- - Mass Cytometry
- data_type:
- term:
- $ref: "_terms.yaml#/data_type"
- enum:
- - image
- - Single Cell Image
- - Raw IMC Data
- - Single Channel IMC Image
- - Antibody Panel Added
- data_format:
- term:
- $ref: "_terms.yaml#/data_format"
- type: string
- experimental_strategy:
- description: "Classification of the slide type with respect to its experimental use."
- enum:
- - Diagnostic Slide
- - Tissue Slide
- cell_type:
- description: "The type of cell being imaged or otherwised analysed."
- type: string
- cell_identifier:
- description: "An alternative identifier for a given cell type."
- type: string
- cell_count:
- description: "Count of the cell type being imaged or otherwise analysed."
- type: integer
- frame_identifier:
- description: "Name, number, or other identifier given to the frame of the slide from which this image was taken."
- type: string
- panel_used:
- description: "Name or other identifier given to the panel used during an IMC run."
- type: string
- protocol_used:
- description: "Name or other identifier given to the protocol used during an IMC run."
- type: string
- run_name:
- description: "Name, number, or other identifier given to the run that generated this slide image."
- type: string
- slides:
- $ref: "_definitions.yaml#/to_one"
- core_metadata_collections:
- $ref: "_definitions.yaml#/to_many"
diff --git a/datadictionary/gdcdictionary/schemas/specimen.yaml b/datadictionary/gdcdictionary/schemas/specimen.yaml
new file mode 100644
index 00000000..4e953bdf
--- /dev/null
+++ b/datadictionary/gdcdictionary/schemas/specimen.yaml
@@ -0,0 +1,545 @@
+$schema: http://json-schema.org/draft-04/schema#
+additionalProperties: false
+category: Biospecimen
+description: " Sample for analysis.\n \n A sample to be used for analysis.\n\
+ \ http://hl7.org/fhir/specimen"
+id: specimen
+links:
+- backref: specimen
+ label: Specimen_parent_specimen
+ multiplicity: many_to_many
+ name: parent_specimen
+ required: false
+ target_type: specimen
+- backref: specimen
+ label: Specimen_subject_patient
+ multiplicity: many_to_many
+ name: subject_patient
+ required: false
+ target_type: patient
+namespace: http://hl7.org/fhir
+program: '*'
+project: '*'
+properties:
+ accessionIdentifier_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ accessionIdentifier_use:
+ description: Identifier assigned by the lab.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: Identifier assigned by the lab.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ accessionIdentifier_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ collection_bodySite_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ collection_bodySite_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ collection_bodySite_coding_0_system:
+ description: Anatomical collection site.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ collection_collectedDateTime:
+ description: Collection time.
+ type:
+ - string
+ - 'null'
+ collection_collectedPeriod_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ collection_collectedPeriod_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ collection_collector_reference:
+ description: Who collected the specimen.. Literal reference, Relative, internal
+ or absolute URL.
+ type:
+ - string
+ - 'null'
+ collection_collector_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ collection_duration_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ collection_duration_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.
+ type:
+ - string
+ - 'null'
+ collection_duration_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ collection_duration_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ collection_duration_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ collection_fastingStatusCodeableConcept_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ collection_fastingStatusCodeableConcept_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ collection_fastingStatusCodeableConcept_coding_0_system:
+ description: Whether or how long patient abstained from food and/or drink.. Code
+ defined by a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ collection_fastingStatusDuration_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ collection_fastingStatusDuration_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.
+ type:
+ - string
+ - 'null'
+ collection_fastingStatusDuration_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ collection_fastingStatusDuration_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ collection_fastingStatusDuration_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ collection_method_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ collection_method_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ collection_method_coding_0_system:
+ description: Technique used to perform collection.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ collection_quantity_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ collection_quantity_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ collection_quantity_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ collection_quantity_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ collection_quantity_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ condition_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ condition_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ condition_0_coding_0_system:
+ description: State of the specimen.. Code defined by a terminology system.. Identity
+ of the terminology system.
+ type:
+ - string
+ - 'null'
+ container_0_additiveCodeableConcept_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ container_0_additiveCodeableConcept_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ container_0_additiveCodeableConcept_coding_0_system:
+ description: Additive associated with container.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ container_0_additiveReference_reference:
+ description: Additive associated with container.. Literal reference, Relative,
+ internal or absolute URL.
+ type:
+ - string
+ - 'null'
+ container_0_additiveReference_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ container_0_capacity_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ container_0_capacity_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ container_0_capacity_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ container_0_capacity_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ container_0_capacity_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ container_0_description:
+ description: Textual description of the container.
+ type:
+ - string
+ - 'null'
+ container_0_identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ container_0_identifier_0_use:
+ description: Id for the container.. The purpose of this identifier.. http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: Id for the container.. The purpose of this identifier.. http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ container_0_identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ container_0_specimenQuantity_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ container_0_specimenQuantity_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ container_0_specimenQuantity_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ container_0_specimenQuantity_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ container_0_specimenQuantity_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ container_0_type_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ container_0_type_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ container_0_type_coding_0_system:
+ description: Kind of container directly associated with specimen.. Code defined
+ by a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ created_datetime:
+ $ref: _definitions.yaml#/datetime
+ id:
+ description: Logical id of this artifact.
+ type:
+ - string
+ - 'null'
+ identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ identifier_0_use:
+ description: External Identifier.. The purpose of this identifier.. http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: External Identifier.. The purpose of this identifier.. http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ language:
+ description: Language of the resource content.
+ type:
+ - string
+ - 'null'
+ processing_0_additive_0_reference:
+ description: Material used in the processing step.. Literal reference, Relative,
+ internal or absolute URL.
+ type:
+ - string
+ - 'null'
+ processing_0_additive_0_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ processing_0_description:
+ description: Textual description of procedure.
+ type:
+ - string
+ - 'null'
+ processing_0_procedure_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ processing_0_procedure_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ processing_0_procedure_coding_0_system:
+ description: Indicates the treatment step applied to the specimen.. Code defined
+ by a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ processing_0_timeDateTime:
+ description: Date and time of specimen processing.
+ type:
+ - string
+ - 'null'
+ processing_0_timePeriod_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ processing_0_timePeriod_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ project_id:
+ $ref: _definitions.yaml#/project_id
+ receivedTime:
+ description: The time when specimen was received for processing.
+ type:
+ - string
+ - 'null'
+ resource_type:
+ description: ''
+ type:
+ - string
+ - 'null'
+ state:
+ $ref: _definitions.yaml#/state
+ status:
+ description: The availability of the specimen.. http://hl7.org/fhir/specimen-status
+ enum:
+ - available
+ - unavailable
+ - unsatisfactory
+ - entered-in-error
+ term:
+ description: The availability of the specimen.. http://hl7.org/fhir/specimen-status
+ termDef:
+ cde_id: http://hl7.org/fhir/specimen-status
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/specimen-status
+ term_url: http://hl7.org/fhir/specimen-status
+ submitter_id:
+ type:
+ - string
+ - 'null'
+ type_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ type_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ type_coding_0_system:
+ description: Kind of material that forms the specimen.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ updated_datetime:
+ $ref: _definitions.yaml#/datetime
+required:
+- submitter_id
+submittable: true
+systemProperties:
+- id
+- project_id
+- created_datetime
+- updated_datetime
+- state
+title: Specimen
+type: object
+uniqueKeys:
+- - id
+- - project_id
+ - submitter_id
+validators: null
diff --git a/datadictionary/gdcdictionary/schemas/submitted_aligned_reads.yaml b/datadictionary/gdcdictionary/schemas/submitted_aligned_reads.yaml
deleted file mode 100644
index 59db76ba..00000000
--- a/datadictionary/gdcdictionary/schemas/submitted_aligned_reads.yaml
+++ /dev/null
@@ -1,95 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "submitted_aligned_reads"
-title: Submitted Aligned Reads
-type: object
-namespace: http://gdc.nci.nih.gov
-category: data_file
-program: '*'
-project: '*'
-description: >
- Data file containing aligned reads that are used as input to GDC workflows.
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - created_datetime
- - updated_datetime
- - state
- - file_state
- - error_type
-
-links:
- - exclusive: false
- required: true
- subgroup:
- - name: read_groups
- backref: submitted_aligned_reads_files # pretty ugly
- label: data_from
- target_type: read_group
- multiplicity: one_to_many
- required: false
- - name: core_metadata_collections
- backref: submitted_aligned_reads_files
- label: data_from
- target_type: core_metadata_collection
- multiplicity: many_to_many
- required: false
-
-required:
- - submitter_id
- - type
- - file_name
- - file_size
- - data_format
- - md5sum
- - data_category
- - data_type
- - experimental_strategy
-
-uniqueKeys:
- - [ id ]
- - [ project_id, submitter_id ]
-
-properties:
- $ref: "_definitions.yaml#/data_file_properties"
- type:
- enum: [ "submitted_aligned_reads" ]
- data_category:
- term:
- $ref: "_terms.yaml#/data_category"
- enum:
- - Sequencing Data
- - Sequencing Reads
- - Raw Sequencing Data
- data_type:
- term:
- $ref: "_terms.yaml#/data_type"
- enum:
- - Aligned Reads
- - Alignment Coordinates
- data_format:
- term:
- $ref: "_terms.yaml#/data_format"
- enum:
- - BAM
- - BED
- experimental_strategy:
- term:
- $ref: "_terms.yaml#/experimental_strategy"
- enum:
- - WGS
- - WXS
- - Low Pass WGS
- - Validation
- - RNA-Seq
- - miRNA-Seq
- - Total RNA-Seq
- - DNA Panel
- read_groups:
- $ref: "_definitions.yaml#/to_many"
- core_metadata_collections:
- $ref: "_definitions.yaml#/to_many"
diff --git a/datadictionary/gdcdictionary/schemas/submitted_copy_number.yaml b/datadictionary/gdcdictionary/schemas/submitted_copy_number.yaml
deleted file mode 100644
index 2bc9d772..00000000
--- a/datadictionary/gdcdictionary/schemas/submitted_copy_number.yaml
+++ /dev/null
@@ -1,91 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "submitted_copy_number"
-title: Submitted Copy Number
-type: object
-namespace: http://gdc.nci.nih.gov
-category: data_file
-program: '*'
-project: '*'
-description: >
- Data file containing normalized copy number information from an aliquot.
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - created_datetime
- - updated_datetime
- - state
- - file_state
- - error_type
-
-links:
- - exclusive: false
- required: true
- subgroup:
- - name: core_metadata_collections
- backref: submitted_copy_number_files
- label: data_from
- target_type: core_metadata_collection
- multiplicity: many_to_many
- required: false
- - exclusive: true
- required: false
- subgroup:
- - name: aliquots
- backref: submitted_copy_number_files
- label: derived_from
- target_type: aliquot
- multiplicity: one_to_one
- required: false
- - name: read_groups
- backref: submitted_copy_number_files
- label: derived_from
- target_type: read_group
- multiplicity: many_to_many
- required: false
-
-required:
- - submitter_id
- - type
- - file_name
- - file_size
- - data_format
- - md5sum
- - data_category
- - data_type
- - experimental_strategy
-
-uniqueKeys:
- - [ id ]
- - [ project_id, submitter_id ]
-
-properties:
- $ref: "_definitions.yaml#/data_file_properties"
- type:
- enum: [ "submitted_copy_number" ]
- data_category:
- term:
- $ref: "_terms.yaml#/data_category"
- type: string
- data_type:
- term:
- $ref: "_terms.yaml#/data_type"
- type: string
- data_format:
- term:
- $ref: "_terms.yaml#/data_format"
- type: string
- experimental_strategy:
- term:
- $ref: "_terms.yaml#/experimental_strategy"
- type: string
- aliquots:
- $ref: "_definitions.yaml#/to_one"
- read_groups:
- $ref: "_definitions.yaml#/to_many"
- core_metadata_collections:
- $ref: "_definitions.yaml#/to_many"
diff --git a/datadictionary/gdcdictionary/schemas/submitted_methylation.yaml b/datadictionary/gdcdictionary/schemas/submitted_methylation.yaml
deleted file mode 100644
index 10408f5f..00000000
--- a/datadictionary/gdcdictionary/schemas/submitted_methylation.yaml
+++ /dev/null
@@ -1,86 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "submitted_methylation"
-title: Submitted Methylation
-type: object
-namespace: https://www.bloodpac.org/
-category: data_file
-program: '*'
-project: '*'
-description: "DNA methylation data files contain information on raw and normalized signal intensities, detection confidence and calculated beta values for methylated and unmethylated probes. DNA methylation is an epigenetic mark which can be associated with transcriptional inactivity when located in promoter regions."
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - created_datetime
- - updated_datetime
- - state
- - file_state
- - error_type
-
-links:
- - exclusive: false
- required: true
- subgroup:
- - name: core_metadata_collections
- backref: submitted_methylation_files
- label: data_from
- target_type: core_metadata_collection
- multiplicity: many_to_many
- required: false
- - name: aliquots
- backref: submitted_methylation_files
- label: data_from
- target_type: aliquot
- multiplicity: many_to_one
- required: false
-
-required:
- - submitter_id
- - type
- - file_name
- - file_size
- - md5sum
- - data_category
- - data_type
- - data_format
-
-uniqueKeys:
- - [ id ]
- - [ project_id, submitter_id ]
-
-properties:
- $ref: "_definitions.yaml#/data_file_properties"
- type:
- enum: [ "submitted_methylation" ]
- data_category:
- term:
- $ref: "_terms.yaml#/data_category"
- enum:
- - Methylation Data
- data_type:
- term:
- $ref: "_terms.yaml#/data_type"
- enum: [ "Methylation Intensity Values" ]
- data_format:
- term:
- $ref: "_terms.yaml#/data_format"
- enum:
- - IDAT
- assay_method:
- enum:
- - Methylation Array
- assay_instrument:
- enum:
- - Illumina
- assay_instrument_model:
- enum:
- - Illumina Infinium HumanMethylation450
- - Illumina Infinium HumanMethylation450K
- aliquots:
- $ref: "_definitions.yaml#/to_one"
- core_metadata_collections:
- $ref: "_definitions.yaml#/to_many"
diff --git a/datadictionary/gdcdictionary/schemas/submitted_somatic_mutation.yaml b/datadictionary/gdcdictionary/schemas/submitted_somatic_mutation.yaml
deleted file mode 100644
index f51b502c..00000000
--- a/datadictionary/gdcdictionary/schemas/submitted_somatic_mutation.yaml
+++ /dev/null
@@ -1,83 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "submitted_somatic_mutation"
-title: Submitted Somatic Mutation
-type: object
-namespace: http://gdc.nci.nih.gov
-category: data_file
-program: '*'
-project: '*'
-description: >
- Data file containing somatic mutation calls from a read group.
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - created_datetime
- - updated_datetime
- - state
- - file_state
- - error_type
-
-links:
- - exclusive: false
- required: true
- subgroup:
- - name: core_metadata_collections
- backref: submitted_somatic_mutations
- label: data_from
- target_type: core_metadata_collection
- multiplicity: many_to_many
- required: false
- - name: read_groups
- backref: submitted_somatic_mutations
- label: derived_from
- target_type: read_group
- multiplicity: many_to_many
- required: false
-
-required:
- - submitter_id
- - type
- - file_name
- - file_size
- - data_format
- - md5sum
- - data_category
- - data_type
- - experimental_strategy
-
-uniqueKeys:
- - [ id ]
- - [ project_id, submitter_id ]
-
-properties:
- $ref: "_definitions.yaml#/data_file_properties"
- type:
- enum: [ "submitted_somatic_mutation" ]
- data_category:
- term:
- $ref: "_terms.yaml#/data_category"
- type: string
- data_type:
- term:
- $ref: "_terms.yaml#/data_type"
- type: string
- data_format:
- term:
- $ref: "_terms.yaml#/data_format"
- type: string
- experimental_strategy:
- term:
- $ref: "_terms.yaml#/experimental_strategy"
- type: string
- total_variants:
- description: "The total number of variants detected carrying a base change difference from the reference genome."
- type: integer
- read_groups:
- $ref: "_definitions.yaml#/to_many"
- core_metadata_collections:
- $ref: "_definitions.yaml#/to_many"
diff --git a/datadictionary/gdcdictionary/schemas/submitted_unaligned_reads.yaml b/datadictionary/gdcdictionary/schemas/submitted_unaligned_reads.yaml
deleted file mode 100644
index 4b72d465..00000000
--- a/datadictionary/gdcdictionary/schemas/submitted_unaligned_reads.yaml
+++ /dev/null
@@ -1,92 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "submitted_unaligned_reads"
-title: Submitted Unaligned Reads
-type: object
-namespace: http://gdc.nci.nih.gov
-category: data_file
-program: '*'
-project: '*'
-description: "Data file containing unaligned reads that have not been GDC Harmonized."
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - created_datetime
- - updated_datetime
- - state
- - file_state
- - error_type
-
-links:
- - exclusive: false
- required: true
- subgroup:
- - name: read_groups
- backref: submitted_unaligned_reads_files # pretty ugly
- label: data_from
- target_type: read_group
- multiplicity: many_to_one
- required: false
- - name: core_metadata_collections
- backref: submitted_unaligned_reads_files
- label: data_from
- target_type: core_metadata_collection
- multiplicity: many_to_many
- required: false
-
-required:
- - submitter_id
- - type
- - file_name
- - file_size
- - md5sum
- - data_category
- - data_type
- - data_format
- - experimental_strategy
-
-uniqueKeys:
- - [ id ]
- - [ project_id, submitter_id ]
-
-properties:
- $ref: "_definitions.yaml#/data_file_properties"
- type:
- enum: [ "submitted_unaligned_reads" ]
- data_category:
- term:
- $ref: "_terms.yaml#/data_category"
- enum:
- - Sequencing Data
- - Sequencing Reads
- - Raw Sequencing Data
- data_type:
- term:
- $ref: "_terms.yaml#/data_type"
- enum: [ "Unaligned Reads" ]
- data_format:
- term:
- $ref: "_terms.yaml#/data_format"
- enum:
- - BAM
- - FASTQ
- experimental_strategy:
- term:
- $ref: "_terms.yaml#/experimental_strategy"
- enum:
- - WGS
- - WXS
- - Low Pass WGS
- - Validation
- - RNA-Seq
- - miRNA-Seq
- - Total RNA-Seq
- - DNA Panel
- read_groups:
- $ref: "_definitions.yaml#/to_one"
- core_metadata_collections:
- $ref: "_definitions.yaml#/to_many"
diff --git a/datadictionary/gdcdictionary/schemas/task.yaml b/datadictionary/gdcdictionary/schemas/task.yaml
new file mode 100644
index 00000000..41535936
--- /dev/null
+++ b/datadictionary/gdcdictionary/schemas/task.yaml
@@ -0,0 +1,1635 @@
+$schema: http://json-schema.org/draft-04/schema#
+additionalProperties: false
+category: Analysis
+description: " A task to be performed.\n http://hl7.org/fhir/task"
+id: task
+links:
+- backref: tasks
+ label: Task_focus_specimen
+ multiplicity: many_to_many
+ name: focus_specimen
+ required: false
+ target_type: specimen
+- backref: tasks
+ label: Task_for_patient
+ multiplicity: many_to_many
+ name: for_patient
+ required: true
+ target_type: patient
+- backref: tasks
+ label: Task_output_document_reference
+ multiplicity: many_to_many
+ name: output_document_reference
+ required: true
+ target_type: document_reference
+namespace: http://hl7.org/fhir
+program: '*'
+project: '*'
+properties:
+ authoredOn:
+ description: Task Creation Date.
+ type:
+ - string
+ - 'null'
+ businessStatus_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ businessStatus_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ businessStatus_coding_0_system:
+ description: E.g. "Specimen collected", "IV prepped".. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ code_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ code_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ code_coding_0_system:
+ description: A name or code (or both) briefly describing what the task involves..
+ Code defined by a terminology system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ created_datetime:
+ $ref: _definitions.yaml#/datetime
+ description:
+ description: Human-readable explanation of task.
+ type:
+ - string
+ - 'null'
+ executionPeriod_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ executionPeriod_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ groupIdentifier_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ groupIdentifier_use:
+ description: Requisition or grouper id.. The purpose of this identifier.. http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: Requisition or grouper id.. The purpose of this identifier.. http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ groupIdentifier_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ id:
+ description: Logical id of this artifact.
+ type:
+ - string
+ - 'null'
+ identifier_0_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ identifier_0_use:
+ description: Task Instance Identifier.. The purpose of this identifier.. http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: Task Instance Identifier.. The purpose of this identifier.. http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ identifier_0_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ input_0_type_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ input_0_type_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ input_0_type_coding_0_system:
+ description: Label for the input.. Code defined by a terminology system.. Identity
+ of the terminology system.
+ type:
+ - string
+ - 'null'
+ input_0_valueAttachment_contentType:
+ description: Mime type of the content, with charset etc..
+ type:
+ - string
+ - 'null'
+ input_0_valueAttachment_creation:
+ description: Date attachment was first created.
+ type:
+ - string
+ - 'null'
+ input_0_valueAttachment_data:
+ description: Data inline, base64ed.
+ type:
+ - string
+ - 'null'
+ input_0_valueAttachment_hash:
+ description: Hash of the data (sha-1, base64ed).
+ type:
+ - string
+ - 'null'
+ input_0_valueAttachment_language:
+ description: Human language of the content (BCP-47).
+ type:
+ - string
+ - 'null'
+ input_0_valueAttachment_size:
+ description: Number of bytes of content (if url provided).
+ type:
+ - number
+ - 'null'
+ input_0_valueAttachment_title:
+ description: Label to display in place of the data.
+ type:
+ - string
+ - 'null'
+ input_0_valueAttachment_url:
+ description: Uri where the data can be found.
+ type:
+ - string
+ - 'null'
+ input_0_valueBase64Binary:
+ description: Content to use in performing the task.
+ type:
+ - string
+ input_0_valueBoolean:
+ description: Content to use in performing the task.
+ type:
+ - boolean
+ input_0_valueCanonical:
+ description: Content to use in performing the task.
+ type:
+ - string
+ input_0_valueCode:
+ description: Content to use in performing the task.
+ type:
+ - string
+ input_0_valueCodeableConcept_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ input_0_valueCodeableConcept_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ input_0_valueCodeableConcept_coding_0_system:
+ description: Content to use in performing the task.. Code defined by a terminology
+ system.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ input_0_valueCoding_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ input_0_valueCoding_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ input_0_valueCoding_system:
+ description: Content to use in performing the task.. Identity of the terminology
+ system.
+ type:
+ - string
+ - 'null'
+ input_0_valueContactDetail_name:
+ description: Name of an individual to contact.
+ type:
+ - string
+ - 'null'
+ input_0_valueContributor_contact_0_name:
+ description: Name of an individual to contact.
+ type:
+ - string
+ - 'null'
+ input_0_valueContributor_name:
+ description: Who contributed the content.
+ type:
+ - string
+ input_0_valueContributor_type:
+ description: The type of contributor.. http://hl7.org/fhir/contributor-type
+ enum:
+ - author
+ - editor
+ - reviewer
+ - endorser
+ term:
+ description: The type of contributor.. http://hl7.org/fhir/contributor-type
+ termDef:
+ cde_id: http://hl7.org/fhir/contributor-type
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/contributor-type
+ term_url: http://hl7.org/fhir/contributor-type
+ input_0_valueCount_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ input_0_valueCount_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.
+ type:
+ - string
+ - 'null'
+ input_0_valueCount_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ input_0_valueCount_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ input_0_valueCount_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ input_0_valueDataRequirement_codeFilter_0_path:
+ description: A code-valued attribute to filter on.
+ type:
+ - string
+ - 'null'
+ input_0_valueDataRequirement_codeFilter_0_searchParam:
+ description: A coded (token) parameter to search on.
+ type:
+ - string
+ - 'null'
+ input_0_valueDataRequirement_codeFilter_0_valueSet:
+ description: Valueset for the filter.
+ type:
+ - string
+ - 'null'
+ input_0_valueDataRequirement_dateFilter_0_path:
+ description: A date-valued attribute to filter on.
+ type:
+ - string
+ - 'null'
+ input_0_valueDataRequirement_dateFilter_0_searchParam:
+ description: A date valued parameter to search on.
+ type:
+ - string
+ - 'null'
+ input_0_valueDataRequirement_dateFilter_0_valueDateTime:
+ description: The value of the filter, as a Period, DateTime, or Duration value.
+ type:
+ - string
+ - 'null'
+ input_0_valueDataRequirement_limit:
+ description: Number of results.
+ type:
+ - number
+ - 'null'
+ input_0_valueDataRequirement_mustSupport:
+ description: Indicates specific structure elements that are referenced by the
+ knowledge module.
+ type:
+ - string
+ - 'null'
+ input_0_valueDataRequirement_profile:
+ description: The profile of the required data.
+ type:
+ - string
+ - 'null'
+ input_0_valueDataRequirement_sort_0_direction:
+ description: The direction of the sort, ascending or descending.. http://hl7.org/fhir/sort-direction
+ enum:
+ - ascending
+ - descending
+ term:
+ description: The direction of the sort, ascending or descending.. http://hl7.org/fhir/sort-direction
+ termDef:
+ cde_id: http://hl7.org/fhir/sort-direction
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/sort-direction
+ term_url: http://hl7.org/fhir/sort-direction
+ input_0_valueDataRequirement_sort_0_path:
+ description: The name of the attribute to perform the sort.
+ type:
+ - string
+ input_0_valueDataRequirement_subjectReference_reference:
+ description: E.g. Patient, Practitioner, RelatedPerson, Organization, Location,
+ Device.. Literal reference, Relative, internal or absolute URL.
+ type:
+ - string
+ - 'null'
+ input_0_valueDataRequirement_subjectReference_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ input_0_valueDataRequirement_type:
+ description: The type of the required data.
+ type:
+ - string
+ input_0_valueDate:
+ description: Content to use in performing the task.
+ type:
+ - string
+ input_0_valueDateTime:
+ description: Content to use in performing the task.
+ type:
+ - string
+ input_0_valueDecimal:
+ description: Content to use in performing the task.
+ type:
+ - number
+ input_0_valueDistance_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ input_0_valueDistance_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.
+ type:
+ - string
+ - 'null'
+ input_0_valueDistance_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ input_0_valueDistance_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ input_0_valueDistance_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ input_0_valueDosage_asNeededBoolean:
+ description: Take "as needed" (for x).
+ type:
+ - boolean
+ - 'null'
+ input_0_valueDosage_maxDosePerAdministration_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ input_0_valueDosage_maxDosePerAdministration_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ input_0_valueDosage_maxDosePerAdministration_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ input_0_valueDosage_maxDosePerAdministration_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ input_0_valueDosage_maxDosePerAdministration_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ input_0_valueDosage_maxDosePerLifetime_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ input_0_valueDosage_maxDosePerLifetime_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ input_0_valueDosage_maxDosePerLifetime_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ input_0_valueDosage_maxDosePerLifetime_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ input_0_valueDosage_maxDosePerLifetime_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ input_0_valueDosage_patientInstruction:
+ description: Patient or consumer oriented instructions.
+ type:
+ - string
+ - 'null'
+ input_0_valueDosage_sequence:
+ description: The order of the dosage instructions.
+ type:
+ - number
+ - 'null'
+ input_0_valueDosage_text:
+ description: Free text dosage instructions e.g. SIG.
+ type:
+ - string
+ - 'null'
+ input_0_valueDosage_timing_event:
+ description: When the event occurs.
+ type:
+ - string
+ - 'null'
+ input_0_valueDuration_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ input_0_valueDuration_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.
+ type:
+ - string
+ - 'null'
+ input_0_valueDuration_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ input_0_valueDuration_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ input_0_valueDuration_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ input_0_valueExpression_description:
+ description: Natural language description of the condition.
+ type:
+ - string
+ - 'null'
+ input_0_valueExpression_expression:
+ description: Expression in specified language.
+ type:
+ - string
+ - 'null'
+ input_0_valueExpression_language:
+ description: The media type of the language for the expression.. http://hl7.org/fhir/expression-language
+ text/cql|text/fhirpath|application/x-fhir-query
+ term:
+ description: The media type of the language for the expression.. http://hl7.org/fhir/expression-language
+ text/cql|text/fhirpath|application/x-fhir-query
+ termDef:
+ cde_id: http://hl7.org/fhir/expression-language
+ cde_version: null
+ source: fhir
+ strength: extensible
+ term: http://hl7.org/fhir/expression-language
+ term_url: http://hl7.org/fhir/expression-language
+ type:
+ - string
+ input_0_valueExpression_name:
+ description: Short name assigned to expression for reuse.
+ type:
+ - string
+ - 'null'
+ input_0_valueExpression_reference:
+ description: Where the expression is found.
+ type:
+ - string
+ - 'null'
+ input_0_valueId:
+ description: Content to use in performing the task.
+ type:
+ - string
+ input_0_valueIdentifier_system:
+ description: The namespace for the identifier value.
+ type:
+ - string
+ - 'null'
+ input_0_valueIdentifier_use:
+ description: Content to use in performing the task.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ enum:
+ - usual
+ - official
+ - temp
+ - secondary
+ - old
+ term:
+ description: Content to use in performing the task.. The purpose of this identifier..
+ http://hl7.org/fhir/identifier-use
+ termDef:
+ cde_id: http://hl7.org/fhir/identifier-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/identifier-use
+ term_url: http://hl7.org/fhir/identifier-use
+ input_0_valueIdentifier_value:
+ description: The value that is unique.
+ type:
+ - string
+ - 'null'
+ input_0_valueInstant:
+ description: Content to use in performing the task.
+ type:
+ - string
+ input_0_valueInteger:
+ description: Content to use in performing the task.
+ type:
+ - number
+ input_0_valueMarkdown:
+ description: Content to use in performing the task.
+ type:
+ - string
+ input_0_valueMeta_lastUpdated:
+ description: When the resource version last changed.
+ type:
+ - string
+ - 'null'
+ input_0_valueMeta_profile:
+ description: Profiles this resource claims to conform to.
+ type:
+ - string
+ - 'null'
+ input_0_valueMeta_security_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ input_0_valueMeta_security_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ input_0_valueMeta_security_0_system:
+ description: Security Labels applied to this resource.. Identity of the terminology
+ system.
+ type:
+ - string
+ - 'null'
+ input_0_valueMeta_source:
+ description: Identifies where the resource comes from.
+ type:
+ - string
+ - 'null'
+ input_0_valueMeta_tag_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ input_0_valueMeta_tag_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ input_0_valueMeta_tag_0_system:
+ description: Tags applied to this resource.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ input_0_valueMeta_versionId:
+ description: Version specific identifier.
+ type:
+ - string
+ - 'null'
+ input_0_valueMoney_currency:
+ description: ISO 4217 Currency Code.
+ type:
+ - string
+ - 'null'
+ input_0_valueMoney_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ input_0_valueOid:
+ description: Content to use in performing the task.
+ type:
+ - string
+ input_0_valueParameterDefinition_documentation:
+ description: A brief description of the parameter.
+ type:
+ - string
+ - 'null'
+ input_0_valueParameterDefinition_max:
+ description: Maximum cardinality (a number of *).
+ type:
+ - string
+ - 'null'
+ input_0_valueParameterDefinition_min:
+ description: Minimum cardinality.
+ type:
+ - number
+ - 'null'
+ input_0_valueParameterDefinition_name:
+ description: Name used to access the parameter value.
+ type:
+ - string
+ - 'null'
+ input_0_valueParameterDefinition_profile:
+ description: What profile the value is expected to be.
+ type:
+ - string
+ - 'null'
+ input_0_valueParameterDefinition_type:
+ description: What type of value.
+ type:
+ - string
+ input_0_valueParameterDefinition_use:
+ description: Whether the parameter is input or output for the module.. http://hl7.org/fhir/operation-parameter-use
+ enum:
+ - in
+ - out
+ term:
+ description: Whether the parameter is input or output for the module.. http://hl7.org/fhir/operation-parameter-use
+ termDef:
+ cde_id: http://hl7.org/fhir/operation-parameter-use
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/operation-parameter-use
+ term_url: http://hl7.org/fhir/operation-parameter-use
+ input_0_valuePeriod_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ input_0_valuePeriod_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ input_0_valuePositiveInt:
+ description: Content to use in performing the task.
+ type:
+ - number
+ input_0_valueQuantity_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ input_0_valueQuantity_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ input_0_valueQuantity_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ input_0_valueQuantity_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ input_0_valueQuantity_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ input_0_valueRange_high_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ input_0_valueRange_high_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ input_0_valueRange_high_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ input_0_valueRange_high_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ input_0_valueRange_high_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ input_0_valueRange_low_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ input_0_valueRange_low_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ input_0_valueRange_low_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ input_0_valueRange_low_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ input_0_valueRange_low_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ input_0_valueRatio_denominator_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ input_0_valueRatio_denominator_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ input_0_valueRatio_denominator_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ input_0_valueRatio_denominator_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ input_0_valueRatio_denominator_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ input_0_valueRatio_numerator_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ input_0_valueRatio_numerator_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ input_0_valueRatio_numerator_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ input_0_valueRatio_numerator_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ input_0_valueRatio_numerator_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ input_0_valueReference_reference:
+ description: Content to use in performing the task.. Literal reference, Relative,
+ internal or absolute URL.
+ type:
+ - string
+ - 'null'
+ input_0_valueReference_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ input_0_valueRelatedArtifact_citation:
+ description: Bibliographic citation for the artifact.
+ type:
+ - string
+ - 'null'
+ input_0_valueRelatedArtifact_display:
+ description: Brief description of the related artifact.
+ type:
+ - string
+ - 'null'
+ input_0_valueRelatedArtifact_document_contentType:
+ description: Mime type of the content, with charset etc..
+ type:
+ - string
+ - 'null'
+ input_0_valueRelatedArtifact_document_creation:
+ description: Date attachment was first created.
+ type:
+ - string
+ - 'null'
+ input_0_valueRelatedArtifact_document_data:
+ description: Data inline, base64ed.
+ type:
+ - string
+ - 'null'
+ input_0_valueRelatedArtifact_document_hash:
+ description: Hash of the data (sha-1, base64ed).
+ type:
+ - string
+ - 'null'
+ input_0_valueRelatedArtifact_document_language:
+ description: Human language of the content (BCP-47).
+ type:
+ - string
+ - 'null'
+ input_0_valueRelatedArtifact_document_size:
+ description: Number of bytes of content (if url provided).
+ type:
+ - number
+ - 'null'
+ input_0_valueRelatedArtifact_document_title:
+ description: Label to display in place of the data.
+ type:
+ - string
+ - 'null'
+ input_0_valueRelatedArtifact_document_url:
+ description: Uri where the data can be found.
+ type:
+ - string
+ - 'null'
+ input_0_valueRelatedArtifact_label:
+ description: Short label.
+ type:
+ - string
+ - 'null'
+ input_0_valueRelatedArtifact_resource:
+ description: What resource is being referenced.
+ type:
+ - string
+ - 'null'
+ input_0_valueRelatedArtifact_type:
+ description: The type of relationship to the related artifact.. http://hl7.org/fhir/related-artifact-type
+ enum:
+ - documentation
+ - justification
+ - citation
+ - predecessor
+ - successor
+ - derived-from
+ - depends-on
+ - composed-of
+ term:
+ description: The type of relationship to the related artifact.. http://hl7.org/fhir/related-artifact-type
+ termDef:
+ cde_id: http://hl7.org/fhir/related-artifact-type
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/related-artifact-type
+ term_url: http://hl7.org/fhir/related-artifact-type
+ input_0_valueRelatedArtifact_url:
+ description: Where the artifact can be accessed.
+ type:
+ - string
+ - 'null'
+ input_0_valueSampledData_data:
+ description: Decimal values with spaces, or "E" | "U" | "L".
+ type:
+ - string
+ - 'null'
+ input_0_valueSampledData_dimensions:
+ description: Number of sample points at each time point.
+ type:
+ - number
+ input_0_valueSampledData_factor:
+ description: Multiply data by this before adding to origin.
+ type:
+ - number
+ - 'null'
+ input_0_valueSampledData_lowerLimit:
+ description: Lower limit of detection.
+ type:
+ - number
+ - 'null'
+ input_0_valueSampledData_origin_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ input_0_valueSampledData_origin_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ input_0_valueSampledData_origin_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ input_0_valueSampledData_origin_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ input_0_valueSampledData_origin_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ input_0_valueSampledData_period:
+ description: Number of milliseconds between samples.
+ type:
+ - number
+ input_0_valueSampledData_upperLimit:
+ description: Upper limit of detection.
+ type:
+ - number
+ - 'null'
+ input_0_valueSignature_data:
+ description: The actual signature content (XML DigSig. JWS, picture, etc.).
+ type:
+ - string
+ - 'null'
+ input_0_valueSignature_onBehalfOf_reference:
+ description: The party represented.. Literal reference, Relative, internal or
+ absolute URL.
+ type:
+ - string
+ - 'null'
+ input_0_valueSignature_onBehalfOf_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ input_0_valueSignature_sigFormat:
+ description: The technical format of the signature.
+ type:
+ - string
+ - 'null'
+ input_0_valueSignature_targetFormat:
+ description: The technical format of the signed resources.
+ type:
+ - string
+ - 'null'
+ input_0_valueSignature_type_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ input_0_valueSignature_type_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ input_0_valueSignature_type_0_system:
+ description: Indication of the reason the entity signed the object(s).. Identity
+ of the terminology system.
+ type:
+ - string
+ - 'null'
+ input_0_valueSignature_when:
+ description: When the signature was created.
+ type:
+ - string
+ input_0_valueSignature_who_reference:
+ description: Who signed.. Literal reference, Relative, internal or absolute URL.
+ type:
+ - string
+ - 'null'
+ input_0_valueSignature_who_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ input_0_valueString:
+ description: Content to use in performing the task.
+ type:
+ - string
+ input_0_valueTime:
+ description: Content to use in performing the task.
+ type:
+ - string
+ input_0_valueTiming_event:
+ description: When the event occurs.
+ type:
+ - string
+ - 'null'
+ input_0_valueTiming_repeat_count:
+ description: Number of times to repeat.
+ type:
+ - number
+ - 'null'
+ input_0_valueTiming_repeat_countMax:
+ description: Maximum number of times to repeat.
+ type:
+ - number
+ - 'null'
+ input_0_valueTiming_repeat_dayOfWeek:
+ description: If one or more days of week is provided, then the action happens
+ only on the specified day(s).. http://hl7.org/fhir/days-of-week
+ enum:
+ - mon
+ - tue
+ - wed
+ - thu
+ - fri
+ - sat
+ - sun
+ term:
+ description: If one or more days of week is provided, then the action happens
+ only on the specified day(s).. http://hl7.org/fhir/days-of-week
+ termDef:
+ cde_id: http://hl7.org/fhir/days-of-week
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/days-of-week
+ term_url: http://hl7.org/fhir/days-of-week
+ input_0_valueTiming_repeat_duration:
+ description: How long when it happens.
+ type:
+ - number
+ - 'null'
+ input_0_valueTiming_repeat_durationMax:
+ description: How long when it happens (Max).
+ type:
+ - number
+ - 'null'
+ input_0_valueTiming_repeat_durationUnit:
+ description: s | min | h | d | wk | mo | a - unit of time (UCUM).
+ type:
+ - string
+ - 'null'
+ input_0_valueTiming_repeat_frequency:
+ description: Event occurs frequency times per period.
+ type:
+ - number
+ - 'null'
+ input_0_valueTiming_repeat_frequencyMax:
+ description: Event occurs up to frequencyMax times per period.
+ type:
+ - number
+ - 'null'
+ input_0_valueTiming_repeat_offset:
+ description: Minutes from event (before or after).
+ type:
+ - number
+ - 'null'
+ input_0_valueTiming_repeat_period:
+ description: Event occurs frequency times per period.
+ type:
+ - number
+ - 'null'
+ input_0_valueTiming_repeat_periodMax:
+ description: Upper limit of period (3-4 hours).
+ type:
+ - number
+ - 'null'
+ input_0_valueTiming_repeat_periodUnit:
+ description: s | min | h | d | wk | mo | a - unit of time (UCUM).
+ type:
+ - string
+ - 'null'
+ input_0_valueTiming_repeat_timeOfDay:
+ description: Time of day for action.
+ type:
+ - string
+ - 'null'
+ input_0_valueTiming_repeat_when:
+ description: Code for time period of occurrence.
+ type:
+ - string
+ - 'null'
+ input_0_valueTriggerDefinition_condition_description:
+ description: Natural language description of the condition.
+ type:
+ - string
+ - 'null'
+ input_0_valueTriggerDefinition_condition_expression:
+ description: Expression in specified language.
+ type:
+ - string
+ - 'null'
+ input_0_valueTriggerDefinition_condition_language:
+ description: The media type of the language for the expression.. http://hl7.org/fhir/expression-language
+ text/cql|text/fhirpath|application/x-fhir-query
+ term:
+ description: The media type of the language for the expression.. http://hl7.org/fhir/expression-language
+ text/cql|text/fhirpath|application/x-fhir-query
+ termDef:
+ cde_id: http://hl7.org/fhir/expression-language
+ cde_version: null
+ source: fhir
+ strength: extensible
+ term: http://hl7.org/fhir/expression-language
+ term_url: http://hl7.org/fhir/expression-language
+ type:
+ - string
+ input_0_valueTriggerDefinition_condition_name:
+ description: Short name assigned to expression for reuse.
+ type:
+ - string
+ - 'null'
+ input_0_valueTriggerDefinition_condition_reference:
+ description: Where the expression is found.
+ type:
+ - string
+ - 'null'
+ input_0_valueTriggerDefinition_data_0_limit:
+ description: Number of results.
+ type:
+ - number
+ - 'null'
+ input_0_valueTriggerDefinition_data_0_mustSupport:
+ description: Indicates specific structure elements that are referenced by the
+ knowledge module.
+ type:
+ - string
+ - 'null'
+ input_0_valueTriggerDefinition_data_0_profile:
+ description: The profile of the required data.
+ type:
+ - string
+ - 'null'
+ input_0_valueTriggerDefinition_data_0_type:
+ description: The type of the required data.
+ type:
+ - string
+ input_0_valueTriggerDefinition_name:
+ description: Name or URI that identifies the event.
+ type:
+ - string
+ - 'null'
+ input_0_valueTriggerDefinition_timingDate:
+ description: Timing of the event.
+ type:
+ - string
+ - 'null'
+ input_0_valueTriggerDefinition_timingDateTime:
+ description: Timing of the event.
+ type:
+ - string
+ - 'null'
+ input_0_valueTriggerDefinition_timingReference_reference:
+ description: Timing of the event.. Literal reference, Relative, internal or absolute
+ URL.
+ type:
+ - string
+ - 'null'
+ input_0_valueTriggerDefinition_timingReference_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ input_0_valueTriggerDefinition_timingTiming_event:
+ description: When the event occurs.
+ type:
+ - string
+ - 'null'
+ input_0_valueTriggerDefinition_type:
+ description: The type of triggering event.. http://hl7.org/fhir/trigger-type
+ enum:
+ - named-event
+ - periodic
+ - data-changed
+ - data-added
+ - data-modified
+ - data-removed
+ - data-accessed
+ - data-access-ended
+ term:
+ description: The type of triggering event.. http://hl7.org/fhir/trigger-type
+ termDef:
+ cde_id: http://hl7.org/fhir/trigger-type
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/trigger-type
+ term_url: http://hl7.org/fhir/trigger-type
+ input_0_valueUnsignedInt:
+ description: Content to use in performing the task.
+ type:
+ - number
+ input_0_valueUri:
+ description: Content to use in performing the task.
+ type:
+ - string
+ input_0_valueUrl:
+ description: Content to use in performing the task.
+ type:
+ - string
+ input_0_valueUsageContext_code_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ input_0_valueUsageContext_code_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ input_0_valueUsageContext_code_system:
+ description: Type of context being specified.. Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ input_0_valueUsageContext_valueQuantity_code:
+ description: Coded form of the unit.
+ type:
+ - string
+ - 'null'
+ input_0_valueUsageContext_valueQuantity_comparator:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ enum:
+ - <
+ - <=
+ - '>='
+ - '>'
+ term:
+ description: How the value should be understood and represented - whether the
+ actual value is greater or less than the stated value due to measurement issues;
+ e.g. if the comparator is "<" , then the real value is < stated value.. http://hl7.org/fhir/quantity-comparator
+ termDef:
+ cde_id: http://hl7.org/fhir/quantity-comparator
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/quantity-comparator
+ term_url: http://hl7.org/fhir/quantity-comparator
+ input_0_valueUsageContext_valueQuantity_system:
+ description: System that defines coded unit form.
+ type:
+ - string
+ - 'null'
+ input_0_valueUsageContext_valueQuantity_unit:
+ description: Unit representation.
+ type:
+ - string
+ - 'null'
+ input_0_valueUsageContext_valueQuantity_value:
+ description: Numerical value (with implicit precision).
+ type:
+ - number
+ - 'null'
+ input_0_valueUsageContext_valueReference_reference:
+ description: Value that defines the context.. Literal reference, Relative, internal
+ or absolute URL.
+ type:
+ - string
+ - 'null'
+ input_0_valueUsageContext_valueReference_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ input_0_valueUuid:
+ description: Content to use in performing the task.
+ type:
+ - string
+ instantiatesCanonical:
+ description: Formal definition of task.
+ type:
+ - string
+ - 'null'
+ instantiatesUri:
+ description: Formal definition of task.
+ type:
+ - string
+ - 'null'
+ intent:
+ description: unknown | proposal | plan | order | original-order | reflex-order
+ | filler-order | instance-order | option.
+ type:
+ - string
+ language:
+ description: Language of the resource content.
+ type:
+ - string
+ - 'null'
+ lastModified:
+ description: Task Last Modified Date.
+ type:
+ - string
+ - 'null'
+ performerType_0_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ performerType_0_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ performerType_0_coding_0_system:
+ description: Requested performer.. Code defined by a terminology system.. Identity
+ of the terminology system.
+ type:
+ - string
+ - 'null'
+ priority:
+ description: Indicates how quickly the Task should be addressed with respect to
+ other requests.. http://hl7.org/fhir/request-priority
+ enum:
+ - routine
+ - urgent
+ - asap
+ - stat
+ term:
+ description: Indicates how quickly the Task should be addressed with respect
+ to other requests.. http://hl7.org/fhir/request-priority
+ termDef:
+ cde_id: http://hl7.org/fhir/request-priority
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/request-priority
+ term_url: http://hl7.org/fhir/request-priority
+ project_id:
+ $ref: _definitions.yaml#/project_id
+ reasonCode_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ reasonCode_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ reasonCode_coding_0_system:
+ description: Why task is needed.. Code defined by a terminology system.. Identity
+ of the terminology system.
+ type:
+ - string
+ - 'null'
+ resource_type:
+ description: ''
+ type:
+ - string
+ - 'null'
+ restriction_period_end:
+ description: End time with inclusive boundary, if not ongoing.
+ type:
+ - string
+ - 'null'
+ restriction_period_start:
+ description: Starting time with inclusive boundary.
+ type:
+ - string
+ - 'null'
+ restriction_recipient_0_reference:
+ description: For whom is fulfillment sought?.. Literal reference, Relative, internal
+ or absolute URL.
+ type:
+ - string
+ - 'null'
+ restriction_recipient_0_type:
+ description: Type the reference refers to (e.g. "Patient").
+ type:
+ - string
+ - 'null'
+ restriction_repetitions:
+ description: How many times to repeat.
+ type:
+ - number
+ - 'null'
+ state:
+ $ref: _definitions.yaml#/state
+ status:
+ description: The current status of the task.. http://hl7.org/fhir/task-status
+ enum:
+ - draft
+ - requested
+ - received
+ - accepted
+ - rejected
+ - ready
+ - cancelled
+ - in-progress
+ - on-hold
+ - failed
+ - completed
+ - entered-in-error
+ term:
+ description: The current status of the task.. http://hl7.org/fhir/task-status
+ termDef:
+ cde_id: http://hl7.org/fhir/task-status
+ cde_version: null
+ source: fhir
+ strength: required
+ term: http://hl7.org/fhir/task-status
+ term_url: http://hl7.org/fhir/task-status
+ statusReason_coding_0_code:
+ description: Symbol in syntax defined by the system.
+ type:
+ - string
+ - 'null'
+ statusReason_coding_0_display:
+ description: Representation defined by the system.
+ type:
+ - string
+ - 'null'
+ statusReason_coding_0_system:
+ description: Reason for current status.. Code defined by a terminology system..
+ Identity of the terminology system.
+ type:
+ - string
+ - 'null'
+ submitter_id:
+ type:
+ - string
+ - 'null'
+ updated_datetime:
+ $ref: _definitions.yaml#/datetime
+required:
+- submitter_id
+- status
+- intent
+submittable: true
+systemProperties:
+- id
+- project_id
+- created_datetime
+- updated_datetime
+- state
+title: Task
+type: object
+uniqueKeys:
+- - id
+- - project_id
+ - submitter_id
+validators: null
diff --git a/datadictionary/gdcdictionary/schemas/treatment.yaml b/datadictionary/gdcdictionary/schemas/treatment.yaml
deleted file mode 100644
index 4ed89da0..00000000
--- a/datadictionary/gdcdictionary/schemas/treatment.yaml
+++ /dev/null
@@ -1,217 +0,0 @@
-$schema: "http://json-schema.org/draft-04/schema#"
-
-id: "treatment"
-title: Treatment
-type: object
-namespace: http://gdc.nci.nih.gov
-category: clinical
-program: '*'
-project: '*'
-description: >
- Record of the administration and intention of therapeutic agents provided to a patient to alter
- the course of a pathologic process.
-additionalProperties: false
-submittable: true
-validators: null
-
-systemProperties:
- - id
- - project_id
- - state
- - created_datetime
- - updated_datetime
-
-required:
- - submitter_id
- - type
-
-links:
- - name: diagnoses
- backref: treatments
- label: describes #need better term here
- target_type: diagnosis
- multiplicity: many_to_one
- required: true
-
-
-
-uniqueKeys:
- #unclear if want submitter ID for clinical
- - [id]
- - [project_id, submitter_id]
-
-properties:
- type:
- enum: [ "treatment" ]
-
- id:
- $ref: "_definitions.yaml#/UUID"
- systemAlias: node_id
-
- state:
- $ref: "_definitions.yaml#/state"
-
- submitter_id:
- type:
- - string
- - "null"
-
- days_to_treatment:
- term:
- $ref: "_terms.yaml#/days_to_treatment"
- type: number
-
- days_to_treatment_end:
- term:
- $ref: "_terms.yaml#/days_to_treatment_end"
- type: number
-
- days_to_treatment_start:
- term:
- $ref: "_terms.yaml#/days_to_treatment_start"
- type: number
-
- therapeutic_agents:
- term:
- $ref: "_terms.yaml#/therapeutic_agents"
- type: string
-
- treatment_anatomic_site:
- term:
- $ref: "_terms.yaml#/treatment_anatomic_site"
- enum:
- - Abdomen, total
- - Arm
- - Ascites
- - Axillary
- - Body, total
- - Bone
- - Bone, non-spine
- - Brain, focal
- - Brain, whole
- - Brain-C2
- - Breast
- - Cervical
- - Chest Wall
- - Effusion
- - Epitrochlear
- - Eye
- - Femoral
- - Gastrointestinal, Colon
- - Gastrointestinal, Gallbladder
- - Gastrointestinal, Intestine
- - Gastrointestinal, Liver
- - Gastrointestinal, NOS
- - Gastrointestinal, Pancreas
- - Gastrointestinal, Rectum
- - Gastrointestinal, Stomach
- - Genitourinary, Bladder
- - Genitourinary, Kidney
- - Genitourinary, NOS
- - Genitourinary, Prostate
- - Genitourinary, Prostate and Seminal Vesicles
- - Head
- - Head, Face, or Neck
- - Hilar
- - Iliac-common
- - Iliac-external
- - Inguinal
- - Internal Mammary Nodes
- - Leg
- - Lung
- - Lymph Nodes
- - Lymph node, distant (specify site)
- - Lymph node, locoregional (specify site)
- - Mantle
- - Mediastinal
- - Mediastinum
- - Mesenteric
- - Occipital
- - Other
- - Paraaortic
- - Parametrium
- - Parotid
- - Pelvis
- - Popliteal
- - Primary tumor site
- - Prostate
- - Prostate Bed
- - Prostate, Seminal Vesicles and Lymph Nodes
- - Rectum
- - Retroperitoneal
- - Sacrum
- - Seminal vesicles
- - Shoulder
- - Skin, lower extremity, local
- - Skin, total
- - Skin, trunk, local
- - Skin, upper extremity, local
- - Spine
- - Spine, whole
- - Splenic
- - Submandibular
- - Supraclavicular
- - Supraclavicular/Axillary Level 3
- - Thorax
- - Trunk
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- treatment_intent_type:
- term:
- $ref: "_terms.yaml#/treatment_intent_type"
- type: string
-
- treatment_or_therapy:
- term:
- $ref: "_terms.yaml#/treatment_or_therapy"
- enum:
- - "yes"
- - "no"
- - unknown
- - not reported
-
- treatment_outcome:
- term:
- $ref: "_terms.yaml#/treatment_outcome"
- enum:
- - Complete Response
- - Partial Response
- - Treatment Ongoing
- - Treatment Stopped Due to Toxicity
- - Unknown
-
- treatment_type:
- term:
- $ref: "_terms.yaml#/treatment_type"
- enum:
- - Ablation
- - Chemotherapy
- - Concurrent Chemoradiation
- - Cryoablation
- - Embolization
- - Hormone Therapy
- - Internal Radiation
- - Immunotherapy (Including Vaccines)
- - Other
- - Pharmaceutical Therapy
- - Radiation Therapy
- - Stem Cell Treatment
- - Surgery
- - Targeted Molecular Therapy
- - Unknown
- - Not Reported
- - Not Allowed To Collect
-
- diagnoses:
- $ref: "_definitions.yaml#/to_one"
-
- project_id:
- $ref: "_definitions.yaml#/project_id"
-
- # ======== Timestamps ========
- created_datetime:
- $ref: "_definitions.yaml#/datetime"
- updated_datetime:
- $ref: "_definitions.yaml#/datetime"
diff --git a/docker-compose.override.yml b/docker-compose.override.yml
new file mode 100644
index 00000000..27de6ae7
--- /dev/null
+++ b/docker-compose.override.yml
@@ -0,0 +1,179 @@
+version: '3'
+
+
+# Settings and configurations that are common for all minio containers
+x-minio-common: &minio-common
+ image: quay.io/minio/minio:RELEASE.2022-09-01T23-53-36Z
+ expose:
+ - "9000"
+ - "9001"
+ environment:
+ MINIO_ROOT_USER: "${MINIO_ROOT_USER}"
+ MINIO_ROOT_PASSWORD: "${MINIO_ROOT_PASSWORD}"
+ MINIO_NOTIFY_WEBHOOK_ENABLE_PRIMARY: on
+ MINIO_NOTIFY_WEBHOOK_ENDPOINT_PRIMARY: http://minio-webhook:3000/file-uploaded
+
+ networks:
+ - devnet
+ healthcheck:
+ test: ["CMD", "curl", "-f", "http://localhost:9000/minio/health/live"]
+ interval: 30s
+ timeout: 20s
+ retries: 3
+
+services:
+# starts 5 docker containers running minio server instances.
+# using nginx reverse proxy, load balancing, you can access
+# it through port 9000.
+ minio-default:
+ <<: *minio-common
+ hostname: minio-default
+ container_name: minio-default
+ command: server --console-address ":9001" http://minio-default/data{1...2}
+ volumes:
+ - data1-1:/data1
+ - data1-2:/data2
+
+ minio-ohsu:
+ <<: *minio-common
+ hostname: minio-ohsu
+ container_name: minio-ohsu
+ command: server --console-address ":9001" http://minio-ohsu/data{1...2}
+ volumes:
+ - data2-1:/data1
+ - data2-2:/data2
+
+ minio-ucl:
+ <<: *minio-common
+ hostname: minio-ucl
+ container_name: minio-ucl
+ command: server --console-address ":9001" http://minio-ucl/data{1...2}
+ volumes:
+ - data3-1:/data1
+ - data3-2:/data2
+
+ minio-manchester:
+ <<: *minio-common
+ hostname: minio-manchester
+ container_name: minio-manchester
+ command: server --console-address ":9001" http://minio-manchester/data{1...2}
+ volumes:
+ - data4-1:/data1
+ - data4-2:/data2
+
+ minio-stanford:
+ <<: *minio-common
+ hostname: minio-stanford
+ container_name: minio-stanford
+ command: server --console-address ":9001" http://minio-stanford/data{1...2}
+ volumes:
+ - data5-1:/data1
+ - data5-2:/data2
+
+
+ #
+ # adds minio dependencies to revproxy
+ #
+ revproxy-service:
+ depends_on:
+ - arborist-service
+ - indexd-service
+ - peregrine-service
+ - sheepdog-service
+ - fence-service
+ - portal-service
+ - pidgin-service
+ - minio-default
+ - minio-ohsu
+ - minio-ucl
+ - minio-manchester
+ - minio-stanford
+
+## create an etl container to hold our command line tools
+ etl-service:
+ build: etl
+ hostname: etl-service
+ container_name: etl-service
+ command: tail -f /dev/null
+ networks:
+ - devnet
+ depends_on:
+ - minio-default
+ - minio-ohsu
+ - minio-ucl
+ - minio-manchester
+ - minio-stanford
+ restart: unless-stopped
+ logging:
+ driver: "json-file"
+ options:
+ max-size: "10m"
+## create a web-hook container handle events
+ minio-webhook:
+ build: minio-webhook
+ hostname: minio-webhook
+ container_name: minio-webhook
+ expose:
+ - "3000"
+ command: uvicorn main:app --reload --host 0.0.0.0 --port 3000
+ networks:
+ - devnet
+ volumes:
+ - ./minio-webhook:/minio-webhook
+ depends_on:
+ - minio-default
+ - minio-ohsu
+ - minio-ucl
+ - minio-manchester
+ - minio-stanford
+ restart: unless-stopped
+ logging:
+ driver: "json-file"
+ options:
+ max-size: "10m"
+
+# expose postgres to host os
+
+ postgres:
+ ports:
+ - 5432:5432
+
+# adds fhir service
+ fhir-service:
+ image: "hapiproject/hapi:v6.1.0"
+ ports:
+ - "8090:8080"
+ volumes:
+ - ./fhir/data:/data/hapi
+ environment:
+ SPRING_CONFIG_LOCATION: 'file:///data/hapi/application.yaml'
+ networks:
+ - devnet
+ logging:
+ driver: "json-file"
+ options:
+ max-size: "10m"
+
+ dash-service:
+ build: dash
+ container_name: dash-service
+ ports:
+ - "8050:8050" # HOST:CONTAINER
+ environment:
+ - DASH_URL_BASE_PATHNAME=/dash/proxy/
+ networks:
+ - devnet
+
+## By default this config uses default local driver,
+## For custom volumes replace with volume driver configuration.
+volumes:
+ data1-1:
+ data1-2:
+ data2-1:
+ data2-2:
+ data3-1:
+ data3-2:
+ data4-1:
+ data4-2:
+ data5-1:
+ data5-2:
diff --git a/docker-compose.yml b/docker-compose.yml
index 378c0956..4fa935af 100644
--- a/docker-compose.yml
+++ b/docker-compose.yml
@@ -11,6 +11,10 @@ services:
- ./scripts/postgres_always.sh:/postgres_always.sh:ro
- ./scripts/postgres_run.sh:/usr/local/bin/postgres_run.sh:ro
restart: unless-stopped
+ logging:
+ driver: "json-file"
+ options:
+ max-size: "10m"
healthcheck:
test: ["CMD-SHELL", "psql -U fence_user -d fence_db -c 'SELECT 1;'"]
interval: 60s
@@ -20,11 +24,14 @@ services:
environment:
- POSTGRES_PASSWORD=postgres
indexd-service:
- image: "quay.io/cdis/indexd:2021.03"
+ image: quay.io/cdis/indexd:2022.12 #"quay.io/cdis/indexd:2022.09"
command: bash indexd_setup.sh
container_name: indexd-service
networks:
- devnet
+ environment:
+ # https://github.com/uc-cdis/indexd/blob/b1eb4824eecf668b82bafe47304d772526d96d10/indexd/app.py#L27
+ - PRESIGNED_FENCE_URL=http://fence-service
volumes:
- ./Secrets/indexd_settings.py:/var/www/indexd/local_settings.py
- ./Secrets/indexd_creds.json:/var/www/indexd/creds.json
@@ -37,8 +44,13 @@ services:
retries: 3
depends_on:
- postgres
+ restart: unless-stopped
+ logging:
+ driver: "json-file"
+ options:
+ max-size: "10m"
fence-service:
- image: "quay.io/cdis/fence:2021.03"
+ image: quay.io/cdis/fence:2023.01 #"quay.io/cdis/fence:integration202211"
command: bash /var/www/fence/fence_setup.sh
container_name: fence-service
networks:
@@ -60,8 +72,13 @@ services:
depends_on:
- postgres
- arborist-service
+ restart: unless-stopped
+ logging:
+ driver: "json-file"
+ options:
+ max-size: "10m"
arborist-service:
- image: "quay.io/cdis/arborist:2021.03"
+ image: "quay.io/cdis/arborist:2022.12"
container_name: arborist-service
entrypoint: bash /go/src/github.com/uc-cdis/arborist/arborist_setup.sh
networks:
@@ -83,22 +100,43 @@ services:
retries: 10
depends_on:
- postgres
+ restart: unless-stopped
+ logging:
+ driver: "json-file"
+ options:
+ max-size: "10m"
metadata-service:
- image: "quay.io/cdis/metadata-service:2021.03"
+ # update image
+ image: quay.io/cdis/metadata-service:2.1.0 #"quay.io/cdis/metadata-service:1.8.0"
container_name: metadata-service
depends_on:
- postgres
+ volumes:
+ # To load:
+ # scripts/discovery_metadata.sh
+ # Also interesting:
+ # /env/bin/python /src/src/mds/populate.py --config /var/local/metadata-service/aggregate_config.json
+ - ./Secrets/metadata/aggregate_config.json:/var/local/metadata-service/aggregate_config.json
environment:
- DB_HOST=postgres
- DB_USER=metadata_user
- DB_PASSWORD=metadata_pass
- DB_DATABASE=metadata
+ - DEBUG=true
+ - USE_AGG_MDS=false
+ - AGG_MDS_NAMESPACE=default_namespace
+ - GEN3_ES_ENDPOINT=http://esproxy-service:9200
command: >
sh -c "/env/bin/alembic upgrade head && /env/bin/uvicorn --host 0.0.0.0 --port 80 mds.asgi:app --reload"
networks:
- devnet
+ restart: unless-stopped
+ logging:
+ driver: "json-file"
+ options:
+ max-size: "10m"
peregrine-service:
- image: "quay.io/cdis/peregrine:2021.03"
+ image: quay.io/cdis/peregrine:2023.01 #"quay.io/cdis/peregrine:2021.03"
container_name: peregrine-service
networks:
- devnet
@@ -110,7 +148,7 @@ services:
- ./scripts/peregrine_setup.sh:/peregrine_setup.sh
- ./datadictionary/gdcdictionary/schemas:/schemas_dir
environment: &env
- DICTIONARY_URL: https://s3.amazonaws.com/dictionary-artifacts/datadictionary/develop/schema.json
+ DICTIONARY_URL: https://aced-public.s3.us-west-2.amazonaws.com/aced-test.json
# PATH_TO_SCHEMA_DIR: /schemas_dir
REQUESTS_CA_BUNDLE: /etc/ssl/certs/ca-certificates.crt
healthcheck:
@@ -122,8 +160,13 @@ services:
depends_on:
- postgres
- sheepdog-service
+ restart: unless-stopped
+ logging:
+ driver: "json-file"
+ options:
+ max-size: "10m"
sheepdog-service:
- image: "quay.io/cdis/sheepdog:2021.03"
+ image: quay.io/cdis/sheepdog:2023.01 #"quay.io/cdis/sheepdog:2021.03"
command: bash /sheepdog_setup.sh
container_name: sheepdog-service
networks:
@@ -142,8 +185,13 @@ services:
retries: 5
depends_on:
- postgres
+ restart: unless-stopped
+ logging:
+ driver: "json-file"
+ options:
+ max-size: "10m"
guppy-service:
- image: "quay.io/cdis/guppy:2021.03"
+ image: quay.io/cdis/guppy:2023.01 #"quay.io/cdis/guppy:2021.03"
container_name: guppy-service
networks:
- devnet
@@ -159,13 +207,20 @@ services:
depends_on:
- arborist-service
- esproxy-service
+ restart: unless-stopped
+ logging:
+ driver: "json-file"
+ options:
+ max-size: "10m"
esproxy-service:
- image: quay.io/cdis/elasticsearch-oss:6.8.12
+ build: elasticsearch-mac
container_name: esproxy-service
environment:
- cluster.name=elasticsearch-cluster
- bootstrap.memory_lock=false
- - "ES_JAVA_OPTS=-Xms1g -Xmx1g"
+ # For apple silicon
+ - bootstrap.system_call_filter=false
+ - "ES_JAVA_OPTS=-Xms6g -Xmx6g"
entrypoint:
- /bin/bash
# mmapfs requires systemctl update - see https://www.elastic.co/guide/en/elasticsearch/reference/current/index-modules-store.html#mmapfs
@@ -193,8 +248,13 @@ services:
- 9300:9300
networks:
- devnet
+ restart: unless-stopped
+ logging:
+ driver: "json-file"
+ options:
+ max-size: "10m"
pidgin-service:
- image: "quay.io/cdis/pidgin:2021.03"
+ image: quay.io/cdis/pidgin:2023.01 #"quay.io/cdis/pidgin:2021.03"
container_name: pidgin-service
networks:
- devnet
@@ -207,10 +267,19 @@ services:
retries: 3
depends_on:
- peregrine-service
+ restart: unless-stopped
+ logging:
+ driver: "json-file"
+ options:
+ max-size: "10m"
portal-service:
- image: "quay.io/cdis/data-portal:2021.03"
+ image: quay.io/cdis/data-portal:2023.01 #"quay.io/cdis/data-portal:3.33.0" # 2021.03
container_name: portal-service
command: ["bash", "/var/www/data-portal/waitForContainers.sh"]
+ deploy:
+ resources:
+ reservations:
+ cpus: "3.0"
networks:
- devnet
volumes:
@@ -218,11 +287,15 @@ services:
- ./Secrets/gitops.json:/data-portal/data/config/gitops.json
- ./Secrets/gitops-logo.png:/data-portal/custom/logo/gitops-logo.png
- ./Secrets/gitops.png:/data-portal/custom/createdby/gitops.png
+ # BRH biomedical research hub graphics
+ # - ./Secrets/gitops-sponsors/createdby_50.png:/data-portal/custom/sponsors/gitops-sponsors/createdby_50.png
+ # - ./Secrets/gitops-sponsors/gen3_50.png:/data-portal/custom/sponsors/gitops-sponsors/gen3_50.png
+ # - ./Secrets/gitops-sponsors/gene_bgy.svg:/data-portal/custom/sponsors/gitops-sponsors/gene_bgy.svg
environment:
- NODE_ENV=dev
#- MOCK_STORE=true
- APP=gitops
- - GDC_SUBPATH=http://revproxy-service/api/v0/submission/
+ - GDC_SUBPATH=https://revproxy-service/api/v0/submission/
healthcheck:
test: ["CMD-SHELL", "curl -f http://localhost"]
interval: 60s
@@ -232,6 +305,11 @@ services:
- postgres
- peregrine-service
- sheepdog-service
+ restart: unless-stopped
+ logging:
+ driver: "json-file"
+ options:
+ max-size: "10m"
jupyter-service:
image: "quay.io/cdis/jupyter-slim:latest"
container_name: jupyter-service
@@ -244,16 +322,26 @@ services:
networks:
- devnet
environment:
- - FRAME_ANCESTORS=http://localhost
+ # - FRAME_ANCESTORS=https://aced-training.compbio.ohsu.edu
+ - FRAME_ANCESTORS=http://staging.aced-idp.org
+ restart: unless-stopped
+ logging:
+ driver: "json-file"
+ options:
+ max-size: "10m"
revproxy-service:
- image: "quay.io/cdis/nginx:2021.03"
+ image: quay.io/cdis/nginx:2023.01 #"quay.io/cdis/nginx:2021.03"
container_name: revproxy-service
networks:
- devnet
volumes:
- ./nginx.conf:/etc/nginx/nginx.conf
+ - ./minio.conf.staging:/etc/nginx/minio.conf
- ./Secrets/TLS/service.crt:/etc/nginx/ssl/nginx.crt
- ./Secrets/TLS/service.key:/etc/nginx/ssl/nginx.key
+ - certdata:/var/www/certbot/
+ - certs:/etc/letsencrypt/
+ - ./Secrets/.well-known:/etc/nginx/.well-known
ports:
- "80:80"
- "443:443"
@@ -270,42 +358,18 @@ services:
- fence-service
- portal-service
- pidgin-service
- tube-service:
- image: "quay.io/cdis/tube:2021.03"
- container_name: tube-service
- command: bash -c "while true; do sleep 5; done"
- networks:
- - devnet
- environment:
- - DICTIONARY_URL=https://s3.amazonaws.com/dictionary-artifacts/datadictionary/develop/schema.json
- - ES_URL=esproxy-service
- - ES_INDEX_NAME=etl
- - HADOOP_URL=hdfs://spark-service:9000
- - HADOOP_HOST=spark-service
+ certbot:
+ image: certbot/certbot
+ container_name: certbot
volumes:
- - ./Secrets/etl_creds.json:/usr/share/gen3/tube/creds.json
- - ./Secrets/etlMapping.yaml:/usr/share/gen3/tube/etlMapping.yaml
- - ./Secrets/user.yaml:/usr/share/gen3/tube/user.yaml
- - ./datadictionary:/tmp/datadictionary
+ - ./certbot/conf:/etc/letsencrypt
+ - ./certbot/www:/var/www/certbot
+ - certdata:/var/www/certbot/
+ - certs:/etc/letsencrypt/
depends_on:
- - postgres
- - esproxy-service
- - spark-service
- spark-service:
- image: "quay.io/cdis/gen3-spark:2021.03"
- container_name: spark-service
- command: bash -c "python run_config.py && hdfs namenode -format && hdfs --daemon start namenode && hdfs --daemon start datanode && yarn --daemon start resourcemanager && yarn --daemon start nodemanager && hdfs dfsadmin -safemode leave && hdfs dfs -mkdir /result && while true; do sleep 5; done"
- expose:
- - 22
- - 8030
- - 8031
- - 8032
- - 9000
- networks:
- - devnet
- environment:
- - HADOOP_URL=hdfs://0.0.0.0:9000
- - HADOOP_HOST=0.0.0.0
+ - revproxy-service
+ - portal-service
+ command: certonly --webroot --webroot-path=/var/www/certbot --email beckmanl@ohsu.edu --agree-tos --force-renewal --no-eff-email -d staging.aced-idp.org -d minio-default-staging.aced-idp.org -d minio-ohsu-staging.aced-idp.org -d minio-ucl-staging.aced-idp.org -d minio-manchester-staging.aced-idp.org -d minio-stanford-staging.aced-idp.org -d minio-default-console-staging.aced-idp.org -d minio-ohsu-console-staging.aced-idp.org -d minio-ucl-console-staging.aced-idp.org -d minio-manchester-console-staging.aced-idp.org -d minio-stanford-console-staging.aced-idp.org
kibana-service:
image: quay.io/cdis/kibana-oss:6.5.4
container_name: kibana-service
@@ -323,3 +387,5 @@ networks:
volumes:
psqldata:
esdata:
+ certdata:
+ certs:
diff --git a/docs/.env-example b/docs/.env-example
new file mode 100644
index 00000000..35d64d2f
--- /dev/null
+++ b/docs/.env-example
@@ -0,0 +1,2 @@
+MINIO_ROOT_USER=
+MINIO_ROOT_PASSWORD=
diff --git a/docs/New_Setup.md b/docs/New_Setup.md
new file mode 100644
index 00000000..4fe78d01
--- /dev/null
+++ b/docs/New_Setup.md
@@ -0,0 +1,151 @@
+These are the steps for setting up a feature/staging localhost
+
+1. Clone compose services checkout to feature/staging
+```sh
+git clone [ADDRESS OF feature/staging]
+git checkout feature/staging
+cd compose-services trainign
+```
+
+2. Change. Etl/dockerfile From python:3.10 to From python:3.9
+Assuming that you’re running python 3.9 locally
+
+3. If on a new machine the etc/hosts additions from https://github.com/ACED-IDP/compose-services-training/blob/feature/staging/docs/aced.md
+(it’s in 2 different places so you should be adding around 10 lines in total)
+```sh
+sudo vim /etl/hosts
+```
+
+4. Do a creds setup like this:
+```sh
+bash ./creds_setup.sh aced-training.compbio.ohsu.edu
+```
+
+5. Drag in all files from examples/Secrets to Secrets directory and overwrite all of the secrets directory files. The ones in examples/Secrets are more up to date
+
+6.
+```sh
+docker compose up -d
+```
+you should see the page with a working fence
+
+7. Generate a credentials.json file and save it to Secrets directory by clicking on the profile button and create API key button and download json file button
+
+8. Replace the 2 certs service.crt and service.key in your TLS folder
+With the ones provided from the development.aced-idp server this may require you to ssh into it, refer to step 11.
+
+9. Make a new program and project following this guide:
+https://github.com/uc-cdis/compose-services/blob/master/docs/using_the_commons.md#programs-and-projects
+
+But instead of using MyFirstProgram for the program name use aced and instead of using MyFirstProject for the project name follow the conventions in the command listed on step 10 for arguments project_code being STUDY and program_name being aced
+
+10. Execute the following command from the data_model directory within a virtual environment:
+
+```sh
+python3 -m venv venv
+source venv/bin/activate
+```
+
+ It uploads files into buckets:
+```sh
+nice -n 10 scripts/upload-files STUDY BUCKET &
+```
+ Load the meta data:
+```sh
+nice -n 10 python3 scripts/emitter.py data load --project_code STUDY --input_path output/STUDY/extractions --db_host localhost --program_name aced --sheepdog_creds_path ../compose-services-training/Secrets/sheepdog_creds.json
+```
+```txt
+STUDY BUCKET
+Alcoholism aced-ohsu
+Alzheimers aced-ucl
+Breast_Cancer aced-manchester
+Colon_Cancer aced-stanford
+Diabetes aced-ucl
+Lung_Cancer aced-manchester
+Prostate_Cancer aced-stanford
+```
+
+11. Ssh into staging via
+```sh
+ssh ubuntu@staging.aced-idp.org
+```
+
+For each copy to . (The current directory) Pay attention where The directory is in Staging.aced-idp.org.
+
+12. Make a new folder called data_model that is a sibling folder to compose-services-training this is the folder that all of the scp are going to
+
+13. Run all of these commands to copy all of the required Packages and files into data model:
+```sh
+scp ubuntu@staging.aced-idp.org:/home/ubuntu/data_model/output/all_studies-2.zip .
+```
+Unzip all of the studies from all studies and put them into the output folder.
+```sh
+scp -r ubuntu@staging.aced-idp.org:/home/ubuntu/data_model/output/dicom .
+```
+Where . Is data_model/output directory
+(This one takes some time and might be optional I’m not sure)
+
+```sh
+scp -r ubuntu@staging.aced-idp.org:/home/ubuntu/data_model/output/dna .
+```
+Where . Is data_model/output directory
+
+```sh
+scp -r ubuntu@staging.aced-idp.org:/home/ubuntu/data_model/output/gen3 .
+```
+Where . Is data_model/output directory
+
+```sh
+scp -r ubuntu@staging.aced-idp.org:/home/ubuntu/data_model/scripts .
+```
+Where . Is data_model directory
+
+```sh
+scp ubuntu@staging.aced-idp.org:/home/ubuntu/data_model/config.yaml .
+```
+Where . Is data_model directory
+
+These two package folders should go in data_model/scripts directory
+```sh
+scp -r ubuntu@staging.aced-idp.org:/home/ubuntu/data_model/venv/lib/python3.9/site-packages/jwt .
+
+scp -r ubuntu@staging.aced-idp.org:/home/ubuntu/data_model/venv/lib/python3.9/site-packages/pelican .
+```
+
+Change the name of data_model/output/STUDY/extract to
+data_model/output/STUDY/extractions (or look for where it is in the code and change it)
+
+Pip install all of the other required non-standard packages that will be shown in
+Emmitter.py at the top of the file
+
+14. Re-enable guppy services in Nginx.conf if you never disabled it disregard this step. L:147-149 or search for guppy and uncomment out the three lines where it is referenced
+
+
+15. Virtual environment for guppy setup this time downloading the etl/requirements.txt
+16.
+```sh
+python3 -m venv venv
+source venv/bin/activate
+pip install -r etl/requirements.txt
+```
+
+16.
+```sh
+bash guppy_setup.sh Secrets/credentials.json
+```
+This should not fail if it does fail consider doing a docker compose restart especially if it talks about permissions errors
+Also consider running this line
+
+```sh
+python3 -m pip uninstall certifi
+python3 -m pip install certifi
+```
+if you’re having problems with certification issues, you may have to do this step.
+
+you will have to wait until esproxy-service is running for guppy setup to work this can take some time. docker compose ps to check if it is running or not.
+
+17. Restart everything with
+```sh
+docker compose restart
+```
+
diff --git a/docs/aced.md b/docs/aced.md
new file mode 100644
index 00000000..33c4a522
--- /dev/null
+++ b/docs/aced.md
@@ -0,0 +1,861 @@
+- [ACED specific changes](#aced-specific-changes)
+ - [New Data Model Setup](#new-data-model-setup)
+ - [Fence (Authentication and Authorization)](#fence-authentication-and-authorization)
+ - [Setup](#setup)
+ - [Windmill's Auth Display](#windmills-auth-display)
+ - [Database Migrations](#database-migrations)
+ - [Authentication](#authentication)
+ - [Certs](#certs)
+ - [Customize Fence](#customize-fence)
+ - [Guppy and Kibana (Data)](#guppy-and-kibana-data)
+ - [Manual TSV Upload](#manual-tsv-upload)
+ - [`metadata` Command Upload](#metadata-command-upload)
+ - [API](#api)
+ - [Re-Enable Guppy](#re-enable-guppy)
+ - [Expose the Kibana Service](#expose-the-kibana-service)
+ - [Refactor the Spark and Tube Services](#refactor-the-spark-and-tube-services)
+ - [Minio (Local Object Store)](#minio-local-object-store)
+ - [Let's Setup Discovery](#lets-setup-discovery)
+ - [Enable jupyter notebooks](#enable-jupyter-notebooks)
+ - [Change the data dictionary](#change-the-data-dictionary)
+ - [Import data files from the study into gen3](#import-data-files-from-the-study-into-gen3)
+ - [Import metadata into gen3](#import-metadata-into-gen3)
+ - [Try some Graphql queries "Graph"](#try-some-graphql-queries-graph)
+ - [Test download](#test-download)
+- [Microservices Reference](#microservices-reference)
+
+# ACED specific changes
+
+> This document assumes you have completed setting up a 'stock' gen3 instance as described in https://github.com/uc-cdis/compose-services
+>
+> Now that you've completed this task, let's explore some ACED specific customizations.
+
+## New Data Model Setup
+
+0. Update `/etc/hosts` to have the following entries:
+
+```
+##
+# Host Database
+#
+# localhost is used to configure the loopback interface
+# when the system is booting. Do not change this entry.
+##
+127.0.0.1 localhost
+255.255.255.255 broadcasthost
+::1 localhost
+# Added by Docker Desktop
+# To allow the same kube context to work on the host and the container:
+127.0.0.1 kubernetes.docker.internal
+# End of section
+
+# ACED data portal
+127.0.0.1 aced-training.compbio.ohsu.edu
+
+# Minio storage
+127.0.0.1 minio.compbio.ohsu.edu
+127.0.0.1 minio-console.compbio.ohsu.edu
+127.0.0.1 minio-default.compbio.ohsu.edu
+127.0.0.1 minio-default-console.compbio.ohsu.edu
+127.0.0.1 minio-ohsu.compbio.ohsu.edu
+127.0.0.1 minio-ohsu-console.compbio.ohsu.edu
+127.0.0.1 minio-ucl.compbio.ohsu.edu
+127.0.0.1 minio-ucl-console.compbio.ohsu.edu
+127.0.0.1 minio-manchester.compbio.ohsu.edu
+127.0.0.1 minio-manchester-console.compbio.ohsu.edu
+127.0.0.1 minio-stanford.compbio.ohsu.edu
+127.0.0.1 minio-stanford-console.compbio.ohsu.edu
+```
+
+Docker resource limits are insufficient as default. Change to the following and restart Docker engine:
+- 7 CPUs
+- 32.5 GB Memory
+- 2 GB Swap
+- 512 GB Disk image size
+
+
+1. Fetch code
+```sh
+git clone git@github.com:ACED-IDP/compose-services-training.git
+cd compose-services-training
+bash ./creds_setup.sh aced-training.compbio.ohsu.edu
+```
+
+From the staging environment, make the following changes to run locally.
+
+```sh
+
+diff --git a/docker-compose.yml b/docker-compose.yml
+index 06380cb..17b9e4c 100644
+--- a/docker-compose.yml
++++ b/docker-compose.yml
+@@ -278,12 +278,14 @@ services:
+ - devnet
+ volumes:
+ - ./nginx.conf:/etc/nginx/nginx.conf
+- - ./minio.conf.staging:/etc/nginx/minio.conf
+ - ./Secrets/TLS/service.crt:/etc/nginx/ssl/nginx.crt
+ - ./Secrets/TLS/service.key:/etc/nginx/ssl/nginx.key
+- - certdata:/var/www/certbot/
+- - certs:/etc/letsencrypt/
+ - ./Secrets/.well-known:/etc/nginx/.well-known
++ # use `minio.conf` for local desktop dev
++ - ./minio.conf:/etc/nginx/minio.conf
++ # comment out for local desktop dev
++# - certdata:/var/www/certbot/
++# - certs:/etc/letsencrypt/
+ ports:
+ - "80:80"
+ - "443:443"
+
+
+diff --git a/nginx.conf b/nginx.conf
+index 2c12722..b7224fa 100644
+--- a/nginx.conf
++++ b/nginx.conf
+@@ -38,13 +38,13 @@ http {
+
+ listen 443 ssl;
+
+- # Original certificates
+- # ssl_certificate /etc/nginx/ssl/nginx.crt;
+- # ssl_certificate_key /etc/nginx/ssl/nginx.key;
++# Original certificates - uncomment for desktop development
++ ssl_certificate /etc/nginx/ssl/nginx.crt;
++ ssl_certificate_key /etc/nginx/ssl/nginx.key;
+
+- # New Certbot certificates
+- ssl_certificate /etc/letsencrypt/live/staging.aced-idp.org/fullchain.pem;
+- ssl_certificate_key /etc/letsencrypt/live/staging.aced-idp.org/privkey.pem;
++# # New Certbot certificates
++# ssl_certificate /etc/letsencrypt/live/staging.aced-idp.org/fullchain.pem;
++# ssl_certificate_key /etc/letsencrypt/live/staging.aced-idp.org/privkey.pem;
+
+ set $access_token "";
+ set $csrf_check "ok-tokenauth";
+
+```
+
+2. Copy contents of Secrets directory
+```sh
+cp example/Secrets
+```
+
+3. Copy google, microsoft `client` values from `development.aced-idp.org` in `fence-config.yml`:
+```yaml
+ google:
+ client_id: ''
+ client_secret: ''
+ microsoft:
+ client_id: ''
+ client_secret: ''
+```
+
+Logon to user-training.compbio.ohsu.edu/profile, create a credentials.json file, save it in Secrets
+
+4. Copy `user.yaml` from staging and replace e-mail address with the one logged into profile.
+
+5. Create new program and projects from the projects defined in user.yaml
+
+```sh
+ python scripts/load.py init --gen3_credentials_file ../compose-services/Secrets/credentials.json --user_path ../compose-services/Secrets/user.yaml
+```
+
+ https://aced-training.compbio.ohsu.edu/aced-Alcoholism
+
+
+6. Add to fence-config.yml (from Walsh):
+```yaml
+ALLOWED_DATA_UPLOAD_BUCKETS:
+ - aced-default
+ - aced-ohsu
+ - aced-ucl
+ - aced-manchester
+ - aced-stanford
+```
+
+7. Setup minio:
+```sh
+cat etl/setup-minio.sh | dc exec -T etl-service bash
+```
+
+9. See `data_model/scripts/gen3_etl.sh` for instructions on how to load the data:
+
+## Fence (Authentication and Authorization)
+
+> Fence is the main authentication mechanism for Gen3. Let's add some ACED specifics ...
+
+### Setup
+
+> Let's create an instance that should be hosted on aced-training.compbio.ohsu.edu
+
+You should have already seen this in the compose services docs for `bash ./creds_setup.sh aced-training.compbio.ohsu.edu`
+
+To test locally, update your `/etc/hosts` file.
+
+```
+# testing
+127.0.0.1 aced-training.compbio.ohsu.edu
+127.0.0.1 minio.compbio.ohsu.edu
+127.0.0.1 minio-console.compbio.ohsu.edu
+```
+### Windmill's Auth Display
+
+add to Secrets/gitops.json
+```json
+"showArboristAuthzOnProfile": true,
+"showFenceAuthzOnProfile": false
+```
+
+You should now see detailed authorization in the profile screen.
+
+
+
+### Database Migrations
+
+> The fence service will automatically apply a database migration on startup. We don't want to do that every time, let's turn that off
+
+In Secrets/fence-config.yaml
+```
+ENABLE_DB_MIGRATION: false
+```
+
+Now, when you re-start fence-service, you should see this message
+```
+fence-service | [2022-09-06 21:27:31,812][ fence][ INFO] NOT running database migration.
+```
+
+### Authentication
+
+> For testing, we won't configure OAuth, we will use a default user "test"
+
+* Let's turn off auth: Secrets/fence-config.yaml#L48-L49
+
+```yaml
+# if true, will automatically login a user with username "test"
+MOCK_AUTH: true
+```
+
+* Then adjust the user mapping to make the "test" user admin. In Secrets/user.yaml, change all occurrences of `username1@gmail.com` to `test`
+
+* Then restart fence.
+```sh
+docker-compose stop fence-service ; docker-compose rm -f fence-service ; docker-compose up -d fence-service ;
+```
+
+### Certs
+
+* If you are on an exastack node (or AWS instance) and want to use the official certificate, please ask for access:
+
+```
+https://ohsuitg-my.sharepoint.com/ XXXXX
+```
+
+* Once you have access, to install the certificate in gen3 follow these steps, assuming certs directory in ~/compbio-tls
+
+```sh
+cp /home/ubuntu/compbio-tls/compbio-tls/compbio.ohsu.edu-2022.interim-bundle.pem ./Secrets/TLS/service.crt
+cp /home/ubuntu/compbio-tls/compbio-tls/compbio.ohsu.edu-2022.key ./Secrets/TLS/service.key
+```
+
+### Customize Fence
+
+PRs to allow user to upload file to any of the buckets fence manages.
+https://github.com/uc-cdis/gen3sdk-python/pull/158
+https://github.com/uc-cdis/fence/pull/1048
+
+> These steps assume the PRs have _not_ been merged to main.
+
+* Clone fence in the compose-services dir. checkout the `alternate-data_upload_bucket` branch
+
+```sh
+git clone git@github.com:uc-cdis/fence.git
+```
+
+* Alter docker-compose.
+
+```diff
+- image: "quay.io/cdis/fence:2021.03"
++ #image: "quay.io/cdis/fence:2021.03"
++ build: fence
+```
+
+* update etl/requirements.txt to point to client side PR
+
+```diff
+# etl/requirements.txt
+-gen3
++# gen3
++https://github.com/ohsu-comp-bio/gen3sdk-python/archive/feat/alternate-data_upload_bucket.zip
+```
+
+## Guppy and Kibana (Data)
+
+* Per instructions, to disable guppy and kibana see https://github.com/uc-cdis/compose-services/blob/master/docs/setup.md#start-running-your-local-gen3-docker-compose-environment
+
+* Create a program (MyFirstProgram) and project (MyFirstProject). See https://github.com/uc-cdis/compose-services/blob/master/docs/using_the_commons.md#programs-and-projects
+
+* Activate the virtual environment and install required packages.
+
+```sh
+python3 -m venv venv
+source venv/bin/activate
+pip install -r etl/requirements.txt
+```
+
+* Let's generate some synthetic meta data
+
+```sh
+export TEST_DATA_PATH="$(pwd)/tests/fixtures/projects/MyFirstProject"
+mkdir -p "$TEST_DATA_PATH"
+
+docker run -it -v "${TEST_DATA_PATH}:/mnt/data" --rm --name=dsim --entrypoint=data-simulator quay.io/cdis/data-simulator:master simulate --url https://s3.amazonaws.com/dictionary-artifacts/datadictionary/develop/schema.json --path /mnt/data --program MyFirstProgram --project MyFirstProject --max_samples 10
+```
+* Let's run setup-minio.sh
+```sh
+docker compose exec -it etl-service bash
+#copy paste the contents of /etl/setup-minio.#!/bin/sh and press enter
+# then press control d to exit the container ssh
+```
+
+* Now, let's generate corresponding data files
+
+```sh
+./etl/generate_simulated_files --project_path tests/fixtures/projects/MyFirstProject --output_path tests/fixtures/projects/MyFirstProject/DATA
+
+./etl/file --gen3_credentials_file Secrets/credentials.json upload --project_path tests/fixtures/projects/MyFirstProject/
+```
+### Manual TSV Upload
+
+* Load the data manually by following the [instructions](https://gen3.org/resources/user/submit-data/#begin-metadata-tsv-submissions)
+(Note that the data we will be using is in JSON form.) This will be a good opportunity to discover data dependency order. Navigate to the "Submit Data" page. Load the data, following the hierarchy displayed in the "Toogle View"
+
+
+
+* When complete, the graph should look like this.
+
+
+
+### `metadata` Command Upload
+
+* Alternatively, to upload the metadata with the `metadata` command enter the following:
+
+```sh
+./etl/metadata --gen3_credentials_file Secrets/credentials.json load --project MyFirstProject --program MyFirstProgram --data_directory tests/fixtures/projects/MyFirstProject/
+```
+
+### API
+
+This may be a good time to examine the Gen3 API. You will need an API key first.
+
+
+
+For example, view the `metadata` script, where `credentials.json` is the key file downloaded above.
+
+List the schema entities:
+
+```sh
+./etl/metadata --gen3_credentials_file Secrets/credentials.json ls | jq .
+
+{
+"programs": {
+"MyFirstProgram": [
+"MyFirstProject",
+"MySecondProject"
+]
+},
+"entities": [
+"program",
+"project",
+"acknowledgement",
+"core_metadata_collection",
+"experiment",
+"keyword",
+"publication",
+"experimental_metadata",
+"submitted_copy_number",
+"case",
+"demographic",
+"diagnosis",
+"sample",
+"exposure",
+"family_history",
+"clinical_test",
+"aliquot",
+"slide",
+"treatment",
+"slide_image",
+"submitted_methylation",
+"read_group",
+"slide_count",
+"submitted_aligned_reads",
+"submitted_somatic_mutation",
+"submitted_unaligned_reads",
+"aligned_reads_index",
+"read_group_qc"
+]
+}
+
+```
+
+### Re-Enable Guppy
+
+* (Re)Read the [documentation](https://github.com/uc-cdis/compose-services/blob/master/docs/using_the_commons.md#configuring-guppy-for-exploration-page)
+* Rollback comment out of guppy in nginx.conf
+* Rollback comment out of kibana in docker-compose.yml
+* Run `bash guppy_setup.sh` - this should run the spark, tube service and launch the guppy service.
+
+### Expose the Kibana Service
+
+* Add the kibana path to `nginx.conf`
+
+```diff
++++ b/nginx.conf
+@@ -276,5 +276,17 @@ http {
+location /lw-workspace/ {
+return 302 /lw-workspace/proxy;
+}
++
++ location /kibana {
++ proxy_http_version 1.1;
++ proxy_set_header Upgrade $http_upgrade;
++ proxy_set_header Connection 'upgrade';
++ proxy_set_header Host $host;
++ proxy_cache_bypass $http_upgrade;
++
++ proxy_pass http://kibana-service:5601/;
++ rewrite ^/kibana/(.*)$ /$1 break;
++ }
++
+}
+}
+```
+
+* add the path to docker-compose
+
+```diff
+kibana-service:
+image: quay.io/cdis/kibana-oss:6.5.4
+container_name: kibana-service
+environment:
+- SERVER_NAME=kibana-service
+- ELASTICSEARCH_URL=http://esproxy-service:9200
++ - SERVER_BASEPATH=/kibana
+ports:
+- 5601:5601
+networks:
+
+```
+
+### Refactor the Spark and Tube Services
+
+* Update the docker compose to disable the spark and tube service
+
+```diff
+@@ -283,42 +287,42 @@ services:
+- fence-service
+- portal-service
+- pidgin-service
+- tube-service:
+- image: "quay.io/cdis/tube:2021.03"
+- container_name: tube-service
+- command: bash -c "while true; do sleep 5; done"
+- networks:
+- - devnet
+
+...
+```
+
+* Run the `tube-lite` replacement of spark and tube
+
+```sh
+./etl/tube_lite --credentials_path Secrets/credentials.json --elastic http://localhost:9200
+```
+
+* Alter `guppy-setup.sh` to run the tube_lite
+
+```diff
+diff --git a/guppy_setup.sh b/guppy_setup.sh
+index 559668d..5081eb1 100644
+--- a/guppy_setup.sh
++++ b/guppy_setup.sh
+@@ -1,16 +1,9 @@
+#!/bin/bash
+# Script to create and re-create es indices and setup guppy
+
+-sleep 2
+-docker exec esproxy-service curl -X DELETE http://localhost:9200/etl_0
+-sleep 2
+-docker exec esproxy-service curl -X DELETE http://localhost:9200/file_0
+-sleep 2
+-docker exec esproxy-service curl -X DELETE http://localhost:9200/file-array-config_0
+-sleep 2
+-docker exec esproxy-service curl -X DELETE http://localhost:9200/etl-array-config_0
+-sleep 2
+-docker exec tube-service bash -c "python run_config.py && python run_etl.py"
+-
++docker exec esproxy-service curl -X DELETE http://localhost:9200/gen3.aced.*
+docker container stop guppy-service
++
++./etl/tube_lite --credentials_path $1 --elastic http://localhost:9200
++
+docker container start guppy-service
+
+```
+
+* Examine the results using [kibana](http://localhost:5601/app/kibana#/dev_tools/console?_g=())
+
+
+
+* Examine the results in the [portal](http://localhost/files)
+
+
+
+## Minio (Local Object Store)
+
+Root credentials for the Minio consoles are stored within environmental variables defined in the `.env` file. Docker passes these values to the Minio services upon the `docker-compose up` command. If the values are changed simply rerun `docker-compose up` and the updated values will be passed to the Minio services.
+
+* Add the following to `/etc/hosts` for local host testing.
+
+```
+127.0.0.1 minio-default.compbio.ohsu.edu
+127.0.0.1 minio-default-console.compbio.ohsu.edu
+127.0.0.1 minio-ohsu.compbio.ohsu.edu
+127.0.0.1 minio-ohsu-console.compbio.ohsu.edu
+127.0.0.1 minio-ucl.compbio.ohsu.edu
+127.0.0.1 minio-ucl-console.compbio.ohsu.edu
+127.0.0.1 minio-manchester.compbio.ohsu.edu
+127.0.0.1 minio-manchester-console.compbio.ohsu.edu
+127.0.0.1 minio-stanford.compbio.ohsu.edu
+127.0.0.1 minio-stanford-console.compbio.ohsu.edu
+```
+
+* Add the minio.conf file to the revproxy-service
+
+```diff
+$ git diff docker-compose.yml
+diff --git a/docker-compose.yml b/docker-compose.yml
+index 62c536d..0a0f03f 100644
+--- a/docker-compose.yml
++++ b/docker-compose.yml
+@@ -269,6 +269,7 @@ services:
+- devnet
+volumes:
+- ./nginx.conf:/etc/nginx/nginx.conf
++ - ./minio.conf:/etc/nginx/minio.conf
+- ./Secrets/TLS/service.crt:/etc/nginx/ssl/nginx.crt
+- ./Secrets/TLS/service.key:/etc/nginx/ssl/nginx.key
+ports:
+```
+
+* Add the minio configuration to `docker-compose-override.yml`
+
+* Start the service
+
+```sh
+dc up -d ; dc logs -f minio-default
+```
+
+* Setup minio
+```yaml
+cat etl/setup-minio.sh | docker exec -i etl-service bash
+
+```
+
+* Examine logs
+
+```sh
+$ dc logs minio-default
+minio1-service | Formatting 1st pool, 1 set(s), 2 drives per set.
+minio1-service | WARNING: Host minio1-service:9000 has more than 1 drives of set. A host failure will result in data becoming unavailable.
+minio1-service | MinIO Object Storage Server
+minio1-service | Copyright: 2015-2022 MinIO, Inc.
+minio1-service | License: GNU AGPLv3
+minio1-service | Version: RELEASE.2022-09-01T23-53-36Z (go1.18.5 linux/arm64)
+minio1-service |
+minio1-service | Status: 2 Online, 0 Offline.
+minio1-service | API: http://172.19.0.2:9000 http://127.0.0.1:9000
+minio1-service | Console: http://172.19.0.2:9001 http://127.0.0.1:9001
+minio1-service |
+minio1-service | Documentation: https://docs.min.io
+```
+
+* Verify connection
+
+* `curl -I http://minio-default.compbio.ohsu.edu/minio/health/live`
+* open http://minio-default-console.compbio.ohsu.edu
+* repeat for other minio-* servers
+
+* Enable fence URL signing
+* see AWS_CREDENTIALS, S3_BUCKETS in Secrets/fence-config.yml
+
+* Now, let's empty the project, then re-create the project end to end.
+
+```sh
+# empty the project
+./etl/metadata --gen3_credentials_file Secrets/credentials.json empty --program MyFirstProgram --project MyFirstProject
+# upload the data files
+./etl/file --gen3_credentials_file Secrets/credentials.json upload --project_path tests/fixtures/projects/MyFirstProject/
+# upload the meta data
+./etl/metadata --gen3_credentials_file Secrets/credentials.json load --project MyFirstProject --program MyFirstProgram --data_directory tests/fixtures/projects/MyFirstProject/
+# re-create the elastic search indices
+bash ./guppy_setup.sh Secrets/credentials.json
+
+```
+
+## Let's Setup Discovery
+
+* Update the metadata service image
+
+```diff
+metadata-service:
+- image: "quay.io/cdis/metadata-service:2021.03"
++ image: "quay.io/cdis/metadata-service:1.8.0"
+container_name: metadata-service
+depends_on:
+- postgres
++ volumes:
++ # /env/bin/python /src/src/mds/populate.py --config /var/local/metadata-service/aggregate_config.json
++ - ./Secrets/metadata/aggregate_config.json:/var/local/metadata-service/aggregate_config.json
+environment:
+- DB_HOST=postgres
+- DB_USER=metadata_user
+- DB_PASSWORD=metadata_pass
+- DB_DATABASE=metadata
++ - USE_AGG_MDS=true
++ - GEN3_ES_ENDPOINT=http://esproxy-service:9200
+command: >
+```
+
+* Update gitops.json
+
+See example/Secrets
+
+## Enable jupyter notebooks
+
+* All we need to do is specify trusted domain for the jupyter notebook in docker-compose.yml
+
+```diff
+environment:
+- - FRAME_ANCESTORS=http://localhost
++ - FRAME_ANCESTORS=http://aced-training.compbio.ohsu.edu
+revproxy-service:
+```
+
+## Change the data dictionary
+
+* Update the data dictionary. In this case we've updated it from the data_dictionary repo.
+ * Note: This is not the only potential source of changes, see https://github.com/uc-cdis/compose-services/blob/master/docs/using_the_commons.md#changing-the-data-dictionary
+
+```commandline
+ cp ../data_model/DATA/gen3/*.yaml datadictionary/gdcdictionary/schemas/
+ # "compile" the yaml into a single json file
+ python3 etl/compile.py datadictionary/gdcdictionary/schemas/ --out datadictionary/
+ # copy the resulting datadictionary/aced.json to S3
+ # verify you can read https://aced-public.s3.us-west-2.amazonaws.com/aced.json
+```
+
+* Set the new datadictionary URL in docker-compose
+```commandline
+git diff docker-compose.yml
+diff --git a/docker-compose.yml b/docker-compose.yml
+index 7d139a0..6280178 100644
+--- a/docker-compose.yml
++++ b/docker-compose.yml
+@@ -125,7 +125,7 @@ services:
+ - ./scripts/peregrine_setup.sh:/peregrine_setup.sh
+ - ./datadictionary/gdcdictionary/schemas:/schemas_dir
+ environment: &env
+- DICTIONARY_URL: https://s3.amazonaws.com/dictionary-artifacts/datadictionary/develop/schema.json
++ DICTIONARY_URL: https://aced-public.s3.us-west-2.amazonaws.com/aced.json
+
+```
+
+* add custom graphql section to
+* Review services
+ * restart services as necessary, typically peregrine, sheepdog and portal
+ * comment out guppy in nginx.conf until we re-build guppy
+ * see https://github.com/uc-cdis/compose-services/blob/master/docs/using_the_commons.md#changing-the-data-dictionary*
+
+
+### Import data files from the study into gen3
+
+Make sure you have created all minio buckets.
+
+```
+cat etl/setup-minio.sh | docker exec -i etl-service bash
+
+>>> Added `default` successfully.
+Added `ohsu` successfully.
+Added `ucl` successfully.
+Added `manchester` successfully.
+Added `stanford` successfully.
+Bucket created successfully `default/aced-default`.
+Bucket created successfully `default/aced-public`.
+Bucket created successfully `ohsu/aced-ohsu`.
+Bucket created successfully `ucl/aced-ucl`.
+Bucket created successfully `manchester/aced-manchester`.
+Bucket created successfully `stanford/aced-stanford`.
+...
+Added user `test` successfully.
+...
+Policy `readwrite` is set on user `test`
+...
+Access permission for `default/aced-public` is set to `public`
+...
+Successfully added arn:minio:sqs::PRIMARY:webhook
+
+
+```
+
+```commandline
+ python3 etl/file --gen3_credentials_file Secrets/credentials.json upload-pfb --pfb_path output/research_study_Alzheimers.pfb > file-object_ids.ndjson
+```
+
+* This will upload the files into gen3's indexd & mino's default_bucket.
+* Indexd's object ids are saved to file-object_ids.ndjson
+
+### Import metadata into gen3
+
+```commandline
+ python3 etl/import_pfb.py
+Importing output/research_study_Alzheimers.pfb into MyFirstProgram-MyFirstProject project node d8668bf7-ed94-5fe2-bc2c-24a9ede57d4f
+2022-10-10 14:50:29.497221
+total_count 10000 Practitioner 2022-10-10 14:50:49.200088
+...
+total_count 450000 DocumentReference 2022-10-10 15:22:35.035110
+total_count 460000 DocumentReference 2022-10-10 15:23:22.449487
+Elapsed time: 0:32:58.921382 total_count 461412
+
+```
+
+### Try some Graphql queries "Graph"
+
+````commandline
+{
+ program {
+ id
+ name
+ projects {
+ id
+ code
+ ResearchStudies {
+ id
+ state
+ title
+ ResearchSubjects(first: 1) {
+ id
+ Patients {
+ id
+ Specimen {
+ Tasks {
+ id
+ DocumentReferences {
+ ga4gh_drs_uri
+ }
+ }
+ }
+ Observations {
+ id
+ category_coding_observation_category
+ code_text
+ valueString
+ valueQuantity_unit
+ valueQuantity_value
+ component_0_code_coding_loinc_org
+ component_0_code_text
+ component_0_valueQuantity_unit
+ component_0_valueQuantity_value
+ component_1_code_coding_loinc_org
+ component_1_code_text
+ component_1_valueQuantity_unit
+ component_1_valueQuantity_value
+ }
+ }
+ }
+ }
+ }
+ }
+}
+
+
+````
+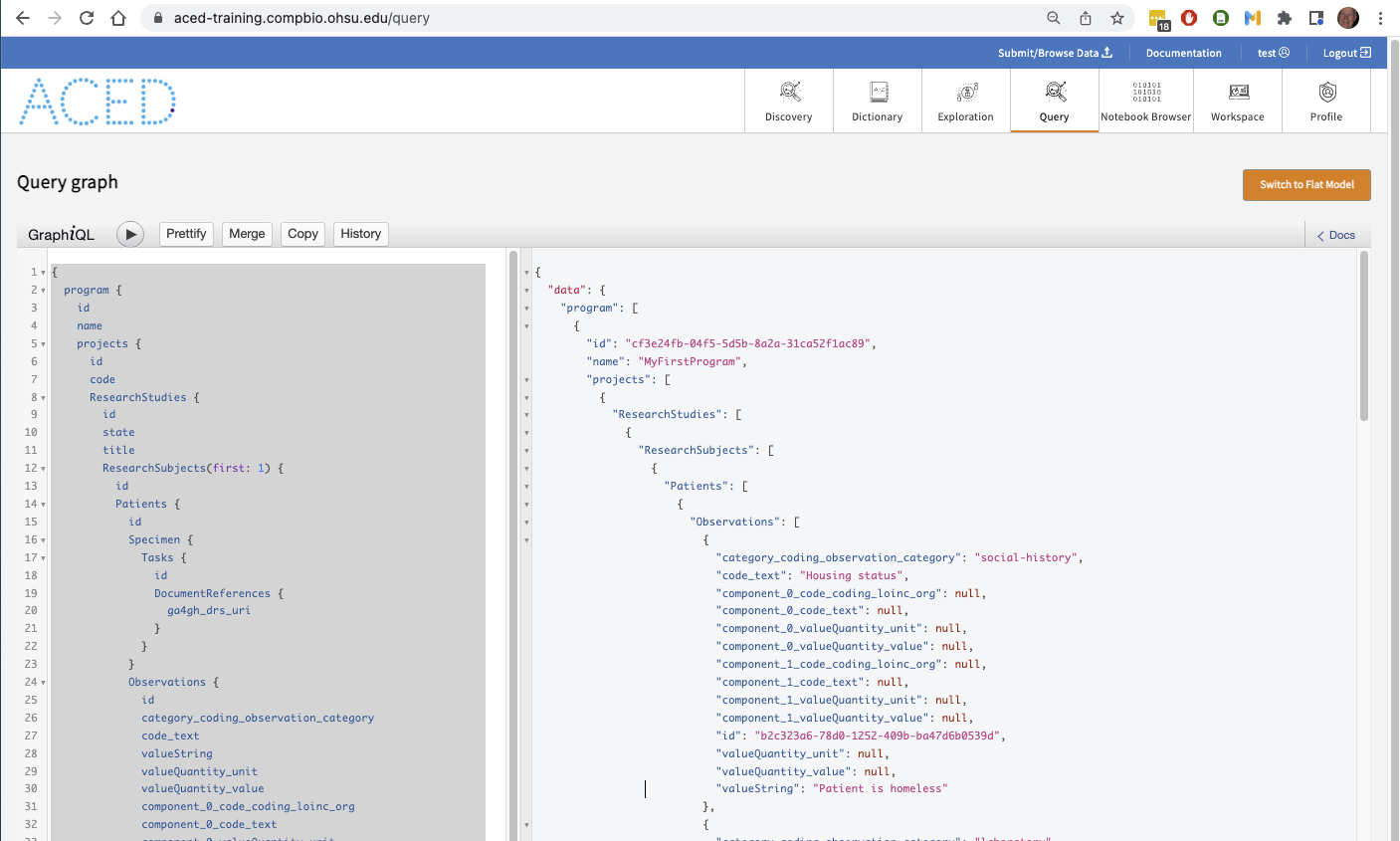
+
+```commandline
+{
+ DocumentReference(id: "6c37767e-6b9a-5376-bf93-b814d6a24fc1") {
+ id
+ object_id
+ content_attachment_md5
+ content_attachment_size
+ content_attachment_url
+ Tasks {
+ id
+ Specimen {
+ id
+ Patients {
+ id
+ }
+ }
+ }
+ }
+}
+```
+
+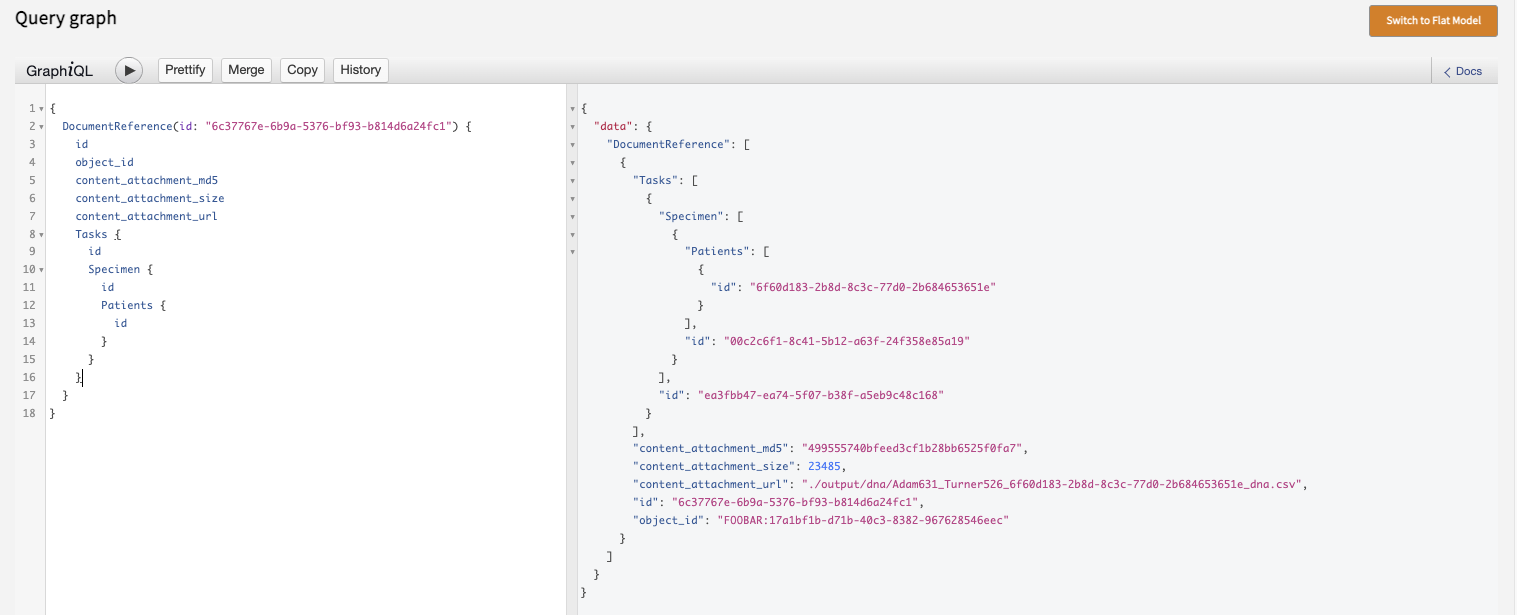
+
+### Test download
+
+```commandline
+$ python3 etl/file --gen3_credentials_file credentials-aced-training-local.json drs-download --did FOOBAR:17a1bf1b-d71b-40c3-8382-967628546eec --file_name /tmp/ttt
+2022-10-10 15:42:48,120 cli INFO : https://aced-training.compbio.ohsu.edu
+2022-10-10 15:42:48,487 download DEBUG : START
+2022-10-10 15:42:48,487 download DEBUG : ('process', 'https://minio-default.compbio.ohsu.edu/aced-default/FOOBAR%3A17a1bf1b-d71b-40c3-8382-967628546eec/output/dna/Adam631_Turner526_6f60d183-2b8d-8c3c-77d0-2b684653651e_dna.csv?X-Amz-Algorithm=AWS4-HMAC-SHA256&X-Amz-Credential=test%2F20221010%2Fus-east-1%2Fs3%2Faws4_request&X-Amz-Date=20221010T224248Z&X-Amz-Expires=3600&X-Amz-SignedHeaders=host&client_id=&user_id=1&username=test&X-Amz-Signature=f35fd6d6bc079845d812720c972804511df9f2aad9c739c6af629b64c9881240')
+2022-10-10 15:42:48,548 download DEBUG : ('get', 'https://minio-default.compbio.ohsu.edu/aced-default/FOOBAR%3A17a1bf1b-d71b-40c3-8382-967628546eec/output/dna/Adam631_Turner526_6f60d183-2b8d-8c3c-77d0-2b684653651e_dna.csv?X-Amz-Algorithm=AWS4-HMAC-SHA256&X-Amz-Credential=test%2F20221010%2Fus-east-1%2Fs3%2Faws4_request&X-Amz-Date=20221010T224248Z&X-Amz-Expires=3600&X-Amz-SignedHeaders=host&client_id=&user_id=1&username=test&X-Amz-Signature=f35fd6d6bc079845d812720c972804511df9f2aad9c739c6af629b64c9881240',
+
+)
+2022-10-10 15:42:48,550 download DEBUG : ('download', 4341499264, '/Users/walsbr/aced/compose-services/Adam631_Turner526_6f60d183-2b8d-8c3c-77d0-2b684653651e_dna.csv2q8u7a6g/Adam631_Turner526_6f60d183-2b8d-8c3c-77d0-2b684653651e_dna.csv.part0')
+2022-10-10 15:42:48,560 download DEBUG : ('md5', 4341499264, 'Adam631_Turner526_6f60d183-2b8d-8c3c-77d0-2b684653651e_dna.csv', '499555740bfeed3cf1b28bb6525f0fa7', b'SZVVdAv+7Tzxsou2Ul8Ppw==')
+2022-10-10 15:42:48,566 download INFO : 0.07946720800000007 seconds [True]
+
+```
+
+# Microservices Reference
+
+- `Sheepdog`: Handles the submitting and downloading of metadata to and from the GraphQL database.
+- `Peregrine`: Sends GraphQL queries to the PostgreSQL backend and returns results in JSON format.
+- `Fence`: Authenticates (AuthN) and authorizes (AuthZ) users. Generates tokens in `credentials.json`.
+- `Indexd`: Creates permanent ID's (GUID) for every newly submitted data object.
+- `Windmill`: Frontend for the exploration, submission, and downloading of data.
+
+# Let's Encrypt Certification
+
+To obtain SSL certificates for the `aced-idp.org` domains we'll run the following `certbot` command:
+
+```sh
+sudo certbot certonly --server https://acme-v02.api.letsencrypt.org/directory --manual --preferred-challenges dns -d 'aced-idp.org,*.aced-idp.org'
+```
+
+`certbot` may ask that one or more `TXT` records be deployed by the DNS provider in order to verify domain ownership. Proceed with the deployments and if all goes well the certificate files will be created in the `/etc/letsencrypt/live/aced-idp.org/` directory.
+
+`docker-compose` expects these certificate files to be in the `Secrets/TLS` directory so we'll copy and change ownership to the `ubuntu` user:
+
+```sh
+cd compose-services-training
+sudo cp /etc/letsencrypt/live/aced-idp.org/privkey.pem Secrets/TLS/service.key
+sudo cp /etc/letsencrypt/live/aced-idp.org/fullchain.pem Secrets/TLS/service.crt
+sudo chown ubuntu:ubuntu Secrets/TLS/service.{crt,key}
+```
+
+We're ready to start the project with `docker-compose`:
+
+```sh
+docker-compose up -d
+```
+
+Once all services are running the SSL certificates should be inspected to verify the correct domains and information were used.
+
+
+
+Make a note of the `Validity` value as a renewal will be required before that date to keep the HTTPS certification active for the domains. Unfortunately the manual configuration outlined above prevents us from using the automatic `certbot renew` command. Future work may focus on automating the `certbot` commands in a `cron` or `systemd` framework to allow for easier deployments.
diff --git a/docs/aws-ubuntu-setup.md b/docs/aws-ubuntu-setup.md
new file mode 100644
index 00000000..4e8a690e
--- /dev/null
+++ b/docs/aws-ubuntu-setup.md
@@ -0,0 +1,40 @@
+sudo apt-get remove docker docker-engine docker.io containerd runc
+
+sudo apt-get update
+sudo apt-get install \
+ ca-certificates \
+ curl \
+ gnupg \
+ lsb-release
+
+sudo mkdir -p /etc/apt/keyrings
+curl -fsSL https://download.docker.com/linux/ubuntu/gpg | sudo gpg --dearmor -o /etc/apt/keyrings/docker.gpg
+
+echo \
+ "deb [arch=$(dpkg --print-architecture) signed-by=/etc/apt/keyrings/docker.gpg] https://download.docker.com/linux/ubuntu \
+ $(lsb_release -cs) stable" | sudo tee /etc/apt/sources.list.d/docker.list > /dev/null
+
+sudo apt-get update
+
+sudo apt-get install docker-ce docker-ce-cli containerd.io docker-compose-plugin
+
+sudo docker run hello-world
+
+sudo groupadd docker
+sudo usermod -aG docker $USER
+
+sudo apt-get install jq
+sudo apt install unzip
+
+
+sudo apt install python3.9
+sudo apt update
+sudo apt install software-properties-common
+sudo add-apt-repository ppa:deadsnakes/ppa
+sudo apt install python3.9
+sudo apt-get install python3.9-dev python3.9-venv
+
+cd compose-services-training
+
+python3.9 -m venv venv
+
diff --git a/docs/graph-view-complete.png b/docs/graph-view-complete.png
new file mode 100644
index 00000000..cfb708eb
Binary files /dev/null and b/docs/graph-view-complete.png differ
diff --git a/docs/graph-view.png b/docs/graph-view.png
new file mode 100644
index 00000000..397ab7cd
Binary files /dev/null and b/docs/graph-view.png differ
diff --git a/docs/kibana.png b/docs/kibana.png
new file mode 100644
index 00000000..00410301
Binary files /dev/null and b/docs/kibana.png differ
diff --git a/docs/portal-tube-results.png b/docs/portal-tube-results.png
new file mode 100644
index 00000000..de5c1db3
Binary files /dev/null and b/docs/portal-tube-results.png differ
diff --git a/docs/profile-create-key.png b/docs/profile-create-key.png
new file mode 100644
index 00000000..55ec2bd2
Binary files /dev/null and b/docs/profile-create-key.png differ
diff --git a/docs/profile.png b/docs/profile.png
new file mode 100644
index 00000000..2e9fb0ab
Binary files /dev/null and b/docs/profile.png differ
diff --git a/elasticsearch-mac/Dockerfile b/elasticsearch-mac/Dockerfile
new file mode 100644
index 00000000..1769f179
--- /dev/null
+++ b/elasticsearch-mac/Dockerfile
@@ -0,0 +1,47 @@
+# Dockerfile-elastic.yml
+FROM arm64v8/openjdk:8-jdk-buster
+
+WORKDIR /usr
+
+# add a non root user
+RUN useradd -m elasticuser
+
+# Download extras
+RUN apt update
+RUN apt install -y curl nano
+
+# Download elasticsearch
+RUN curl -L -O https://artifacts.elastic.co/downloads/elasticsearch/elasticsearch-6.8.23.tar.gz
+RUN tar -xvf elasticsearch-6.8.23.tar.gz
+
+# Add config
+ADD elasticsearch.yml /usr/elasticsearch-6.8.23/config/elasticsearch.yml
+
+# Add scripts
+ADD docker-entrypoint.sh /usr/local/bin
+
+# create an empty directory where docker compose will mount a data volume
+RUN mkdir /usr/elasticsearch-6.8.23/data
+
+RUN ln -s /usr/elasticsearch-6.8.23 /usr/elasticsearch
+RUN ln -s /usr/elasticsearch-6.8.23 /usr/share/elasticsearch
+
+
+RUN chown -R elasticuser: .
+USER elasticuser
+
+ENV JAVA_HOME=/usr/local/openjdk-18/bin/java
+ENV discovery.type=single-node
+ENV xpack.security.enabled=false
+ENV xpack.ml.enabled=false
+
+RUN cd elasticsearch-6.8.23/bin
+RUN chown -R elasticuser: elasticsearch-6.8.23
+
+EXPOSE 9200 9300
+#RUN ls -1 /usr/elasticsearch
+#RUN ls -1 /usr/elasticsearch/config
+#RUN cat /usr/share/elasticsearch/config/elasticsearch.yml
+
+CMD cd /usr/elasticsearch-6.8.23/bin/ && ./elasticsearch
+
diff --git a/elasticsearch-mac/README.md b/elasticsearch-mac/README.md
new file mode 100644
index 00000000..0b971207
--- /dev/null
+++ b/elasticsearch-mac/README.md
@@ -0,0 +1,37 @@
+# elasticsearch mac
+
+The released build for elasticsearch `quay.io/cdis/elasticsearch-oss:6.8.12` tends to run poorly on mac silicon.
+
+Symptoms include:
+* cpu utilization at 100%
+* timeouts
+* not starting
+
+This custom build targets mac silicon. see https://stackoverflow.com/a/70713284
+
+
+## getting started
+
+Make this change to docker compose.
+
+```commandline
+ esproxy-service:
+- image: quay.io/cdis/elasticsearch-oss:6.8.12
++ # image: quay.io/cdis/elasticsearch-oss:6.8.12
++ build: elasticsearch-mac
+ container_name: esproxy-service
+ environment:
+ - cluster.name=elasticsearch-cluster
+
+```
+
+Stop and rm the exiting elastic search.
+
+```
+dc stop esproxy-service ; dc rm -f esproxy-service ;
+```
+
+Build and launch the new image
+```commandline
+dc build esproxy-service ; dc up -d esproxy-service
+```
diff --git a/elasticsearch-mac/docker-entrypoint.sh b/elasticsearch-mac/docker-entrypoint.sh
new file mode 100755
index 00000000..96abf2e0
--- /dev/null
+++ b/elasticsearch-mac/docker-entrypoint.sh
@@ -0,0 +1,106 @@
+#!/bin/bash
+set -e
+
+# Files created by Elasticsearch should always be group writable too
+umask 0002
+
+run_as_other_user_if_needed() {
+ if [[ "$(id -u)" == "0" ]]; then
+ # If running as root, drop to specified UID and run command
+ exec chroot --userspec=1000 / "${@}"
+ else
+ # Either we are running in Openshift with random uid and are a member of the root group
+ # or with a custom --user
+ exec "${@}"
+ fi
+}
+
+# Allow user specify custom CMD, maybe bin/elasticsearch itself
+# for example to directly specify `-E` style parameters for elasticsearch on k8s
+# or simply to run /bin/bash to check the image
+if [[ "$1" != "eswrapper" ]]; then
+ if [[ "$(id -u)" == "0" ]] && [[ "$1" == *elasticsearch* ]]; then
+ # centos:7 chroot doesn't have `--skip-chdir` option and
+ # changes our CWD. Rewrite CMD args to replace $1 with
+ # `elasticsearch` explicitly, so that we are backwards
+ # compatible with the docs from previous Elasticsearch
+ # versions<6 and configuration option D:
+ # https://www.elastic.co/guide/en/elasticsearch/reference/5.6/docker.html#_d_override_the_image_8217_s_default_ulink_url_https_docs_docker_com_engine_reference_run_cmd_default_command_or_options_cmd_ulink
+ # Without this, user could specify `elasticsearch -E x.y=z` but not
+ # `bin/elasticsearch -E x.y=z`
+ set -- "elasticsearch" "${@:2}"
+ # Use chroot to switch to UID 1000
+ exec chroot --userspec=1000 / "$@"
+ else
+ # User probably wants to exec something else, like /bin/bash
+ exec "$@"
+ fi
+fi
+
+# Parse Docker env vars to customize Elasticsearch
+#
+# e.g. Setting the env var cluster.name=testcluster
+#
+# will cause Elasticsearch to be invoked with -Ecluster.name=testcluster
+#
+# see https://www.elastic.co/guide/en/elasticsearch/reference/current/settings.html#_setting_default_settings
+
+declare -a es_opts
+
+while IFS='=' read -r envvar_key envvar_value
+do
+ # Elasticsearch env vars need to have at least two dot separated lowercase words, e.g. `cluster.name`
+ if [[ "$envvar_key" =~ ^[a-z_]+\.[a-z_]+ ]]; then
+ if [[ ! -z $envvar_value ]]; then
+ es_opt="-E${envvar_key}=${envvar_value}"
+ es_opts+=("${es_opt}")
+ fi
+ fi
+done < <(env)
+
+# The virtual file /proc/self/cgroup should list the current cgroup
+# membership. For each hierarchy, you can follow the cgroup path from
+# this file to the cgroup filesystem (usually /sys/fs/cgroup/) and
+# introspect the statistics for the cgroup for the given
+# hierarchy. Alas, Docker breaks this by mounting the container
+# statistics at the root while leaving the cgroup paths as the actual
+# paths. Therefore, Elasticsearch provides a mechanism to override
+# reading the cgroup path from /proc/self/cgroup and instead uses the
+# cgroup path defined the JVM system property
+# es.cgroups.hierarchy.override. Therefore, we set this value here so
+# that cgroup statistics are available for the container this process
+# will run in.
+export ES_JAVA_OPTS="-Des.cgroups.hierarchy.override=/ $ES_JAVA_OPTS"
+
+# Determine if x-pack is enabled
+if bin/elasticsearch-plugin list -s | grep -q x-pack; then
+ # Setting ELASTIC_PASSWORD is mandatory on the *first* node (unless
+ # LDAP is used). As we have no way of knowing if this is the first
+ # node at this step, we can't enforce the presence of this env
+ # var.
+ if [[ -n "$ELASTIC_PASSWORD" ]]; then
+ [[ -f config/elasticsearch.keystore ]] || run_as_other_user_if_needed "bin/elasticsearch-keystore" "create"
+ run_as_other_user_if_needed echo "$ELASTIC_PASSWORD" | bin/elasticsearch-keystore add -x 'bootstrap.password'
+ fi
+
+ # ALLOW_INSECURE_DEFAULT_TLS_CERT=true permits the use of a
+ # pre-bundled self signed cert for transport TLS.
+ # This should be used strictly on non-production environments.
+ if [[ "$ALLOW_INSECURE_DEFAULT_TLS_CERT" == "true" ]]; then
+ es_opts+=( '-Expack.security.authc.token.enabled=false'
+ '-Expack.ssl.verification_mode=certificate'
+ '-Expack.ssl.key=x-pack/node01/node01.key'
+ '-Expack.ssl.certificate=x-pack/node01/node01.crt'
+ '-Expack.ssl.certificate_authorities=x-pack/ca/ca.crt'
+ )
+ fi
+fi
+
+if [[ "$(id -u)" == "0" ]]; then
+ # If requested and running as root, mutate the ownership of bind-mounts
+ if [[ -n "$TAKE_FILE_OWNERSHIP" ]]; then
+ chown -R 1000:0 /usr/share/elasticsearch/{data,logs}
+ fi
+fi
+
+run_as_other_user_if_needed /usr/share/elasticsearch/bin/elasticsearch "${es_opts[@]}"
\ No newline at end of file
diff --git a/elasticsearch-mac/elasticsearch.yml b/elasticsearch-mac/elasticsearch.yml
new file mode 100644
index 00000000..ab9417f9
--- /dev/null
+++ b/elasticsearch-mac/elasticsearch.yml
@@ -0,0 +1,6 @@
+# elasticsearch.yml
+xpack.security.enabled: false
+xpack.ml.enabled: false
+
+http.host: 0.0.0.0
+transport.host: 0.0.0.0
diff --git a/etl/Dockerfile b/etl/Dockerfile
new file mode 100644
index 00000000..1b4e753d
--- /dev/null
+++ b/etl/Dockerfile
@@ -0,0 +1,16 @@
+# set base image (host OS)
+FROM python:3.9
+
+# set the working directory in the container
+WORKDIR /etl
+
+# copy the dependencies file to the working directory
+COPY requirements.txt .
+
+# install dependencies
+RUN pip install -r requirements.txt
+
+
+RUN wget https://dl.min.io/client/mc/release/linux-amd64/mc
+RUN chmod +x mc
+RUN mv mc /usr/local/bin
diff --git a/etl/README-aws.md b/etl/README-aws.md
new file mode 100644
index 00000000..78fa5421
--- /dev/null
+++ b/etl/README-aws.md
@@ -0,0 +1,39 @@
+cat >> ~./bashrc
+alias dc='docker compose'
+export PS1="\e[0;32m[\u@staging \W]\$ \e[0m"
+
+
+sudo apt-get remove docker docker-engine docker.io containerd runc
+sudo apt-get update
+sudo apt-get install ca-certificates curl gnupg lsb-release
+sudo mkdir -p /etc/apt/keyrings
+curl -fsSL https://download.docker.com/linux/ubuntu/gpg | sudo gpg --dearmor -o /etc/apt/keyrings/docker.gpg
+echo "deb [arch=$(dpkg --print-architecture) signed-by=/etc/apt/keyrings/docker.gpg] https://download.docker.com/linux/ubuntu \
+$(lsb_release -cs) stable" | sudo tee /etc/apt/sources.list.d/docker.list > /dev/null
+sudo apt-get update
+sudo apt-get install docker-ce docker-ce-cli containerd.io docker-compose-plugin
+sudo usermod -aG docker $USER
+
+
+sudo apt update
+sudo apt install software-properties-common
+sudo add-apt-repository ppa:deadsnakes/ppa
+sudo apt install python3.9
+python3.9 --version
+sudo apt-get install python3.9-venv
+
+sudo apt install unzip
+sudo apt install jq
+
+
+
+alias h='history | grep $1'
+
+alias h='history | grep $1'
+alias dc='docker-compose'
+rs () {
+ docker-compose stop $1
+ docker-compose rm -f $1
+ docker-compose up -d $1
+ docker-compose logs -f $1
+}
\ No newline at end of file
diff --git a/etl/README.md b/etl/README.md
new file mode 100644
index 00000000..fdf9eb53
--- /dev/null
+++ b/etl/README.md
@@ -0,0 +1,43 @@
+# etl
+
+A collections of scripts to automate Gen3 metadata processing.
+
+
+## metadata
+
+```commandline
+
+Usage: metadata [OPTIONS] COMMAND [ARGS]...
+
+ Metadata loader.
+
+Options:
+ --gen3_credentials_file TEXT API credentials file downloaded from gen3
+ profile. [default: credentials.json]
+
+ --help Show this message and exit.
+
+Commands:
+ drop-program Drops empty program
+ drop-project Drops empty project
+ empty Empties project, deletes all metadata.
+ load Loads metadata into project
+ ls Introspects schema and returns types in order.
+
+```
+
+
+## Some useful shortcuts
+
+```commandline
+
+cat etl/truncate_imported_tables.sql | dc exec -T postgres psql -U postgres
+
+dc stop peregrine-service ; dc rm -f peregrine-service ; dc up -d peregrine-servhistoryice
+dc stop sheepdog-service ; dc rm -f sheepdog-service ; dc up -d sheepdog-service
+
+dc stop portal-service ; dc rm -f portal-service ; dc up -d portal-service
+dc logs -f portal-service
+
+
+```
\ No newline at end of file
diff --git a/etl/__init__.py b/etl/__init__.py
new file mode 100644
index 00000000..e69de29b
diff --git a/etl/compile.py b/etl/compile.py
new file mode 100644
index 00000000..49a6c094
--- /dev/null
+++ b/etl/compile.py
@@ -0,0 +1,27 @@
+"""Converts datadictionary yaml files to json schema"""
+
+
+from pathlib import Path
+
+import click
+from dictionaryutils import dump_schemas_from_dir
+import json
+import os
+
+
+@click.command()
+@click.argument('schema')
+@click.option('--out', default="aced.json", help="Output path")
+def convert(schema, out):
+ """Converts yaml files to json schema"""
+ if os.path.isdir(out):
+ out = os.path.join(out, "aced.json")
+ schema = Path(schema)
+ assert schema.is_dir(), f"{schema} should be a path"
+ click.echo(f"Writing schema into {out}...")
+ with open(out, "w") as f:
+ json.dump(dump_schemas_from_dir(schema), f)
+
+
+if __name__ == '__main__':
+ convert()
diff --git a/etl/dictionary.py b/etl/dictionary.py
new file mode 100644
index 00000000..3227dcf4
--- /dev/null
+++ b/etl/dictionary.py
@@ -0,0 +1,190 @@
+# see https://github.com/uc-cdis/pelican/blob/master/pelican/dictionary.py
+
+import itertools
+from collections import defaultdict
+
+from dictionaryutils import DataDictionary, dictionary
+
+
+def init_dictionary(url):
+ d = DataDictionary(url=url)
+ dictionary.init(d)
+ # the gdcdatamodel expects dictionary initiated on load, so this can't be
+ # imported on module level
+ from gdcdatamodel import models as md
+
+ return d, md
+
+
+class DataDictionaryTraversal:
+ def __init__(self, model):
+ self.model = model
+
+ def get_nodes(self):
+ return self.model.Node.__subclasses__()
+
+ def get_edges(self):
+ return self.model.Edge.__subclasses__()
+
+ def get_node_table_by_label(self):
+ nodes = self.get_nodes()
+ node_tables = {str(node.label): node.__tablename__ for node in nodes}
+ return node_tables
+
+ def get_node_label_by_table(self):
+ nodes = self.get_nodes()
+ node_tables = {node.__tablename__: str(node.label) for node in nodes}
+ return node_tables
+
+ def get_edge_table_by_labels(self):
+ edges = self.get_edges()
+ edge_tables = {
+ (
+ self.model.Node.get_subclass_named(edge.__src_class__).label,
+ self.model.Node.get_subclass_named(edge.__dst_class__).label,
+ ): edge.__tablename__
+ for edge in edges
+ }
+ return edge_tables
+
+ def get_edge_labels_by_table(self):
+ edges = self.get_edges()
+ edge_tables = {
+ edge.__tablename__: {
+ "src": self.model.Node.get_subclass_named(edge.__src_class__).label,
+ "dst": self.model.Node.get_subclass_named(edge.__dst_class__).label,
+ }
+ for edge in edges
+ }
+ return edge_tables
+
+ def get_edges_by_node(self):
+ edges = self.get_edges()
+ it = defaultdict(list)
+
+ for edge in edges:
+ it[self.model.Node.get_subclass_named(edge.__src_class__).label].append(
+ edge.__tablename__
+ )
+
+ return it
+
+ def _get_bfs(self, node_name):
+ queue = [node_name]
+
+ visited = {}
+
+ r = []
+
+ while queue:
+ s = queue.pop(0)
+
+ node = self.model.Node.get_subclass(s).__name__
+ edges = self.model.Edge._get_edges_with_dst(node)
+
+ r.append(s)
+
+ for i in [
+ self.model.Node.get_subclass_named(e.__src_class__).get_label()
+ for e in edges
+ ]:
+ if i not in visited:
+ queue.append(i)
+ visited[i] = True
+
+ return r
+
+ def _topology_order(self, node_name, source_edges, target_class):
+ stack, path = [node_name], []
+
+ while stack:
+ vertex = stack[-1]
+
+ node_class = self.model.Node.get_subclass(vertex)
+ assert (
+ node_class
+ ), f"Node name '{vertex}' does not exist in the graph data model. Maybe you need to provide a 'root_node' (see Pelican documentation)?"
+ node = node_class.__name__
+ edges = getattr(self.model.Edge, source_edges)(node)
+
+ children = [
+ self.model.Node.get_subclass_named(getattr(e, target_class)).get_label()
+ for e in edges
+ ]
+
+ visited_children = [child for child in children if child not in path]
+
+ if not visited_children:
+ path.insert(0, vertex)
+ stack.pop()
+ else:
+ stack.append(visited_children[0])
+
+ return path
+
+ def get_upward_path(self, node_name):
+ return self._topology_order(node_name, "_get_edges_with_src", "__dst_class__")
+
+ def get_downward_path(self, node_name):
+ return self._topology_order(node_name, "_get_edges_with_dst", "__src_class__")
+
+ def full_traverse_path(self, node_name, extra_nodes=None, include_upward=False):
+ if include_upward:
+ upward_path = list(
+ zip(itertools.repeat(False), self.get_upward_path(node_name))
+ )
+ downward_path = list(
+ zip(itertools.repeat(True), self.get_downward_path(node_name))
+ )[1:]
+ if extra_nodes:
+ path = (
+ upward_path
+ + list(zip(itertools.repeat(True), extra_nodes))
+ + downward_path
+ )
+ else:
+ path = upward_path + downward_path
+ else:
+ downward_path = list(
+ zip(itertools.repeat(True), self.get_downward_path(node_name))
+ )
+ path = downward_path
+
+ return path
+
+
+def convert_to_node(x, is_base64):
+ obj = x["object"]
+ to_update = {}
+ for name, value in obj.items():
+ if value and is_base64[x["name"]][name]:
+ to_update[name] = decode_enum(value)
+
+ obj.update(to_update)
+
+ r = {
+ "created": datetime.now(),
+ "acl": json.dumps({}),
+ "_sysan": json.dumps({}),
+ "_props": json.dumps(obj),
+ "node_id": x["id"],
+ }
+
+ return r
+
+
+def convert_to_edge(x, edge_tables):
+ return [
+ (
+ edge_tables[(x["name"], i["dst_name"])],
+ {
+ "created": datetime.now(),
+ "acl": json.dumps({}),
+ "_sysan": json.dumps({}),
+ "_props": json.dumps({}),
+ "src_id": x["id"],
+ "dst_id": i["dst_id"],
+ },
+ )
+ for i in x["relations"]
+ ]
diff --git a/etl/download.py b/etl/download.py
new file mode 100644
index 00000000..a5a487a7
--- /dev/null
+++ b/etl/download.py
@@ -0,0 +1,137 @@
+import asyncio
+import base64
+import os.path
+import shutil
+import threading
+from dataclasses import dataclass
+from typing import List
+
+import aiofiles
+import aiohttp
+from tempfile import TemporaryDirectory
+import sys
+from urllib.parse import urlparse
+import hashlib
+import logging
+
+# logging.basicConfig(format="%(processName)s %(threadName)s: %(message)s", encoding='utf-8', level=logging.DEBUG)
+
+
+def _logger(name):
+ logger = logging.getLogger(name)
+ logger.setLevel(logging.DEBUG)
+ return logger
+
+
+async def get_content_length(url):
+ async with aiohttp.ClientSession() as session:
+ async with session.head(url) as request:
+ _logger(__name__).debug(('head', url, request))
+ return request.content_length
+
+
+def parts_generator(size, start=0, part_size=10 * 1024 ** 2):
+ while size - start > part_size:
+ yield start, start + part_size
+ start += part_size
+ yield start, size
+
+
+async def async_download(url, headers, save_path):
+ async with aiohttp.ClientSession(headers=headers) as session:
+ async with session.get(url) as request:
+ _logger(__name__).debug(('get', url, request))
+ file = await aiofiles.open(save_path, 'wb')
+ await file.write(await request.content.read())
+ _logger(__name__).debug(('download', threading.get_ident(), save_path))
+
+
+class Wrapped(object):
+ """Wrap the read() method and calculate MD5"""
+ def __init__(self, file_, md5_hash):
+ self._file = file_
+ self._md5_hash = md5_hash
+
+ def read(self, size):
+ buffer = self._file.read(size)
+ self._md5_hash.update(buffer)
+ return buffer
+
+ def __getattr__(self, attr):
+ return getattr(self._file, attr)
+
+
+async def process(url, size, expected_md5):
+ _logger(__name__).debug(('process', url))
+ filename = os.path.basename(urlparse(url).path)
+ tmp_dir = TemporaryDirectory(prefix=filename, dir=os.path.abspath('.'))
+ # size = await get_content_length(url)
+ tasks = []
+ file_parts = []
+ for number, sizes in enumerate(parts_generator(size)):
+ part_file_name = os.path.join(tmp_dir.name, f'{filename}.part{number}')
+ file_parts.append(part_file_name)
+ tasks.append(async_download(url, {'Range': f'bytes={sizes[0]}-{sizes[1]}'}, part_file_name))
+ await asyncio.gather(*tasks)
+ with open(filename, 'wb') as wfd:
+ md5_hash = hashlib.md5()
+ for f in file_parts:
+ with open(f, 'rb') as fd:
+ wrapped_fd = Wrapped(fd, md5_hash)
+ shutil.copyfileobj(wrapped_fd, wfd)
+ actual_md5 = md5_hash.hexdigest()
+ # compare calculated md5 vs expected
+ assert expected_md5 == actual_md5, f"Actual md5 {actual_md5} does not match expected {expected_md5}"
+ base64_md5 = base64.b64encode(bytes.fromhex(actual_md5))
+ _logger(__name__).debug(('md5', threading.get_ident(), filename, actual_md5, base64_md5))
+ return True
+
+
+@dataclass
+class DownloadURL(object):
+ """Information about the file to be downloaded."""
+ url: str
+ """Signed url."""
+ md5: str
+ """Needed for integrity check."""
+ size: int
+ """Needed for multi part download."""
+
+
+async def _download(urls: List[DownloadURL]):
+ """Download urls."""
+ results = await asyncio.gather(*[process(url.url, url.size, url.md5) for url in urls])
+ return results
+
+
+def download(urls: List[DownloadURL]):
+ """Setup async loop and download urls."""
+ import time
+ start_code = time.monotonic()
+ _logger(__name__).debug('START')
+ loop = asyncio.get_event_loop()
+ results = loop.run_until_complete(_download(urls))
+ _logger(__name__).info(f'{time.monotonic() - start_code} seconds {results}')
+
+
+# async def main():
+# # if len(sys.argv) <= 1:
+# # print('Add URLS')
+# # exit(1)
+# # urls = sys.argv[1:]
+# _logger(__name__).debug('BEFORE GATHER')
+# # ./etl/file --gen3_credentials_file credentials-aced-training-local.json get-index --did 40be30f4-4a21-47f1-ab64-745bf04cadd5
+# urls = [ 'https://minio-default.compbio.ohsu.edu/aced-default/40be30f4-4a21-47f1-ab64-745bf04cadd5/tmp/DATA/758278b648?X-Amz-Algorithm=AWS4-HMAC-SHA256&X-Amz-Credential=test%2F20220922%2Fus-east-1%2Fs3%2Faws4_request&X-Amz-Date=20220922T211915Z&X-Amz-Expires=3600&X-Amz-SignedHeaders=host&client_id=&user_id=1&username=test&X-Amz-Signature=5a5b83cc97dd372cee50c44331ead025c9eca8f0638c405ec228c9f3be590b3c']
+# md5 = 'b7c91b3c87cded71bdcd2684cdb268f7'
+# size = 10794
+# await asyncio.gather(*[process(url, size, md5) for url in urls])
+#
+#
+# if __name__ == '__main__':
+# import time
+#
+# start_code = time.monotonic()
+# _logger(__name__).debug('START')
+# loop = asyncio.get_event_loop()
+# loop.run_until_complete(main())
+# _logger(__name__).info(f'{time.monotonic() - start_code} seconds!')
diff --git a/etl/examples.gql b/etl/examples.gql
new file mode 100644
index 00000000..82219e97
--- /dev/null
+++ b/etl/examples.gql
@@ -0,0 +1,466 @@
+
+# #. Drill down hierarchy
+# {
+# program {
+# projects {
+# id
+# project_ResearchStudies {
+# title
+# study_ResearchSubjects {
+# individual_Patient {
+# id
+# multipleBirthBoolean
+# multipleBirthInteger
+# birthDate
+# deceasedBoolean
+# gender
+# subject_Specimen {
+# id
+# }
+# subject_Observations {
+# id
+# }
+# subject_DiagnosticReports {
+# id
+# }
+# }
+# }
+# }
+# }
+# }
+# }
+
+# # this works
+# {
+# Specimen {
+# subject_Patient {
+# id
+# }
+# }
+# }
+
+# query by parent edge
+# # Patients with maritalStatus
+# {
+# Patient(with_links_any:"maritalStatus_CodeableConcept") {
+# maritalStatus_CodeableConcept {
+# coding_0_code
+# }
+# }
+# }
+
+# # query by child edge
+# # Patients with specimens
+# {
+# Patient(with_links_any:"subject_Specimen") {
+# subject_Specimen {
+# id
+# }
+# }
+# }
+
+# # Patients with specimens and observations
+# {
+# Patient(with_links_any:"subject_Specimen", with_links:"subject_Observations", first:1) {
+# subject_Specimen(first:1) {
+# id
+# }
+# subject_Observations(first:1) {
+# id
+# status
+# state
+# }
+# }
+# }
+
+# #codes with observations
+# {
+# CodeableConcept(first:1, with_links_any:"code_Observations") {
+# id
+# coding_0_display
+# code_Observations{
+# id
+# }
+# }
+# }
+
+# #observations with codes
+# {
+# Observation(id:"e8f791cf-e2b2-6db2-b527-861f2fd57a93") {
+# id
+# code_CodeableConcept {
+# id
+# }
+# }
+# }
+
+
+
+# {
+# Observation(with_links_any:"subject_Patient") {
+# id
+# subject_Patient {id}
+# }
+# }
+
+# {
+# Patient(first:1) {
+# _subject_Observations_count
+# subject_Observations {
+# id
+# category {
+# id
+# }
+# }
+# }
+# }
+
+# {
+# Observation(id:"00b9e3b8-e877-b1e8-e12b-c88759053781") {
+# id
+# type
+# }
+# }
+
+# #"Total Cholesterol"
+# {
+# CodeableConcept(id:"f16d1c2b-5ff4-39c5-9152-054a6cb4c4ea") {
+# id
+# coding_0_display
+# }
+# }
+
+# {
+# Condition {
+# Patient
+# }
+# }
+
+# # condition with code
+# {
+# Condition(id:"5aaa2f7c-77fd-15fb-9dc9-030c325f7c64") {
+# code{id}
+# category{id}
+# }
+# }
+
+# # code with condition
+# {
+# CodeableConcept(with_links_any:"code_Conditions") {
+# id
+# coding_0_display
+# code_Conditions(first:1) {
+# id
+# }
+# }
+# }
+
+
+# {
+# DiagnosticReport(id:"242fa5aa-6187-c328-39e5-97bcca40490d") {
+# conclusion
+# encounter_Encounter {
+# _encounter_Observations_count
+# }
+# _result_Observation_count
+# _specimen_Specimen_count
+# _subject_Patient_count
+# }
+# }
+
+# {
+# Patient {
+# _subject_DiagnosticReports_count
+# }
+# }
+
+
+# {
+# CodeableConcept(id: "f12977c2-a689-3b08-876b-26bb05050586") {
+# id
+# coding_0_code
+# coding_0_system
+# clinicalStatus_Conditions(id:"c407e64d-f0ac-bbef-18a1-38890973ce43") {
+# id
+# code_CodeableConcept {
+# id
+# }
+# bodySite_CodeableConcept {
+# id
+# }
+# onsetDateTime
+# }
+
+# }
+# }
+
+# {
+# CodeableConcept(id: "c4fbc26f-c4bd-3406-9427-c195084059a6") {
+# id
+# coding_0_code
+# coding_0_display
+# code_Conditions(id:"c407e64d-f0ac-bbef-18a1-38890973ce43") {
+# id
+# }
+# }
+# }
+
+
+# THIS WORKS:get a document reference, and it's code
+# {
+# DocumentReference(first:1) {
+# id
+# type_CodeableConcept {
+# id
+# coding_0_display
+# }
+# category_CodeableConcept {
+# id
+# coding_0_display
+# }
+# }
+# }
+# returns
+# DocumentReference "id": 990de556-dfb4-45c4-f119-73c979720201"
+# CodeableConcept "id": "6e1f3684-e1cd-39fe-b4a5-cb1119bbaae8"
+
+# {
+# DocumentReference(first:1, offset:0) {
+# id
+# project_id
+# object_id
+# type_CodeableConcept {
+# coding_0_display
+# }
+# subject_Patient {
+# id
+# }
+# content_0_attachment_url
+# }
+# }
+
+# # THIS FAILS:
+# {
+# CodeableConcept(id: "6e1f3684-e1cd-39fe-b4a5-cb1119bbaae8") {
+# id
+# category_DocumentReferences(id: "990de556-dfb4-45c4-f119-73c979720201"){id}
+# }
+# }
+# # should return DocumentReference "id": "990de556-dfb4-45c4-f119-73c979720201" returns []
+
+# {
+# CodeableConcept(id: "6e1f3684-e1cd-39fe-b4a5-cb1119bbaae8") {
+# id
+# coding_0_display
+# category_DocumentReferences(id:"990de556-dfb4-45c4-f119-73c979720201"){
+# id
+# }
+# }
+# }
+
+
+# # actual codable concept
+# {
+# CodeableConcept(id: "14dbf741-d496-372e-8a6e-0a66f4f08916") {
+# id
+# _type_DocumentReferences_count
+# }
+# }
+# # returns 48354 _type_DocumentReferences_count
+
+# # get a diagnostic report
+# {
+# DiagnosticReport(first:1) {
+# id
+# category_CodeableConcept{id, coding_0_display}
+# code_CodeableConcept{id, coding_0_display}
+# }
+# }
+# # CodeableConcept "14dbf741-d496-372e-8a6e-0a66f4f08916"
+# # DiagnosticReport "00074d49-3a71-27d0-1e08-2418a566fed5"
+
+# # this works
+# {
+# CodeableConcept(id:"14dbf741-d496-372e-8a6e-0a66f4f08916")
+# {
+# code_DiagnosticReports(id: "00074d49-3a71-27d0-1e08-2418a566fed5") {
+# id
+# }
+# category_DiagnosticReports(id: "00074d49-3a71-27d0-1e08-2418a566fed5") {
+# id
+# }
+# }
+# }
+
+
+# {
+# node(id:"14dbf741-d496-372e-8a6e-0a66f4f08916") {
+# id
+# type
+# }
+# }
+
+# The psqlgraph information
+# {
+# "label": "DocumentReference_type_CodeableConcept",
+# "tablename": "edge_5942e705_DoDotyCoCo",
+# "srcclass": "Documentreference",
+# "dstclass": "Codeableconcept",
+# "srcdstassoc": "type_CodeableConcept",
+# "dstsrcassoc": "type_DocumentReferences",
+# "srctable": "node_DocumentReference",
+# "dsttable": "node_CodeableConcept"
+# }
+
+
+
+
+
+# {
+# Condition(id:"c407e64d-f0ac-bbef-18a1-38890973ce43") {
+# id
+# code_CodeableConcept(id: "c4fbc26f-c4bd-3406-9427-c195084059a6") {
+# id
+# }
+# }
+# }
+
+
+# #From parent, the Patient "6f60d183-2b8d-8c3c-77d0-2b684653651e"
+# # Retrieve the child, the Specimen "00c2c6f1-8c41-5b12-a63f-24f358e85a19"
+# {
+# Patient(id:"6f60d183-2b8d-8c3c-77d0-2b684653651e") {
+# id
+# subject_Specimen {
+# id
+# }
+# }
+# }
+
+#From child, the Specimen "00c2c6f1-8c41-5b12-a63f-24f358e85a19"
+# Retrieve the parent, the Patient "6f60d183-2b8d-8c3c-77d0-2b684653651e"
+# {
+# Specimen(id: "00c2c6f1-8c41-5b12-a63f-24f358e85a19") {
+# id
+# subject_Patient {
+# id
+# }
+# }
+# }
+
+
+#From child, the Specimen "00c2c6f1-8c41-5b12-a63f-24f358e85a19"
+# Retrieve the parent, the CodeableConcept
+# {
+# Specimen(id: "00c2c6f1-8c41-5b12-a63f-24f358e85a19") {
+# id
+# #returns empty list
+# type_CodeableConcept {
+# id
+# }
+# #returns empty list
+# condition_CodeableConcept {
+# id
+# }
+# }
+# }
+
+# # From the parent, the CodeableConcept
+# {
+# CodeableConcept(with_links_any: "type_Specimen") {
+# type_Specimen{id}
+# }
+# }
+
+# Works: child->parent
+# {
+# Condition(id:"47a56aed-fe69-df4c-3448-eaec0f0906c6") {
+# id
+# subject_Patient {
+# id # "6f60d183-2b8d-8c3c-77d0-2b684653651e"
+# }
+# }
+# }
+
+# # Works: parent->child
+# {
+# Patient(id: "6f60d183-2b8d-8c3c-77d0-2b684653651e") {
+# subject_Conditions(id:"47a56aed-fe69-df4c-3448-eaec0f0906c6") {
+# id
+# }
+# }
+# }
+
+
+# Fails: child->parent
+# {
+# Condition(id:"47a56aed-fe69-df4c-3448-eaec0f0906c6") {
+# id
+# #returns empty list
+# code_CodeableConcept {
+# id
+# }
+# #returns empty list
+# category_CodeableConcept {
+# id
+# }
+# }
+# }
+
+# # Works
+# {
+# CodeableConcept(id: "c11b07a0-7288-3973-9659-b4e2aeeefebe") {
+# code_Conditions(id: "47a56aed-fe69-df4c-3448-eaec0f0906c6") {
+# id
+# }
+# }
+# }
+
+
+# {
+# CodeableConcept(id: "55fa011b-3d22-3d9a-bf1d-a434810ff966") {
+# category_Conditions(id: "47a56aed-fe69-df4c-3448-eaec0f0906c6") {
+# id
+# }
+# }
+# }
+
+
+# # THIS WORKS:get a document reference, and it's code
+# {
+# DocumentReference(first:1) {
+# id
+# type_CodeableConcept {
+# id
+# coding_0_display
+# }
+# category_CodeableConcept {
+# id
+# coding_0_display
+# }
+# }
+# }
+
+#This fails
+# {
+# Procedure(id:"50b0c419-23ea-5ac2-426e-b5f4c4387cda") {
+# code_CodeableConcept {id}
+# category_CodeableConcept {id}
+# id
+# }
+# }
+
+
+
+{
+ Condition(first:1){
+ id
+ verificationStatus_coding_0_display
+ clinicalStatus_coding_0_display
+ code_coding_0_display
+ category_0_coding_0_display
+ }
+}
+
+
+
diff --git a/etl/file b/etl/file
new file mode 100755
index 00000000..db9a40ff
--- /dev/null
+++ b/etl/file
@@ -0,0 +1,309 @@
+#!/usr/bin/env python3
+import json
+import os
+import logging
+from pathlib import Path
+
+import click
+import fastavro
+import jwt
+import requests
+import urllib.parse
+
+from gen3.auth import Gen3Auth
+from gen3.file import Gen3File
+from gen3.index import Gen3Index
+
+from gen3.submission import Gen3Submission
+
+from cdislogging import get_logger as get_gen3_logger
+import base64
+
+log_fmt = "%(asctime)s %(name)s %(levelname)s : %(message)s"
+
+# set logging to warning, since gen3.submission logs a verbose INFO message on each call :-()
+logging.basicConfig(level=logging.WARNING, format=log_fmt)
+# set gen3's logger as well
+get_gen3_logger('__name__', log_level='warn', format=log_fmt)
+
+
+def get_logger_(name):
+ """Return logger with level set to info"""
+ logger = logging.getLogger(name)
+ logger.setLevel(logging.INFO)
+ return logger
+
+
+def extract_endpoint(gen3_credentials_file):
+ """Get base url of jwt issuer claim."""
+ with open(gen3_credentials_file) as input_stream:
+ api_key = json.load(input_stream)['api_key']
+ claims = jwt.decode(api_key, options={"verify_signature": False})
+ assert 'iss' in claims
+ return claims['iss'].replace('/user', '')
+
+
+@click.group()
+@click.option('--gen3_credentials_file', default='credentials.json', show_default=True,
+ help='API credentials file downloaded from gen3 profile.')
+@click.pass_context
+def cli(ctx, gen3_credentials_file):
+ """File uploader."""
+
+ endpoint = extract_endpoint(gen3_credentials_file)
+ get_logger_("cli").info(endpoint)
+ get_logger_("cli").debug(f"Read {gen3_credentials_file} endpoint {endpoint}")
+ auth = Gen3Auth(endpoint, refresh_file=gen3_credentials_file)
+ ctx.ensure_object(dict)
+ ctx.obj['submission_client'] = Gen3Submission(endpoint, auth)
+ ctx.obj['file_client'] = Gen3File(endpoint, auth)
+ ctx.obj['index_client'] = Gen3Index(endpoint, auth)
+
+ ctx.obj['endpoint'] = endpoint
+ ctx.obj['programs'] = [link.split('/')[-1] for link in ctx.obj['submission_client'].get_programs()['links']]
+
+
+def file_attributes(file_name):
+ """Calculate the hash and size."""
+ import hashlib
+
+ md5_hash = hashlib.md5()
+
+ with open(file_name, "rb") as f:
+ # Read and update hash in chunks of 4K
+ for byte_block in iter(lambda: f.read(4096), b""):
+ md5_hash.update(byte_block)
+
+ return md5_hash.hexdigest(), os.lstat(file_name).st_size
+
+
+@cli.command()
+@click.option('--did', default=None, show_default=True,
+ help='GUID from indexd')
+@click.pass_context
+def get_index(ctx, did):
+ """Read index."""
+ assert did, "Missing did (guid) parameter"
+ result = ctx.obj['index_client'].get_record(did)
+ print("record", result)
+ result = ctx.obj['file_client'].get_presigned_url(did)
+ print("presigned_url", result)
+
+
+
+@cli.command()
+@click.option('--bucket_name', default='aced-default', show_default=True,
+ help='File name to upload')
+@click.option('--pfb_path', required=True, default=None, show_default=True,
+ help='Path to pfb file')
+@click.option('--program', default='MyFirstProgram', show_default=True,
+ help='Gen3 program')
+@click.option('--project', default='MyFirstProject', show_default=True,
+ help='Gen3 project')
+@click.pass_context
+def upload_pfb(ctx, bucket_name, pfb_path, program, project):
+ """Filter DocumentReference records found in PFB to gen3 managed bucket, update hashes and size."""
+
+ index_client = ctx.obj['index_client']
+ file_client = ctx.obj['file_client']
+
+ """Finds DocumentReferences with attachment urls."""
+ for record in pfb_reader(pfb_path):
+ if record['name'] == 'DocumentReference':
+ if record['object']['content_attachment_url']:
+ object_name = record['object']['content_attachment_url'].lstrip('./')
+ content_attachment_md5 = record['object']['content_attachment_md5']
+ content_attachment_size = record['object']['content_attachment_size']
+ record_id = record['id']
+ # print(record['id'], object_name, content_attachment_md5, content_attachment_size)
+ assert 'name' in record, record.keys()
+ record_name = record['name']
+ assert 'submitter_id' in record['object'], record.keys()
+ submitter_id = record['object']['submitter_id']
+ document = file_client.upload_file(object_name, bucket=bucket_name)
+ assert 'guid' in document, document
+ assert 'url' in document, document
+ signed_url = urllib.parse.unquote(document['url'])
+ guid = document['guid']
+ hashes = {'md5': content_attachment_md5}
+ metadata = {
+ **{
+ 'datanode_type': record_name,
+ 'datanode_submitter_id': submitter_id,
+ 'datanode_object_id': guid
+ },
+ **hashes}
+
+ # open the file and upload it to the signed url returned above
+ with open(object_name, 'rb') as data:
+ # When you use this header, Amazon S3 checks the object against the provided MD5 value and,
+ # if they do not match, returns an error.
+ content_md5 = base64.b64encode(bytes.fromhex(content_attachment_md5))
+ headers = {'Content-MD5': content_md5}
+ # Our meta data
+ for key, value in metadata.items():
+ headers[f"x-amz-meta-{key}"] = value
+ r = requests.put(signed_url, data=data, headers=headers)
+ assert r.status_code == 200, (signed_url, r.text)
+ get_logger_("upload").info(f"Successfully uploaded file \"{object_name}\" to GUID {guid}")
+
+ # update the indexd record with urls, authz, size and hashes
+ indexd_record = index_client.get_record(guid)
+ assert 'rev' in indexd_record, record
+ rev = indexd_record['rev']
+ r = index_client.update_blank(guid, rev, hashes=hashes, size=content_attachment_size)
+
+ urls = [f"s3://{bucket_name}/{guid}/{object_name}"]
+ authz = [f'/programs/{program}/projects/{project}']
+
+ result = index_client.update_record(guid=guid, version=rev, file_name=object_name,
+ authz=authz, urls=urls,
+ metadata=metadata)
+ assert result # TODO - better check
+ json.dump()
+ print(json.dumps({'id': record_id, 'name': record_name, 'object_id': guid}))
+
+
+@cli.command()
+@click.option('--bucket_name', default='aced-default', show_default=True,
+ help='File name to upload')
+@click.option('--project_path', required=True, default=None, show_default=True,
+ help='Path to simulated metadata data see: shorturl.at/jqvZ5')
+@click.option('--program', default='MyFirstProgram', show_default=True,
+ help='Gen3 program')
+@click.option('--project', default='MyFirstProject', show_default=True,
+ help='Gen3 project')
+@click.pass_context
+def upload(ctx, bucket_name, project_path, program, project):
+ """Filter data_files found in generated synthetic data to gen3 managed bucket, update hashes and size."""
+
+ index_client = ctx.obj['index_client']
+ file_client = ctx.obj['file_client']
+
+ # this code reads files created by gen3's test meta data
+ # https://github.com/uc-cdis/compose-services/blob/master/docs/using_the_commons.md#generating-test-metadata
+ project_path = Path(project_path)
+ for synthetic_data_path in list(project_path.glob('**/*.json')):
+ updated_records = []
+ for record in json.load(open(synthetic_data_path, "r")):
+ if 'file_name' not in record:
+ break
+
+ object_name = record['file_name'].lstrip('/')
+
+ # create a record in gen3, get a signed url
+ document = file_client.upload_file(object_name, bucket=bucket_name)
+ assert 'guid' in document, document
+ assert 'url' in document, document
+ signed_url = urllib.parse.unquote(document['url'])
+ guid = document['guid']
+ hashes = {'md5': record['md5sum']}
+ metadata = {
+ **{
+ 'datanode_type': record['type'],
+ 'datanode_submitter_id': record['submitter_id'],
+ 'datanode_object_id': guid
+ },
+ **hashes}
+
+ # open the file and upload it to the signed url returned above
+ with open(record['file_name'], 'rb') as data:
+ # When you use this header, Amazon S3 checks the object against the provided MD5 value and,
+ # if they do not match, returns an error.
+ content_md5 = base64.b64encode(bytes.fromhex(record['md5sum']))
+ headers = {'Content-MD5': content_md5}
+ # Our meta data
+ for key, value in metadata.items():
+ headers[f"x-amz-meta-{key}"] = value
+ r = requests.put(signed_url, data=data, headers=headers)
+ assert r.status_code == 200, (signed_url, r.text)
+ get_logger_("upload").info(f"Successfully uploaded file \"{record['file_name']}\" to GUID {guid}")
+
+ # update the indexd record with urls, authz, size and hashes
+ indexd_record = index_client.get_record(guid)
+ assert 'rev' in indexd_record, record
+ rev = indexd_record['rev']
+ r = index_client.update_blank(guid, rev, hashes=hashes, size=record["file_size"])
+
+ urls = [f"s3://{bucket_name}/{guid}/{object_name}"]
+ authz = [f'/programs/{program}/projects/{project}']
+
+ result = index_client.update_record(guid=guid, version=rev, file_name=record['file_name'],
+ authz=authz, urls=urls,
+ metadata=metadata)
+ assert result # TODO - better check
+
+ record['object_id'] = guid
+ updated_records.append(record)
+
+ if len(updated_records) > 0:
+ with open(synthetic_data_path, "w") as fp:
+ json.dump(updated_records, fp, indent=4)
+ print(f"Uploaded data and update object_id in records in {synthetic_data_path}")
+
+
+def pfb_reader(pfb_path):
+ """Yields records from pfb."""
+ reader = fastavro.read.reader
+ with open(pfb_path, 'rb') as fo:
+ for record in reader(fo):
+ yield record
+
+
+@cli.command()
+@click.option('--did', default=None, show_default=True,
+ help='GUID from indexd')
+@click.option('--file_name', default=None, show_default=True,
+ help='output path')
+@click.pass_context
+def drs_download(ctx, did, file_name):
+ """
+ https://github.com/ga4gh/fasp-clients/blob/55dad8373637765bae43a1c670afc5f2b7b302b8/src/fasp/loc/gen3drsclient.py#L60
+ """
+ assert did, "Missing did (guid) parameter"
+ result = ctx.obj['index_client'].get_record(did)
+
+ assert 'hashes' in result, f'Expected "hashes" {result}'
+ assert 'md5' in result['hashes'], f'Expected "hashes.md5" {result}'
+ assert 'size' in result, f'Expected "size" {result}'
+
+ md5 = result['hashes']['md5']
+ size = result['size']
+ result = ctx.obj['file_client'].get_presigned_url(did)
+ assert 'url' in result, f'Expected "url" {result}'
+ presigned_url = result['url']
+ from download import download, DownloadURL
+ download_url = DownloadURL(url=presigned_url, md5=md5, size=size)
+ download(urls=[download_url])
+
+#
+# @cli.command()
+# @click.option('--did', default=None, show_default=True,
+# help='GUID from indexd')
+# @click.option('--file_name', default=None, show_default=True,
+# help='output path')
+# @click.pass_context
+# def download(ctx, did, file_name):
+# """Download file_name."""
+# assert did, "Missing did (guid) parameter"
+# result = ctx.obj['file_client'].get_presigned_url(did)
+# assert 'url' in result, result
+# presigned_url = result['url']
+# if not file_name:
+# result = ctx.obj['index_client'].get_record(did)
+# file_name = result['file_name']
+#
+# # NOTE the stream=True parameter below
+# # see https://stackoverflow.com/questions/16694907/download-large-file-in-python-with-requests
+# with requests.get(presigned_url, allow_redirects=True, stream=True) as r:
+# r.raise_for_status()
+# with open(file_name, 'wb') as f:
+# for chunk in r.iter_content(chunk_size=8192):
+# # If you have chunk encoded response uncomment if
+# # and set chunk_size parameter to None.
+# #if chunk:
+# f.write(chunk)
+
+
+if __name__ == '__main__':
+ cli()
diff --git a/etl/generate_simulated_files b/etl/generate_simulated_files
new file mode 100755
index 00000000..e46ae86f
--- /dev/null
+++ b/etl/generate_simulated_files
@@ -0,0 +1,81 @@
+#!/usr/bin/env python3
+import json
+import os
+import logging
+from pathlib import Path
+from random import randint
+import pathlib
+import click
+
+
+from faker import Faker
+from faker_biology.bioseq import Bioseq
+from cdislogging import get_logger as get_gen3_logger
+
+
+fake = Faker()
+fake.add_provider(Bioseq)
+
+
+def get_logger_(name):
+ """Return logger with level set to info"""
+ logger = logging.getLogger(name)
+ # logger.setLevel(logging.INFO)
+ return logger
+
+
+def file_attributes(file_name):
+ """Calculate the hash and size."""
+ import hashlib
+
+ md5_hash = hashlib.md5()
+
+ with open(file_name, "rb") as f:
+ # Read and update hash in chunks of 4K
+ for byte_block in iter(lambda: f.read(4096), b""):
+ md5_hash.update(byte_block)
+
+ return md5_hash.hexdigest(), os.lstat(file_name).st_size
+
+
+@click.command()
+@click.option('--project_path', required=True, default=None, show_default=True,
+ help='Path to synthetic data')
+@click.option('--output_path', required=True, default=None, show_default=True,
+ help='Path to synthetic data')
+@click.pass_context
+def cli(ctx, project_path, output_path):
+ """Read synthetic meta data, generate synthetic data file for indexd,
+ updates datanode with file_name, _size and md5."""
+ ctx.ensure_object(dict)
+ project_path = Path(project_path)
+ pathlib.Path(output_path).mkdir(parents=True, exist_ok=True)
+ for synthetic_data_path in list(project_path.glob('**/*.json')):
+ updated_records = []
+ for record in json.load(open(synthetic_data_path, "r")):
+ if 'file_name' not in record:
+ break
+ synthetic_file_path = f"{record['file_name'].replace('tests/fixtures/projects/MyFirstProject/DATA/', '')}"
+ if output_path not in synthetic_file_path:
+ synthetic_file_path = f"{output_path}/{synthetic_file_path}".replace('//', '/')
+ with open(synthetic_file_path, "w") as fp:
+ # write random number of lines of dna of random length
+ fp.writelines([fake.dna(randint(80, 256)) for x in range(randint(20, 100))])
+ md5, file_size = file_attributes(synthetic_file_path)
+ record["md5sum"] = md5
+ record["file_size"] = file_size
+ record["file_name"] = synthetic_file_path
+ updated_records.append(record)
+ if len(updated_records) > 0:
+ with open(synthetic_data_path, "w") as fp:
+ json.dump(updated_records, fp, indent=4)
+ print(f"Created synthetic files for records in {synthetic_data_path}")
+
+
+if __name__ == '__main__':
+ # set logging to warning, since gen3.submission logs a verbose INFO message on each call :-()
+ log_fmt = "%(asctime)s %(name)s %(levelname)s : %(message)s"
+ logging.basicConfig(level=logging.WARNING, format=log_fmt)
+ # set gen3's logger as well
+ get_gen3_logger('__name__', log_level='warn', format=log_fmt)
+ cli()
diff --git a/etl/import_pfb.py b/etl/import_pfb.py
new file mode 100644
index 00000000..e8e92058
--- /dev/null
+++ b/etl/import_pfb.py
@@ -0,0 +1,271 @@
+import json
+from datetime import datetime
+
+import psycopg2
+from psycopg2.sql import Identifier, SQL
+
+from fastavro import reader
+from pfb.base import handle_schema_field_unicode, is_enum, decode_enum
+from pelican.dictionary import init_dictionary, DataDictionaryTraversal
+
+
+def create_node_dict(node_id, node_name, values, edges):
+ node_dict = {
+ "id": node_id,
+ "name": node_name,
+ "object": values,
+ "relations": edges[node_id] if node_id in edges else [],
+ }
+
+ return node_dict
+
+
+def split_by_n(input_list, n=10000):
+ return [input_list[x : x + n] for x in range(0, len(input_list), n)]
+
+
+def get_ids_from_table(db, table, ids, id_column):
+ data = None
+
+ if not table or not ids or not id_column:
+ # TODO we need to use gen3logging
+ print(
+ f"[WARNING] Got a false-y input to a query. table: {table}, ids: {ids}, id_column: {id_column}"
+ )
+ return data
+
+ for ids_chunk in split_by_n(ids):
+ try:
+ current_chunk_data = (
+ db.option(
+ "query",
+ "SELECT * FROM {} WHERE {} IN ('{}')".format(
+ table, id_column, "','".join(ids_chunk)
+ ),
+ )
+ .option("fetchsize", "10000")
+ .load()
+ )
+
+ if data:
+ data = data.union(current_chunk_data)
+ else:
+ data = current_chunk_data
+ except TypeError:
+ print(
+ f"[ERROR] Query got invalid inputs: table: {table}, ids: {ids}, id_column: {id_column}."
+ f"Split: {split_by_n(ids)}"
+ )
+ pass
+ else:
+ print(
+ f"[WARNING] Got a false-y ids_chunk by splitting ids: {ids}. Split: {split_by_n(ids)}"
+ )
+
+ return data if data and data.first() else None
+
+
+def convert_to_node(x, is_base64, project_id):
+ obj = x["object"]
+ to_update = {}
+ for name, value in obj.items():
+ if name in is_base64[x["name"]]:
+ if value and is_base64[x["name"]][name]:
+ to_update[name] = decode_enum(value)
+ # TODO get parameter from CLI
+ to_update['project_id'] = project_id
+ obj.update(to_update)
+
+ r = {
+ "created": datetime.now(),
+ "acl": json.dumps({}),
+ "_sysan": json.dumps({}),
+ "_props": json.dumps(obj),
+ "node_id": x["id"],
+ }
+
+ return r
+
+
+def convert_to_edge(x, edge_tables):
+ return [
+ (
+ edge_tables[(x["name"], i["dst_name"])],
+ {
+ "created": datetime.now(),
+ "acl": json.dumps({}),
+ "_sysan": json.dumps({}),
+ "_props": json.dumps({}),
+ "src_id": x["id"],
+ "dst_id": i["dst_id"],
+ },
+ )
+ for i in x["relations"]
+ ]
+
+
+def write_table(cur, table, node, dry_run=True):
+ """Write node to table."""
+ # TODO - improve speed https://naysan.ca/2020/05/09/pandas-to-postgresql-using-psycopg2-bulk-insert-performance-benchmark/
+ columns = ','.join(node.keys())
+ place_holders = ','.join(['%s'] * len(node.keys()))
+ values = [node[column] for column in node.keys()]
+ if 'node_id' in node.keys():
+ # vertex
+ insert = f'insert into "{table}" ({columns}) values ({place_holders}) on conflict(node_id) do nothing;'
+ else:
+ # edge
+ insert = f'insert into "{table}" ({columns}) values ({place_holders}) on conflict(src_id, dst_id) do nothing;'
+ if not dry_run:
+ cur.execute(SQL(insert), values)
+
+
+def import_pfb_job(pfb_file, project_id, project_node_id, ddt, conn, dry_run, document_reference_object_ids):
+ """Import the PFB into the database."""
+ start_time = datetime.now()
+ print(start_time)
+
+ with open(pfb_file, "rb") as schema_field:
+ avro_reader = reader(schema_field)
+ writer_schema = avro_reader.writer_schema
+
+ schema = []
+ for schema_field in writer_schema["fields"]:
+ if schema_field["name"] == "object":
+ it = iter(schema_field["type"])
+ # skip metadata
+ next(it)
+ for node in it:
+ schema.append(node)
+ for field in node["fields"]:
+ handle_schema_field_unicode(field, encode=False)
+
+ _is_base64 = {}
+
+ for node in schema:
+ _is_base64[node["name"]] = fields = {}
+ for field in node["fields"]:
+ fields[field["name"]] = is_enum(field["type"])
+
+ edge_table_by_labels = ddt.get_edge_table_by_labels()
+
+ insert_count = 0
+ total_count = 0
+ batch_size = 10000
+ cur = conn.cursor()
+ for record in avro_reader:
+ if record['name'] == 'Metadata':
+ continue
+ if record['name'] == 'ResearchStudy':
+ # link the ResearchStudy to the gen3 project
+ record['relations'] = [{"dst_id": project_node_id, "dst_name": "project"}]
+ if record['name'] == 'DocumentReference':
+ # link the ResearchStudy to the gen3 project
+ if record['id'] in document_reference_object_ids:
+ record['object']['object_id'] = document_reference_object_ids[record['id']]
+ # print(f"Set document_reference_object_ids {record['object']['object_id']}")
+
+ write_table(
+ cur=cur,
+ table=ddt.get_node_table_by_label()[record['name']],
+ node=convert_to_node(record, _is_base64, project_id),
+ dry_run=dry_run
+ )
+ edge_tuples = convert_to_edge(record, edge_table_by_labels)
+ if edge_tuples:
+ for edge_tuple in edge_tuples:
+ edge_table = edge_tuple[0]
+ edge = edge_tuple[1]
+ write_table(
+ cur=cur,
+ table=edge_table,
+ node=edge,
+ dry_run=dry_run
+ )
+ insert_count += 1
+ if insert_count == batch_size:
+ conn.commit()
+ total_count += insert_count
+ insert_count = 0
+ print("total_count {} {} {}".format(total_count, record['name'], datetime.now()))
+ conn.commit()
+ total_count += insert_count
+ time_elapsed = datetime.now() - start_time
+ print("Elapsed time: {} total_count {}".format(time_elapsed, total_count))
+ cur.close()
+ conn.close()
+ return
+
+
+if __name__ == "__main__":
+
+ # TODO - get this from the command line
+ dry_run = False
+ # TODO - get this from the command line
+ dictionary_url = 'https://aced-public.s3.us-west-2.amazonaws.com/aced.json' # os.environ["DICTIONARY_URL"]
+
+ # TODO - get this from the command line
+ with open("Secrets/sheepdog_creds.json") as pelican_creds_file:
+ sheepdog_creds = json.load(pelican_creds_file)
+
+ # DB_URL = "jdbc:postgresql://{}/{}".format(
+ # sheepdog_creds["db_host"], sheepdog_creds["db_database"]
+ # )
+ DB_USER = sheepdog_creds["db_username"]
+ DB_PASS = sheepdog_creds["db_password"]
+
+ # TODO - get this from the command line
+ DB_NAME = 'metadata_db'
+
+ # TODO - get this from the command line
+ program_name = 'MyFirstProgram'
+ project_code = 'MyFirstProject'
+
+ # NEW_DB_NAME = input_data_json["db"]
+
+ conn = psycopg2.connect(
+ database=DB_NAME,
+ user=DB_USER,
+ password=DB_PASS,
+ # TODO - get this from the command line
+ host='localhost',
+ # port=DATABASE_CONFIG.get('port'),
+ )
+
+ cur = conn.cursor()
+ cur.execute("select node_id, _props from \"node_program\";")
+ programs = cur.fetchall()
+ programs = [{'node_id': p[0], '_props': p[1]} for p in programs]
+ program = next(iter([p for p in programs if p['_props']['name'] == program_name]), None)
+ assert program, f"{program_name} not found in node_program"
+ cur.execute("select node_id, _props from \"node_project\";")
+ projects = cur.fetchall()
+ projects = [{'node_id': p[0], '_props': p[1]} for p in projects]
+ project_node_id = next(iter([p['node_id'] for p in projects if p['_props']['code'] == project_code]), None)
+ assert project_node_id, f"{project_code} not found in node_project"
+ project_id = f"{program_name}-{project_code}"
+
+ # TODO - get this from the command line
+ pfb_file = 'output/research_study_Alzheimers.pfb'
+ print(f"Importing {pfb_file} into {project_id} project node {project_node_id}")
+
+ dictionary, model = init_dictionary(url=dictionary_url)
+ ddt = DataDictionaryTraversal(model)
+
+ # TODO - get this from the command line
+ # get the object ids created by file upload-pfb
+ document_reference_object_ids = {}
+ with open('file-object_ids.ndjson') as f:
+ for line in f.readlines():
+ document_reference_object_id = json.loads(line)
+ document_reference_object_ids[document_reference_object_id['id']] = document_reference_object_id['object_id']
+
+ import_pfb_job(
+ pfb_file=pfb_file,
+ project_id=project_id,
+ project_node_id=project_node_id,
+ ddt=ddt,
+ conn=conn,
+ dry_run=dry_run,
+ document_reference_object_ids=document_reference_object_ids
+ )
diff --git a/etl/metadata b/etl/metadata
new file mode 100755
index 00000000..611f7415
--- /dev/null
+++ b/etl/metadata
@@ -0,0 +1,541 @@
+#!/usr/bin/env python3
+import csv
+import gzip
+import io
+import json
+import os
+import itertools
+import logging
+from collections import defaultdict
+from glob import glob
+
+import fastavro
+from gen3.auth import Gen3Auth
+from gen3.metadata import Gen3Metadata
+from gen3.submission import Gen3Submission
+import click
+import jwt
+import yaml
+
+from cdislogging import get_logger as get_gen3_logger
+from dictionary import init_dictionary, DataDictionaryTraversal, convert_to_node
+
+log_fmt = "%(asctime)s %(name)s %(levelname)s : %(message)s"
+
+# set logging to warning, since gen3.submission logs a verbose INFO message on each call :-()
+logging.basicConfig(level=logging.WARNING, format=log_fmt)
+# set gen3's logger as well
+# get_gen3_logger('__name__', log_level='warn', format=log_fmt)
+
+
+def get_logger_(name):
+ """Return logger with level set to info"""
+ logger = logging.getLogger(name)
+ logger.setLevel(logging.INFO)
+ return logger
+
+
+def delete_all(submission_client, program, project, batch_size=200, types=[]):
+ """Delete all nodes in types hierarchy, skips program and project."""
+ for t in types:
+ try:
+ if t in ['program', 'project']:
+ continue
+ delete_type(submission_client, program, project, batch_size, t)
+ except Exception as e:
+ print(e)
+ raise e
+
+
+def delete_type(submission_client, program, project, batch_size, t):
+ """Delete all instances of a type."""
+ logger = get_logger_("delete_type")
+ response = submission_client.export_node(program, project, node_type=t, fileformat='json')
+ # # pool = mp.Pool(mp.cpu_count())
+
+ def collect_result(delete_response):
+ delete_response = delete_response.json()
+ assert delete_response['code'] == 200, delete_response
+ # logger.info('deleted {} {}'.format(t, delete_response['message']))
+
+ if 'data' not in response or len(response['data']) == 0:
+ logger.warning(f'No {t} to delete {response}')
+ else:
+ for ids in grouper(batch_size, [n['id'] for n in response['data']]):
+ logger.info(f'deleting {program}-{project}.{t} {len(ids)} items.')
+ ids = ','.join(ids)
+ collect_result(submission_client.delete_record(program, project, ids))
+ # # pool.apply_async(submission_client.delete_record, args=(program, project, ids), callback=collect_result)
+ # Close Pool and let all the processes complete
+ # postpones the execution of next line of code until all processes in the queue are done
+ # # pool.close()
+ # # pool.join()
+
+
+class JsonReader:
+ """Read json and return dict iterator."""
+
+ def __init__(self, path):
+ """Open file."""
+ if path.endswith(".json.gz"):
+ self.fp = io.TextIOWrapper(io.BufferedReader(gzip.GzipFile(path)))
+ else:
+ self.fp = open(path, "r", encoding='utf-8')
+ self.items = None
+
+ def __iter__(self):
+ """Return self."""
+ return self
+
+ def __next__(self):
+ """Iterate to next row."""
+ try:
+ if self.items and len(self.items) > 0:
+ return self.items.pop(0)
+ if self.items:
+ raise IndexError()
+ line = self.fp.readline()
+ if len(line) < 1:
+ raise IndexError()
+ return json.loads(line)
+ except json.decoder.JSONDecodeError:
+ self.fp.seek(0, 0)
+ obj_ = json.load(self.fp)
+ self.items = obj_
+ if not isinstance(self.items, list):
+ self.items = [self.items]
+ return self.items.pop(0)
+ except IndexError:
+ raise StopIteration()
+
+
+def reader(path, **kwargs):
+ """Wrap gzip if necessary."""
+ if path.endswith(".json.gz"):
+ return JsonReader(path)
+ elif path.endswith(".gz"):
+ return io.TextIOWrapper(
+ io.BufferedReader(gzip.GzipFile(path))
+ )
+ elif path.endswith(".csv"):
+ return csv.DictReader(open(path, "r", encoding='utf-8'), **kwargs)
+ elif path.endswith(".tsv"):
+ return csv.DictReader(open(path, "r", encoding='utf-8'), delimiter="\t", **kwargs)
+ elif path.endswith(".json"):
+ return JsonReader(path)
+ elif path.endswith(".ndjson"):
+ return JsonReader(path)
+ else:
+ return open(path, "r", encoding='utf-8')
+
+
+def grouper(n, iterable):
+ """Chunk iterable into n size chunks."""
+ it = iter(iterable)
+ while True:
+ chunk = tuple(itertools.islice(it, n))
+ if not chunk:
+ return
+ yield chunk
+
+
+def create_node(submission_client, program_name, project_code, node):
+ """Create node(s)."""
+ logger = get_logger_("create_node")
+
+ try:
+ nodes = node
+ if not isinstance(node, (list,)):
+ nodes = [node]
+ response = None
+ response = submission_client.submit_record(program_name, project_code, nodes)
+ return response
+ except Exception as e:
+ logger.error(f"create_node: error {e}")
+ logger.error(f"create_node: error {nodes}")
+ if response:
+ for entity in response.get('entities', []):
+ for error in entity.get('errors', []):
+ logger.error('{} {} {}'.format(error['type'], entity['type'], entity))
+ for error in response.get('transactional_errors', []):
+ logger.error(' transactional_error {}'.format(error))
+ logger.error(json.dumps(response))
+ raise e
+
+
+@click.group()
+@click.option('--gen3_credentials_file', default='Secrets/credentials.json', show_default=True,
+ help='API credentials file downloaded from gen3 profile.')
+@click.pass_context
+def cli(ctx, gen3_credentials_file):
+ """Metadata loader."""
+
+ endpoint = extract_endpoint(gen3_credentials_file)
+ get_logger_("cli").debug(f"Read {gen3_credentials_file} endpoint {endpoint}")
+ auth = Gen3Auth(endpoint, refresh_file=gen3_credentials_file)
+ submission_client = Gen3Submission(endpoint, auth)
+ ctx.ensure_object(dict)
+ ctx.obj['submission_client'] = submission_client
+ ctx.obj['discovery_client'] = Gen3Metadata(endpoint, auth)
+ ctx.obj['endpoint'] = endpoint
+ ctx.obj['programs'] = [link.split('/')[-1] for link in submission_client.get_programs()['links']]
+
+
+@cli.command()
+@click.option('--pfb_path', default=None, required=True, show_default=True,
+ help='directory that contains research_studyXXXX.pfb')
+@click.option('--program', default=None, show_default=True,
+ help='Gen3 "program"')
+@click.option('--project', default=None, show_default=True,
+ help='Gen3 "project"')
+@click.option('--batch_size', default=10, show_default=True,
+ help='number of records to process per call')
+@click.pass_context
+def load_pfb(ctx, pfb_path, program, project, batch_size):
+ """Loads metadata into project"""
+ logger = get_logger_("load_pfb")
+ submission_client = ctx.obj['submission_client']
+ dictionary, model = init_dictionary(url='https://aced-public.s3.us-west-2.amazonaws.com/aced.json')
+ ddt = DataDictionaryTraversal(model)
+ node_table_by_label = ddt.get_node_table_by_label()
+ # {'root': 'node_root', 'data_release': 'node_datarelease', 'DocumentReference': 'node_DocumentReference',
+ # 'Observation': 'node_Observation', 'Patient': 'node_Patient', 'Practitioner': 'node_Practitioner',
+ # 'MedicationRequest': 'node_MedicationRequest', 'program': 'node_program', 'project': 'node_project',
+ # 'DiagnosticReport': 'node_DiagnosticReport', 'Task': 'node_Task', 'Condition': 'node_Condition',
+ # 'Location': 'node_Location', 'Encounter': 'node_Encounter', 'Procedure': 'node_Procedure',
+ # 'Organization': 'node_Organization', 'ImagingStudy': 'node_ImagingStudy', 'ResearchStudy': 'node_ResearchStudy',
+ # 'Specimen': 'node_Specimen', 'Medication': 'node_Medication', 'ResearchSubject': 'node_ResearchSubject',
+ # 'core_metadata_collection': 'node_coremetadatacollection'}
+ edge_table_by_labels = ddt.get_edge_table_by_labels()
+ # {('data_release', 'root'): 'edge_datareleasedescribesroot',
+ # ('DocumentReference', 'Patient'): 'edge_DocumentReferencePatientsPatient',
+ # ('DocumentReference', 'Organization'): 'edge_ec062b7b_DoOrOr',
+ # ('Observation', 'ResearchStudy'): 'edge_0b10b6fb_ObReRe',
+ # ('Observation', 'Specimen'): 'edge_ObservationSpecimenSpecimen',
+ # ('Observation', 'Patient'): 'edge_ObservationPatientsPatient',
+ # ('Observation', 'Encounter'): 'edge_ObservationEncountersEncounter',
+ # ('Patient', 'Organization'): 'edge_PatientOrganizationsOrganization',
+ # ('MedicationRequest', 'Patient'): 'edge_MedicationRequestPatientsPatient',
+ # ('MedicationRequest', 'Encounter'): 'edge_8ffbdcaa_MeEnEn',
+ # ('MedicationRequest', 'Medication'): 'edge_df7c93a6_MeMeMe', ('project', 'program'): 'edge_projectmemberofprogram',
+ # ('DiagnosticReport', 'Patient'): 'edge_DiagnosticReportPatientsPatient',
+ # ('DiagnosticReport', 'Encounter'): 'edge_DiagnosticReportEncountersEncounter',
+ # ('DiagnosticReport', 'Practitioner'): 'edge_398dc3a8_DiPrPr', ('Task', 'Patient'): 'edge_TaskPatientsPatient',
+ # ('Task', 'Specimen'): 'edge_TaskSpecimenSpecimen', ('Task', 'DocumentReference'): 'edge_42e2e26b_TaDoDo',
+ # ('Condition', 'Patient'): 'edge_ConditionPatientsPatient',
+ # ('Condition', 'Encounter'): 'edge_ConditionEncountersEncounter',
+ # ('Location', 'Organization'): 'edge_LocationOrganizationsOrganization',
+ # ('Encounter', 'Patient'): 'edge_EncounterPatientsPatient',
+ # ('Procedure', 'Patient'): 'edge_ProcedurePatientsPatient',
+ # ('Procedure', 'Encounter'): 'edge_ProcedureEncountersEncounter',
+ # ('Organization', 'Organization'): 'edge_85f2b993_OrOrOr',
+ # ('ImagingStudy', 'Patient'): 'edge_ImagingStudyPatientsPatient',
+ # ('ImagingStudy', 'Encounter'): 'edge_ImagingStudyEncountersEncounter',
+ # ('ResearchStudy', 'ResearchStudy'): 'edge_e2d4f866_ReReRe',
+ # ('ResearchStudy', 'Practitioner'): 'edge_876d23c4_RePrPr',
+ # ('ResearchStudy', 'Organization'): 'edge_8abff393_ReOrOr',
+ # ('ResearchStudy', 'project'): 'edge_ResearchStudyprojectsproject',
+ # ('Specimen', 'Patient'): 'edge_SpecimenPatientsPatient',
+ # ('ResearchSubject', 'Patient'): 'edge_ResearchSubjectPatientsPatient',
+ # ('ResearchSubject', 'ResearchStudy'): 'edge_7ecdea32_ReReRe',
+ # ('core_metadata_collection', 'project'): 'edge_9197510c_comecodafrpr'}
+
+ for records in grouper(batch_size, pfb_reader(pfb_path)):
+ nodes = []
+ for record in records:
+ if record['name'] == 'Metadata':
+ continue
+ print(record['name'], record['id'], record)
+ convert_to_node(record)
+ break
+ break
+ # collect_result(create_node(submission_client, program, project, nodes))
+
+
+def pfb_reader(pfb_path):
+ """Yields records from pfb."""
+ reader_ = fastavro.read.reader
+ with open(pfb_path, 'rb') as fo:
+ for record in reader_(fo):
+ yield record
+
+
+def upload_metadata(path, program, project, submission_client, batch_size):
+ """Read gen3 json and write to gen3."""
+
+ logger = get_logger_("upload_metadata")
+
+ def collect_result(response_):
+ is_error = False
+ for entity in response_['entities']:
+ for error in entity.get('errors', []):
+ logger.error('{} {} {}'.format(error['type'], entity['type'], entity))
+ is_error = True
+ for error in response_['transactional_errors']:
+ logger.error('transactional_error {}'.format(error))
+ logger.error(json.dumps(response_))
+ is_error = True
+ if is_error:
+ logger.debug(response_)
+
+ for p in glob(path):
+ logger.info(f"Uploading {p}")
+ for lines in grouper(batch_size, reader(p)):
+ nodes = [line for line in lines]
+
+ if nodes[0]['type'] == 'project':
+ for node in nodes:
+ logger.debug('creating program')
+ response = submission_client.create_program(
+ {'name': program, 'dbgap_accession_number': program, 'type': 'program'})
+ assert response, 'could not parse response {}'.format(response)
+ # assert 'code' in response, f'Unexpected response {response}'
+ # assert response['code'] == 200, 'could not create {} program'.format(response)
+ assert 'id' in response, 'could not create {} program'.format(response)
+ assert program in response['name'], 'could not create {} program'.format(response)
+
+ response = submission_client.create_project(program, node)
+ assert response, 'could not parse response'
+ assert 'code' in response, f'Unexpected response {response}'
+ assert response['code'] == 200, 'could not create {} {}'.format(nodes[0]['type'], response)
+ assert 'successful' in response['message'], 'could not create {} {}'.format(nodes[0]['type'],
+ response)
+ logger.info('Created project {}'.format(node['code']))
+ continue
+
+ # if nodes[0]['type'] == 'experiment':
+ # project = nodes[0]['projects'][0]['code']
+
+ collect_result(create_node(submission_client, program, project, nodes))
+
+
+def get_schema(submission_client):
+ """Returns gen3 schema."""
+ schema = submission_client.get_dictionary_all()
+ return schema
+
+
+def nodes_in_load_order(submission_client):
+ """Introspects schema and returns types in order of db load."""
+ schema = get_schema(submission_client)
+ loaded = {}
+
+ def process(current_, depth):
+ loaded[current_['id']] = depth
+
+ def traverse(current_, depth=0, depth_limit=1):
+ if depth > depth_limit:
+ return
+ process(current_, depth)
+ target_type = current_['id']
+ for k in schema.keys():
+ n = schema[k]
+ if 'links' not in n or len(n['links']) == 0:
+ continue
+ links = n['links']
+ if 'subgroup' in links[0]:
+ links = links[0]['subgroup']
+ for link in links:
+ if 'target_type' in link and link['target_type'] == target_type:
+ process(schema[n['id']], depth)
+ for link in links:
+ if 'target_type' in link and link['target_type'] == target_type:
+ traverse(schema[n['id']], depth + 1, depth_limit)
+ depth_limit += 1
+
+ schema_keys = [k for k in schema.keys() if
+ not k.startswith('_') and not schema[k].get('category', None) == 'internal']
+ schema_keys = [schema[schema_key] for schema_key in schema_keys if schema[schema_key].get('links', None) == []][0]
+ traverse(schema_keys)
+
+ load_order_ = []
+ levels = set([v for k, v in loaded.items()])
+ for i in sorted(levels):
+ for k, v in loaded.items():
+ if v == i:
+ load_order_.append(k)
+ return load_order_
+
+
+def extract_endpoint(gen3_credentials_file):
+ """Get base url of jwt issuer claim."""
+ with open(gen3_credentials_file) as input_stream:
+ api_key = json.load(input_stream)['api_key']
+ claims = jwt.decode(api_key, options={"verify_signature": False})
+ assert 'iss' in claims
+ return claims['iss'].replace('/user', '')
+
+
+@cli.command()
+@click.pass_context
+def ls(ctx):
+ """Introspects schema and returns types in order."""
+ submission_client = ctx.obj['submission_client']
+ contents = {'programs': defaultdict(list)}
+
+ programs = submission_client.get_programs()
+
+ for program in ctx.obj['programs']:
+ contents['programs'][program] = [link.split('/')[-1] for link in submission_client.get_projects(program)['links']]
+ contents['entities'] = nodes_in_load_order(submission_client)
+ print(json.dumps(contents))
+
+
+@cli.command()
+@click.option('--program', show_default=True,
+ help='Gen3 "program"')
+@click.option('--project', show_default=True,
+ help='Gen3 "project"')
+@click.option('--batch_size', default=10, show_default=True,
+ help='number of records to process per call')
+@click.pass_context
+def empty(ctx, batch_size, program, project):
+ """Empties project, deletes all metadata."""
+ submission_client = ctx.obj['submission_client']
+ nodes = nodes_in_load_order(submission_client)
+ delete_all(submission_client, program, project, types=reversed(nodes), batch_size=batch_size)
+
+
+@cli.command()
+@click.option('--program', show_default=True,
+ help='Gen3 "program"')
+@click.option('--project', show_default=True,
+ help='Gen3 "project"')
+@click.pass_context
+def drop_project(ctx, program, project):
+ """Drops empty project"""
+ submission_client = ctx.obj['submission_client']
+ submission_client.delete_project(program, project)
+ get_logger_("drop+_project").info(f"Dropped {project}")
+
+
+@cli.command()
+@click.option('--program', show_default=True,
+ help='Gen3 "program"')
+# @click.option('--project', show_default=True,
+# help='Gen3 "project"')
+# @click.option('--data_directory', default=None, required=True, show_default=True,
+# help='Path to project.json')
+@click.pass_context
+def discovery(ctx, program):
+ """Writes project information to discovery metadata-service"""
+ discovery_client = ctx.obj['discovery_client']
+ discovery_descriptions = """
+Alcoholism~9300~Patients from 'Coherent Data Set' https://www.mdpi.com/2079-9292/11/8/1199/htm that were diagnosed with condition(s) of: Alcoholism. Data hosted by: aced-ohsu~aced-ohsu
+Alzheimers~45306~Patients from 'Coherent Data Set' https://www.mdpi.com/2079-9292/11/8/1199/htm that were diagnosed with condition(s) of: Alzheimer's, Familial Alzheimer's. Data hosted by: aced-ucl~aced-ucl
+Breast_Cancer~7105~Patients from 'Coherent Data Set' https://www.mdpi.com/2079-9292/11/8/1199/htm that were diagnosed with condition(s) of: Malignant neoplasm of breast (disorder). Data hosted by: aced-manchester~aced-manchester
+Colon_Cancer~25355~Patients from 'Coherent Data Set' https://www.mdpi.com/2079-9292/11/8/1199/htm that were diagnosed with condition(s) of: Malignant tumor of colon, Polyp of colon. Data hosted by: aced-stanford~aced-stanford
+Diabetes~65051~Patients from 'Coherent Data Set' https://www.mdpi.com/2079-9292/11/8/1199/htm that were diagnosed with condition(s) of: Diabetes. Data hosted by: aced-ucl~aced-ucl
+Lung_Cancer~25355~Patients from 'Coherent Data Set' https://www.mdpi.com/2079-9292/11/8/1199/htm that were diagnosed with condition(s) of: Non-small cell carcinoma of lung,TNM stage 1, Non-small cell lung cancer, Suspected lung cancer. Data hosted by: aced-manchester~aced-manchester
+Prostate_Cancer~35488~Patients from 'Coherent Data Set' https://www.mdpi.com/2079-9292/11/8/1199/htm that were diagnosed with condition(s) of: Metastasis from malignant tumor of prostate, Neoplasm of prostate, arcinoma in situ of prostate. Data hosted by: aced-stanford~aced-stanford""".split('\n')
+
+ for line in discovery_descriptions:
+ if len(line) == 0:
+ continue
+ print(line)
+ (name, _subjects_count, description, location, ) = line.split('~')
+ gen3_discovery = {}
+ # {
+ # "availability_mechanism": "061e83cedc",
+ # "availability_type": "Restricted",
+ # "code": "MyFirstProject",
+ # "date_collected": "b556077e00",
+ # "dbgap_accession_number": "phs-jenkins",
+ # "investigator_affiliation": "2395ceacf1",
+ # "investigator_name": "fd0174c590",
+ # "name": "bf0c3e26f3",
+ # "support_id": "155461fbb9",
+ # "support_source": "0343f459a1",
+ # "type": "project"
+ # }
+ gen3_discovery['tags'] = [
+ {"name": program, "category": "Program"},
+ {"name": f"aced_{name}", "category": "Study Registration"},
+ {"name": location, "category": "Study Location"},
+
+ ]
+ gen3_discovery['name'] = name
+ gen3_discovery['full_name'] = name
+ gen3_discovery['study_description'] = description
+
+ guid = f"aced_{name}"
+
+ gen3_discovery['commons'] = "ACED"
+ gen3_discovery['commons_name'] = "ACED Commons"
+ gen3_discovery['commons_url'] = 'staging.aced-idp.org'
+ gen3_discovery['__manifest'] = 0
+ gen3_discovery['_research_subject_count'] = int(_subjects_count)
+ gen3_discovery['_unique_id'] = guid
+ gen3_discovery['study_id'] = guid
+ discoverable_data = dict(_guid_type="discovery_metadata", gen3_discovery=gen3_discovery)
+ discovery_client.create(guid, discoverable_data, aliases=None, overwrite=True)
+ get_logger_("discovery").info(f"Added {name}")
+
+
+@cli.command()
+@click.option('--program', show_default=True,
+ help='Gen3 "program"')
+def drop_program(ctx, data_directory, program):
+ """Drops empty program"""
+ submission_client = ctx.obj['submission_client']
+ submission_client.delete_progranm(program)
+
+
+@cli.command()
+@click.option('--program', show_default=True, required=True,
+ help='Gen3 "program"')
+@click.option('--manifest', show_default=True, required=True,
+ help='Study names, conditions, expected counts, etc. e.g. coherent_studies.manifest.yaml')
+@click.pass_context
+def create_program(ctx, program, manifest):
+ """Creates program and projects."""
+ submission_client = ctx.obj['submission_client']
+ response = submission_client.create_program({'name': program, 'dbgap_accession_number': program, 'type': 'program'})
+ get_logger_('create_program').info(response)
+ study_manifests = yaml.load(open(manifest), yaml.SafeLoader)
+ for name, values in study_manifests.items():
+ response = submission_client.create_project(program, {
+ "type": "project",
+ "code": name,
+ "dbgap_accession_number": name,
+ "name": name
+ })
+ get_logger_('create_program').info(response)
+
+
+@cli.command()
+@click.option('--data_directory', default=None, required=True, show_default=True,
+ help='directory that contains .json')
+@click.option('--program', default=None, show_default=True,
+ help='Gen3 "program"')
+@click.option('--project', default=None, show_default=True,
+ help='Gen3 "project"')
+@click.option('--batch_size', default=10, show_default=True,
+ help='number of records to process per call')
+@click.pass_context
+def load(ctx, data_directory, program, project, batch_size):
+ """Loads metadata into project"""
+
+ submission_client = ctx.obj['submission_client']
+ if not program:
+ assert len(ctx.obj['programs']) == 1, f"No program provided and multiple programs {ctx.obj['programs']}"
+ program = ctx.obj['programs'][0]
+ if not project:
+ projects = [link.split('/')[-1] for link in submission_client.get_projects(program)['links']]
+ if len(projects) == 1:
+ project = projects[0]
+ else:
+ get_logger_('load').warning(f"No program provided and multiple programs {ctx.obj['programs']} {projects}")
+
+ nodes = nodes_in_load_order(submission_client)
+
+ for entity in nodes:
+ filename = f"{data_directory}/{entity}.json"
+ if os.path.isfile(filename):
+ upload_metadata(submission_client=submission_client, path=filename, program=program, project=project,
+ batch_size=batch_size)
+
+
+if __name__ == '__main__':
+ cli()
+
diff --git a/etl/requirements.txt b/etl/requirements.txt
new file mode 100644
index 00000000..cec9a2e1
--- /dev/null
+++ b/etl/requirements.txt
@@ -0,0 +1,27 @@
+pyjwt
+Click
+jsonpath-ng
+certifi
+faker-biology
+elasticsearch==6.8.2
+more-itertools
+
+
+# "stock" gen3 - the version on https://pypi.org/project/gen3/ 4.14.0 does not have the bucket change
+# gen3
+
+# uchicago official repo - has the bucket change https://github.com/uc-cdis/gen3sdk-python/pull/160
+https://github.com/uc-cdis/gen3sdk-python/archive/master.zip
+
+# to test:
+# from gen3.file import Gen3File
+# import inspect
+# assert 'bucket' in [parameter_name for parameter_name in inspect.signature(Gen3File.upload_file).parameters]
+# print('ok')
+
+
+# our fork of pelican that supports 3.9
+git+https://github.com/ACED-IDP/pelican.git@chore/python-version
+# to test:
+# python3 -c "from pelican.dictionary import init_dictionary, DataDictionaryTraversal; print('ok')"
+
diff --git a/etl/setup-minio.sh b/etl/setup-minio.sh
new file mode 100644
index 00000000..eaa74ad3
--- /dev/null
+++ b/etl/setup-minio.sh
@@ -0,0 +1,44 @@
+export MINIO_TEST_USER=test
+export MINIO_TEST_PASSWORD=testtest
+export MINIO_ADMIN_USER=minioadmin
+export MINIO_ADMIN_PASSWORD=minio\$1234
+
+# setup aliases
+mc alias set default http://minio-default:9000 $MINIO_ADMIN_USER $MINIO_ADMIN_PASSWORD
+mc alias set ohsu http://minio-ohsu:9000 $MINIO_ADMIN_USER $MINIO_ADMIN_PASSWORD
+mc alias set ucl http://minio-ucl:9000 $MINIO_ADMIN_USER $MINIO_ADMIN_PASSWORD
+mc alias set manchester http://minio-manchester:9000 $MINIO_ADMIN_USER $MINIO_ADMIN_PASSWORD
+mc alias set stanford http://minio-stanford:9000 $MINIO_ADMIN_USER $MINIO_ADMIN_PASSWORD
+
+# make buckets
+mc mb default/aced-default
+mc mb default/aced-public
+mc mb ohsu/aced-ohsu
+mc mb ucl/aced-ucl
+mc mb manchester/aced-manchester
+mc mb stanford/aced-stanford
+
+
+# add users
+mc admin user add default $MINIO_TEST_USER $MINIO_TEST_PASSWORD
+mc admin user add ohsu $MINIO_TEST_USER $MINIO_TEST_PASSWORD
+mc admin user add ucl $MINIO_TEST_USER $MINIO_TEST_PASSWORD
+mc admin user add manchester $MINIO_TEST_USER $MINIO_TEST_PASSWORD
+mc admin user add stanford $MINIO_TEST_USER $MINIO_TEST_PASSWORD
+
+# add policies
+mc admin policy set default readwrite user=$MINIO_TEST_USER
+mc admin policy set ohsu readwrite user=$MINIO_TEST_USER
+mc admin policy set ucl readwrite user=$MINIO_TEST_USER
+mc admin policy set manchester readwrite user=$MINIO_TEST_USER
+mc admin policy set stanford readwrite user=$MINIO_TEST_USER
+# public bucket
+mc policy set public default/aced-public
+
+# when objects added to bucket, call webhook (see minio-webhook/ for example)
+mc event add default/aced-default arn:minio:sqs::PRIMARY:webhook
+mc event add ohsu/aced-ohsu arn:minio:sqs::PRIMARY:webhook
+mc event add ucl/aced-ucl arn:minio:sqs::PRIMARY:webhook
+mc event add manchester/aced-manchester arn:minio:sqs::PRIMARY:webhook
+mc event add stanford/aced-stanford arn:minio:sqs::PRIMARY:webhook
+
diff --git a/etl/truncate_imported_tables.sql b/etl/truncate_imported_tables.sql
new file mode 100644
index 00000000..f216a4b9
--- /dev/null
+++ b/etl/truncate_imported_tables.sql
@@ -0,0 +1,32 @@
+-- delete all imported data
+
+\c metadata_db ;
+TRUNCATE "node_condition" CASCADE ;
+TRUNCATE "node_datarelease" CASCADE ;
+TRUNCATE "node_diagnosticreport" CASCADE ;
+TRUNCATE "node_documentreference" CASCADE ;
+TRUNCATE "node_encounter" CASCADE ;
+TRUNCATE "node_imagingstudy" CASCADE ;
+TRUNCATE "node_location" CASCADE ;
+TRUNCATE "node_medication" CASCADE ;
+TRUNCATE "node_medicationrequest" CASCADE ;
+TRUNCATE "node_observation" CASCADE ;
+TRUNCATE "node_organization" CASCADE ;
+TRUNCATE "node_patient" CASCADE ;
+TRUNCATE "node_practitioner" CASCADE ;
+TRUNCATE "node_practitionerrole" CASCADE ;
+TRUNCATE "node_procedure" CASCADE ;
+TRUNCATE "node_researchstudy" CASCADE ;
+TRUNCATE "node_researchsubject" CASCADE ;
+TRUNCATE "node_root" CASCADE ;
+TRUNCATE "node_specimen" CASCADE ;
+TRUNCATE "node_task" CASCADE ;
+
+\c indexd_db;
+TRUNCATE "base_version" CASCADE ;
+TRUNCATE "index_record" CASCADE ;
+TRUNCATE "index_record_authz" CASCADE ;
+TRUNCATE "index_record_hash" CASCADE ;
+TRUNCATE "index_record_metadata" CASCADE ;
+TRUNCATE "index_record_url" CASCADE ;
+
diff --git a/etl/tube_lite b/etl/tube_lite
new file mode 100755
index 00000000..13c1d9b3
--- /dev/null
+++ b/etl/tube_lite
@@ -0,0 +1,698 @@
+#!/usr/bin/env python3
+
+"""A lightweight replacement for gen3's spark/tube."""
+
+from datetime import datetime
+
+import click
+import requests
+from elasticsearch import Elasticsearch
+from elasticsearch.helpers import bulk
+from gen3.auth import Gen3Auth
+from gen3.submission import Gen3Submission
+from jsonpath_ng import parse
+from more_itertools import peekable
+import logging
+
+FORMAT = '%(name)s %(process)d %(asctime)s %(message)s'
+logging.basicConfig(format=FORMAT, level=logging.INFO)
+logger = logging.getLogger("root")
+
+DEFAULT_ELASTIC = "http://esproxy-service:9200"
+DEFAULT_NAMESPACE = "gen3.aced.io"
+
+# gen3 graph-model query
+
+# graphql for FILE
+
+# performance improvement: only retrieve what we need in the index (coordinate with gitops.json)
+FILE_PROPERTIES = """
+ file_id: id
+ project_id
+ data_type
+ data_format
+ file_name
+ file_size
+ object_id
+ ga4gh_drs_uri
+ date
+ md5sum
+ state
+ submitter_id
+ file_category: category_0_coding_0_display
+ content_0_attachment_url
+ content_0_attachment_size
+"""
+
+# performance improvement: only retrieve what we need in the index (coordinate with gitops.json)
+PATIENT_PROPERTIES = """
+ id
+ submitter_id
+ project_id
+ gender
+ maritalStatus_coding_0_display
+ deceasedBoolean
+ deceasedDateTime
+ birthDate
+ patient_resource_type: resource_type
+ submitter_id
+ ombCategory: extension_0_extension_0_valueCoding_display
+ ombCategory_detail: extension_1_extension_0_valueCoding_display
+ us_core_birthsex: extension_3_valueCode
+ disability_adjusted_life_years: extension_5_valueDecimal
+ quality_adjusted_life_years: extension_6_valueDecimal
+
+"""
+
+FILE_GRAPHQL = """
+query ($first: Int!, $offset: Int!) {
+ patient(first: $first, offset: $offset) {
+ $PATIENT_PROPERTIES
+ document_references(first:1000) {
+ $FILE_PROPERTIES
+ }
+ }
+}
+""".replace('$FILE_PROPERTIES', FILE_PROPERTIES)\
+ .replace('$PATIENT_PROPERTIES', PATIENT_PROPERTIES)
+
+# graphql query for CASE
+# PATIENT_GRAPHQL = """
+# query ($first: Int!, $offset: Int!) {
+# patient(first: $first, offset: $offset) {
+# $PATIENT_PROPERTIES
+# }
+# }
+# """.replace('$PATIENT_PROPERTIES', PATIENT_PROPERTIES)\
+# .replace('$FILE_PROPERTIES', FILE_PROPERTIES)
+
+# (with_links:["observations", "conditions", "medication_requests"])
+
+ENCOUNTER_QUERY = """
+query ($first: Int!, $offset: Int!) {
+ encounter(first: $first, offset: $offset) {
+ project_id
+ encounter_id: id
+ encounter_type: type_0_coding_0_display
+ encounter_reason: reasonCode_0_coding_0_display
+ encounter_start: period_start
+ patient: subject_patient {
+ id
+ ombCategory: extension_0_extension_0_valueCoding_display
+ ombCategory_detail: extension_1_extension_0_valueCoding_display
+ us_core_birthsex: extension_3_valueCode
+ disability_adjusted_life_years: extension_5_valueDecimal
+ quality_adjusted_life_years: extension_6_valueDecimal
+ gender
+ maritalStatus: maritalStatus_coding_0_display
+ deceasedBoolean
+ deceasedDateTime
+ birthDate
+ }
+ conditions {
+ condition_id: id
+ code_coding_0_display
+ code_coding_0_system
+ code_coding_0_code
+ category_0_coding_0_display
+ onsetDateTime
+ abatementDateTime
+ verificationStatus_coding_0_code
+ clinicalStatus_coding_0_code
+ }
+ observations {
+ observation_id: id
+ category: category_0_coding_0_display
+ bodySite: bodySite_coding_0_code
+ code_display: code_coding_0_display
+ valueTime
+ valueString
+ valueBoolean
+ valueInteger
+ valueDateTime
+ valuePeriod_end
+ valuePeriod_start
+ valueQuantity_code
+ valueQuantity_unit
+ valueQuantity_value
+ valueCodeableConcept_coding_0_system
+ valueCodeableConcept_coding_0_code
+ valueCodeableConcept_coding_0_display
+ component_0_code_coding_0_code
+ component_0_code_coding_0_display
+ component_0_code_coding_0_system
+ component_0_valueTime
+ component_0_valueTime
+ component_0_valueString
+ component_0_valueBoolean
+ component_0_valueInteger
+ component_0_valueDateTime
+ component_0_valuePeriod_end
+ component_0_valuePeriod_start
+ component_0_valueQuantity_code
+ component_0_valueQuantity_unit
+ component_0_valueQuantity_value
+ component_1_code_coding_0_code
+ component_1_code_coding_0_display
+ component_1_code_coding_0_system
+ component_1_valueTime
+ component_1_valueTime
+ component_1_valueString
+ component_1_valueBoolean
+ component_1_valueInteger
+ component_1_valueDateTime
+ component_1_valuePeriod_end
+ component_1_valuePeriod_start
+ component_1_valueQuantity_code
+ component_1_valueQuantity_unit
+ component_1_valueQuantity_value
+ }
+ medication_requests {
+ medication_id: id
+ medicationCodeableConcept_coding_0_code
+ medicationCodeableConcept_coding_0_system
+ medicationCodeableConcept_coding_0_display
+ }
+ procedures {
+ procedure_id: id
+ code_coding_0_code
+ code_coding_0_system
+ code_coding_0_display
+ }
+ }
+}
+
+"""
+
+PATIENT_GRAPHQL = str(ENCOUNTER_QUERY)
+
+
+def key_values(nested, key, normalize=True):
+ """Return all values of a key in nested, cast to None, and scalar if normalize"""
+ jsonpath_expression = parse(f"$..[{key}]")
+ matches = jsonpath_expression.find(nested)
+ if normalize:
+ if len(matches) == 0:
+ return None
+ if len(matches) == 1:
+ return matches[0].value
+ return [match.value for match in matches]
+
+
+def sum_key_values(nested, key):
+ """Return the sum of values for key."""
+ return sum(key_values(nested, key, normalize=False))
+
+
+def create_index_from_source(_source, _index, _type):
+ """Given an ES source dict, create ES index."""
+ mappings = {}
+ for k, v in _source.items():
+ if type(v).__name__ in ['str', 'NoneType']:
+ mappings[k] = {
+ "type": "keyword"
+ }
+ elif isinstance(v, list):
+ mappings[k] = {
+ "type": "text"
+ }
+ else:
+ # naive, there are probably other types
+ mappings[k] = {"type": "float"}
+ return {
+ "mappings": {_type: {"properties": mappings}}
+ }
+
+
+def submission_client(endpoint, refresh_file):
+ """Create authorized client."""
+ auth = Gen3Auth(endpoint, refresh_file=refresh_file)
+ assert auth, 'should return an auth client'
+ submission_client_ = Gen3Submission(endpoint, auth)
+ assert submission_client_, 'should return a submission client'
+ assert 'delete_program' in dir(submission_client_), 'missing delete_program'
+ assert 'create_program' in dir(submission_client_), 'missing create_program'
+ return submission_client_
+
+
+def drop_file_indexes(elastic=DEFAULT_ELASTIC, name_space=DEFAULT_NAMESPACE):
+ """Drop the es indexes."""
+ return {"method": 'DELETE', "url": f'{elastic}/{name_space}_file_0'}
+
+
+def write_file_array_aliases(elastic=DEFAULT_ELASTIC, name_space=DEFAULT_NAMESPACE):
+ """Write the array aliases."""
+ # EXPECTED_ALIASES = {
+ # ".kibana_1": {
+ # "aliases": {
+ # ".kibana": {}
+ # }
+ # },
+ # "etl-array-config_0": {
+ # "aliases": {
+ # "etl-array-config": {},
+ # "etl_array-config": {},
+ # "time_2022-08-25T01:44:47.115494": {}
+ # }
+ # },
+ # "etl_0": {
+ # "aliases": {
+ # "etl": {},
+ # "time_2022-08-25T01:44:47.115494": {}
+ # }
+ # },
+ # "file-array-config_0": {
+ # "aliases": {
+ # "file-array-config": {},
+ # "file_array-config": {},
+ # "time_2022-08-25T01:44:47.115494": {}
+ # }
+ # },
+ # "file_0": {
+ # "aliases": {
+ # "file": {},
+ # "time_2022-08-25T01:44:47.115494": {}
+ # }
+ # }
+ # }
+ return {
+ "method": 'POST',
+ "url": f'{elastic}/_aliases',
+ "json": {
+ "actions": [{"add": {"index": f"{name_space}_file-array-config_0",
+ "alias": f"{name_space}_array-config"}},
+ {"add": {"index": f"{name_space}_file-array-config_0",
+ "alias": f"file_array-config"}},
+ ]}
+ }
+
+
+def write_patient_array_aliases(elastic=DEFAULT_ELASTIC, name_space=DEFAULT_NAMESPACE):
+ """Write the array aliases."""
+ # EXPECTED_ALIASES = {
+ # ".kibana_1": {
+ # "aliases": {
+ # ".kibana": {}
+ # }
+ # },
+ # "etl-array-config_0": {
+ # "aliases": {
+ # "etl-array-config": {},
+ # "etl_array-config": {},
+ # "time_2022-08-25T01:44:47.115494": {}
+ # }
+ # },
+ # "etl_0": {
+ # "aliases": {
+ # "etl": {},
+ # "time_2022-08-25T01:44:47.115494": {}
+ # }
+ # },
+ # "file-array-config_0": {
+ # "aliases": {
+ # "file-array-config": {},
+ # "file_array-config": {},
+ # "time_2022-08-25T01:44:47.115494": {}
+ # }
+ # },
+ # "file_0": {
+ # "aliases": {
+ # "file": {},
+ # "time_2022-08-25T01:44:47.115494": {}
+ # }
+ # }
+ # }
+ return {
+ "method": 'POST',
+ "url": f'{elastic}/_aliases',
+ "json": {
+ "actions": [
+ {"add": {"index": f"{name_space}_case-array-config_0",
+ "alias": f"{name_space}_array-config"}},
+ {"add": {"index": f"{name_space}_case-array-config_0",
+ "alias": f"etl_array-config"}}
+ ]}
+ }
+
+
+def create_file_indexes(_source, elastic=DEFAULT_ELASTIC, name_space=DEFAULT_NAMESPACE):
+ """Create the es indexes."""
+ _index = f"{name_space}_file_0"
+ _type = "file"
+ return {
+ "method": 'PUT',
+ "url": f'{elastic}/{_index}',
+ "json": create_index_from_source(_source, _index, _type),
+ "index": _index,
+ "type": _type
+ }
+
+
+def write_file_array_config(elastic=DEFAULT_ELASTIC, name_space=DEFAULT_NAMESPACE):
+ """Write the array config."""
+ return {
+ "method": 'PUT',
+ "url": f'{elastic}/{name_space}_file-array-config_0/_doc/file',
+ "json": {"timestamp": datetime.now().isoformat(), "array": ["project_code", "program_name"]}
+ }
+
+
+# gen3.aced.io_array-config gen3.aced.io_file-array-config_0 - - -
+
+# case gen3.aced.io_case_0 - - -
+# etl gen3.aced.io_case_0 - - -
+# file gen3.aced.io_file_0 - - -
+
+# gen3.aced.io_case gen3.aced.io_case_0 - - -
+# gen3.aced.io_file gen3.aced.io_file_0 - - -
+# etl_array-config gen3.aced.io_case-array-config_0 - - -
+# gen3.aced.io_array-config gen3.aced.io_case-array-config_0 - - -
+# .kibana .kibana_1 - - -
+
+
+def write_file_alias_config(elastic=DEFAULT_ELASTIC, name_space=DEFAULT_NAMESPACE):
+ """Write the alias config."""
+ return {
+ "method": 'POST',
+ "url": f'{elastic}/_aliases',
+ "json": {"actions": [{"add": {"index": f"{name_space}_file_0", "alias": f"file"}}]} # {name_space}_
+ }
+
+
+def write_file_data(row, elastic=DEFAULT_ELASTIC, name_space=DEFAULT_NAMESPACE):
+ """Write data."""
+ return {
+ "method": 'POST',
+ "url": f'{elastic}/{name_space}_file_0/file',
+ "json": row
+ }
+
+
+def drop_patient_indexes(elastic=DEFAULT_ELASTIC, name_space=DEFAULT_NAMESPACE):
+ """Drop the es indexes."""
+ return {"method": 'DELETE', "url": f'{elastic}/{name_space}_case_0'}
+
+
+def create_patient_indexes(_source, elastic=DEFAULT_ELASTIC, name_space=DEFAULT_NAMESPACE):
+ """Create the es indexes."""
+ _index = f"{name_space}_case_0"
+ _type = "case"
+ return {
+ "method": 'PUT',
+ "url": f'{elastic}/{_index}',
+ "json": create_index_from_source(_source, _index, _type),
+ "index": _index,
+ "type": _type
+ }
+
+
+def write_patient_array_config(elastic=DEFAULT_ELASTIC, name_space=DEFAULT_NAMESPACE):
+ """Write the array config."""
+ # { "timestamp": "2021-09-18T17:10:49.107081", "array": [ "_file_id" ] }
+ return {
+ "method": 'PUT',
+ "url": f'{elastic}/{name_space}_case-array-config_0/_doc/etl',
+ "json": {"timestamp": datetime.now().isoformat(), "array": ['data_format', 'data_type', '_file_id', 'medications', 'conditions']}
+ }
+
+
+def write_patient_alias_config(elastic=DEFAULT_ELASTIC, name_space=DEFAULT_NAMESPACE):
+ """Write the alias config."""
+ return {
+ "method": 'POST',
+ "url": f'{elastic}/_aliases',
+ "json": {"actions": [{"add": {"index": f"{name_space}_case_0", "alias": f"etl"}}]}
+ }
+
+
+def write_patient_data(row, elastic=DEFAULT_ELASTIC, name_space=DEFAULT_NAMESPACE):
+ """Write data."""
+ return {
+ "method": 'POST',
+ "url": f'{elastic}/{name_space}_case_0/case',
+ "json": row
+ }
+
+
+def read_files(sc, batch_size):
+ """Read file records and their ancestors from gen3, map to elastic search."""
+ first = batch_size
+ offset = 0
+ c = 0
+ while True:
+ graphql_variables = {"first": first, "offset": offset}
+ graph_ql_response = sc.query(FILE_GRAPHQL, variables=graphql_variables)
+ c += 1
+ has_data = False
+
+ if 'data' in graph_ql_response:
+ sum_data = 0
+ for k in graph_ql_response['data']:
+ sum_data += len(graph_ql_response['data'][k])
+ has_data = sum_data > 0
+ if not has_data:
+ break
+ offset += batch_size
+
+ # we get patient with array of document_references
+ for patient in graph_ql_response['data']['patient']:
+ assert 'project_id' in patient, (patient, patient.keys())
+ program, project = patient['project_id'].split('-')
+ for document_reference in patient['document_references']:
+ document_reference["auth_resource_path"] = f"/programs/{program}/projects/{project}"
+ for k, v in patient.items():
+ if k == 'document_references':
+ continue
+ document_reference[f"patient_{k}"] = v
+
+ # map fields hard coded by windmill portal
+ # data_types & data_format
+ if document_reference['content_0_attachment_url']:
+ document_reference['data_type'] = document_reference['content_0_attachment_url'].split('.')[-1]
+ else:
+ document_reference['data_type'] = 'txt'
+
+ if document_reference['data_type'] in ['csv']:
+ document_reference['data_format'] = 'variants'
+ if document_reference['data_type'] in ['dcm']:
+ document_reference['data_format'] = 'imaging'
+ if document_reference['data_type'] in ['txt']:
+ document_reference['data_format'] = 'note'
+
+ document_reference['file_name'] = document_reference['content_0_attachment_url']
+ document_reference['file_size'] = 0
+ if document_reference['content_0_attachment_size']:
+ document_reference['file_size'] = int(document_reference['content_0_attachment_size'])
+
+ yield document_reference
+
+
+def read_patients(sc, batch_size, start):
+ """Read patient records and their descendants from gen3."""
+ first = batch_size
+ offset = 0
+ if start:
+ offset = int(start)
+ c = 0
+ while True:
+ # pagination
+ graphql_variables = {"first": first, "offset": offset}
+ # logger.info((PATIENT_GRAPHQL, graphql_variables))
+ r = sc.query(PATIENT_GRAPHQL, variables=graphql_variables)
+ c += 1
+
+ if 'data' not in r or 'encounter' not in r['data'] or len(r['data']['encounter']) == 0:
+ break
+ offset += batch_size
+ for encounter in r['data']['encounter']:
+ assert 'project_id' in encounter, (encounter, encounter.keys())
+ program, project = encounter['project_id'].split('-')
+ # denormalize by patient
+ patient = dict({'project_id': encounter['project_id']})
+ patient["auth_resource_path"] = f"/programs/{program}/projects/{project}"
+ # get scalars for encounter
+
+ for k, v in encounter.items():
+ if not k.startswith('encounter'):
+ continue
+ patient[k] = v
+ # get embedded subject
+ # expecting only one subject
+ for k, v in encounter['patient'][0].items():
+ patient[f"patient_{k}"] = v
+ patient["_case_id"] = encounter['patient'][0]['id'] # TODO why this variable ??? in gitops.json ?
+ # create an array of condition names
+ patient['conditions'] = [c['code_coding_0_display'] for c in encounter['conditions']]
+ # create an array of medication names
+ patient['medications'] = [c['medicationCodeableConcept_coding_0_display'] for c in encounter['medication_requests']]
+ # denormalize by observation
+ for observation in encounter['observations']:
+ patient_observation = dict(patient)
+ for k, v in observation.items():
+ patient_observation[k] = v
+ yield patient_observation
+
+
+def write_dict(output, d):
+ """Write a dict to the output."""
+ output.write(str(d))
+ output.write("\n")
+
+
+def write_http(session, _params, raise_for_status=True):
+ """Write a dict to the session."""
+ r = session.request(**_params)
+ if raise_for_status:
+ if r.status_code > 300:
+ print("TEXT_PARAMS", _params)
+ print("TEXT", r.text)
+ r.raise_for_status()
+
+
+def write_patient_bulk_http(elastic, index, limit, doc_type, generator):
+ """Use efficient method to write to elastic"""
+ counter = 0
+
+ def _bulker(generator_, counter_=counter):
+ for dict_ in generator_:
+ if limit and counter_ > limit:
+ break # for testing
+ yield {
+ '_index': index,
+ '_op_type': 'index',
+ '_type': doc_type,
+ '_source': dict_
+ }
+ counter_ += 1
+ if counter_ % 100 == 0:
+ logger.info(counter_)
+
+ logger.info('Fetching first record.')
+ generator = peekable(generator)
+ first_patient = generator.peek()
+ logger.info('Creating patient indices.')
+
+ index_dict = create_patient_indexes(first_patient, "FOO")
+ elastic.indices.create(index=index_dict['index'], body=index_dict['json'])
+
+ logger.info(f'Writing bulk to {index} limit {limit}.')
+ result = bulk(client=elastic,
+ actions=(d for d in _bulker(generator)),
+ request_timeout=120)
+
+
+def write_file_bulk_http(elastic, index, limit, doc_type, generator):
+ """Use efficient method to write to elastic"""
+ counter = 0
+
+ def _bulker(generator_, counter_=counter):
+ for dict_ in generator_:
+ if limit and counter_ > limit:
+ break # for testing
+ yield {
+ '_index': index,
+ '_op_type': 'index',
+ '_type': doc_type,
+ '_source': dict_
+ }
+ counter_ += 1
+ if counter_ % 100 == 0:
+ logger.info(counter_)
+
+ logger.info('Fetching first record.')
+ generator = peekable(generator)
+ first_file = generator.peek()
+ logger.info('Creating file indices.')
+
+ index_dict = create_file_indexes(first_file, "FOO")
+ elastic.indices.create(index=index_dict['index'], body=index_dict['json'])
+
+ logger.info(f'Writing bulk to {index} limit {limit}.')
+ result = bulk(client=elastic,
+ actions=(d for d in _bulker(generator)),
+ request_timeout=120)
+
+
+@click.command()
+@click.option('--endpoint', type=str, help='Gen3 host base url.')
+@click.option('--credentials_path', type=str, help='Path to gen3 credentials.')
+@click.option('--batch_size', type=int, default=500, help='Number of records to read from gen3 at a time (500.')
+@click.option('--output_path', type=str, default=None, help='For debugging, write the output to this path')
+@click.option('--elastic', type=str, default=None, help='Write directly to elastic host')
+@click.option('--entity', type=str, default=None, help='One of file | observation')
+@click.option('--limit',
+ default=None,
+ show_default=True,
+ help='Max number of rows per index.')
+@click.option('--start',
+ default=None,
+ show_default=True,
+ help='Max number of rows per index.')
+def etl(credentials_path, endpoint, output_path, batch_size, elastic, limit, entity, start):
+ """Extract file centric index from Gen3, create elastic search index."""
+ # check destination
+ assert output_path or elastic, "Please set either elastic url or output_path file path"
+ assert entity, "Please specify file | observation"
+ # connect to source (gen3)
+ sc = submission_client(endpoint, credentials_path)
+
+ if limit:
+ limit = int(limit)
+ # create a handy little function that writes to either file or session
+ output_stream = None
+ write_method = None
+ if output_path:
+ output_stream = open(output_path, "w")
+ write_method = write_dict
+ else:
+ output_stream = requests.sessions.Session()
+ write_method = write_http
+
+ def _writer(data):
+ """Write to destination"""
+ write_method(output_stream, data)
+
+ _es = Elasticsearch([elastic], request_timeout=120)
+
+ global logger
+ logger = logging.getLogger(entity)
+
+ #
+ # PATIENT centric index
+ #
+ if entity.lower() == 'observation':
+ logger.info(f'Reading patients. batch_size {batch_size}')
+
+ write_patient_bulk_http(elastic=_es, index=f"{DEFAULT_NAMESPACE}_case_0", doc_type='case', limit=limit,
+ generator=read_patients(sc, batch_size, start))
+
+ _writer(write_patient_array_config(elastic))
+ _writer(write_patient_alias_config(elastic))
+
+ _writer(write_patient_array_aliases(elastic))
+ #
+ # FILE centric index
+ #
+
+ # assumes guppy-setup dropped ES indices
+ #_writer(drop_file_indexes(elastic))
+
+ # write data
+
+ if entity.lower() == 'file':
+ logger.info(f'Reading files. batch_size {batch_size}')
+ write_file_bulk_http(elastic=_es, index=f"{DEFAULT_NAMESPACE}_file_0", doc_type='file', limit=limit,
+ generator=read_files(sc, batch_size))
+
+ _writer(write_file_array_config(elastic))
+ _writer(write_file_alias_config(elastic))
+
+ _writer(write_file_array_aliases(elastic))
+
+ # cleanup
+ output_stream.close()
+ logger.info('done')
+
+
+if __name__ == '__main__':
+ etl()
diff --git a/examples/Secrets/aggregate_config.json b/examples/Secrets/aggregate_config.json
new file mode 100644
index 00000000..e6ee200c
--- /dev/null
+++ b/examples/Secrets/aggregate_config.json
@@ -0,0 +1,57 @@
+{
+ "gen3_commons": {
+ "IBD Commons": {
+ "mds_url": "https://ibdgc.datacommons.io",
+ "commons_url" : "ibdgc.datacommons.io",
+ "study_data_field" : "my_metadata",
+ "guid_type" : "my_metadata",
+ "columns_to_fields": {
+ "_subjects_count" : "subjects_count",
+ "study_description" : "brief_summary",
+ "short_name": "dataset_title",
+ "full_name": "dataset_title"
+ }
+ },
+ "BioData Catalyst": {
+ "mds_url": "https://gen3.biodatacatalyst.nhlbi.nih.gov",
+ "commons_url" : "gen3.biodatacatalyst.nhlbi.nih.gov",
+ "columns_to_fields": {
+ "short_name": "name",
+ "_unique_id" : "study_id"
+ }
+ },
+ "MIDRC": {
+ "mds_url": "https://data.midrc.org",
+ "commons_url" : "data.midrc.org",
+ "study_data_field" : "discovery_metadata",
+ "columns_to_fields": {
+ "_subjects_count" : "cases_count",
+ "study_description" : "research_description",
+ "_unique_id": "study_id"
+ }
+ },
+ "NIAID ClinicalData": {
+ "mds_url": "https://accessclinicaldata.niaid.nih.gov",
+ "commons_url" : "accessclinicaldata.niaid.nih.gov",
+ "study_data_field" : "my_metadata",
+ "guid_type" : "my_metadata",
+ "columns_to_fields": {
+ "full_name": "title",
+ "study_id" : "nct_number",
+ "_unique_id": "nct_number",
+ "study_description" : "brief_summary"
+ }
+ },
+ "AnVIL": {
+ "mds_url": "https://internalstaging.theanvil.io",
+ "commons_url": "gen3.theanvil.io",
+ "columns_to_fields": {
+ "name": "name",
+ "full_name": "full_name",
+ "_subjects_count" : "_subjects_count",
+ "_unique_id" : "study_id",
+ "study_description" : "study_description"
+ }
+ }
+ }
+}
\ No newline at end of file
diff --git a/examples/Secrets/etlMapping.yaml b/examples/Secrets/etlMapping.yaml
new file mode 100644
index 00000000..d1b64c0f
--- /dev/null
+++ b/examples/Secrets/etlMapping.yaml
@@ -0,0 +1,86 @@
+mappings:
+ - name: etl
+ doc_type: case
+ type: aggregator
+ root: case
+ props:
+ - name: submitter_id
+ - name: project_id
+ - name: disease_type
+ - name: primary_site
+ flatten_props:
+ - path: demographics
+ props:
+ - name: gender
+ value_mappings:
+ - female: F
+ - male: M
+ - name: race
+ value_mappings:
+ - american indian or alaskan native: Indian
+ - name: ethnicity
+ - name: year_of_birth
+ aggregated_props:
+ - name: _samples_count
+ path: samples
+ fn: count
+ - name: _aliquots_count
+ path: samples.aliquots
+ fn: count
+ - name: _submitted_methylations_count
+ path: samples.aliquots.submitted_methylation_files
+ fn: count
+ - name: _submitted_copy_number_files_on_aliquots_count
+ path: samples.aliquots.submitted_copy_number_files
+ fn: count
+ - name: _read_groups_count
+ path: samples.aliquots.read_groups
+ fn: count
+ - name: _submitted_aligned_reads_count
+ path: samples.aliquots.read_groups.submitted_aligned_reads_files
+ fn: count
+ - name: _submitted_unaligned_reads_count
+ path: samples.aliquots.read_groups.submitted_unaligned_reads_files
+ fn: count
+ - name: _submitted_copy_number_files_on_read_groups_count
+ path: samples.aliquots.read_groups.submitted_copy_number_files
+ fn: count
+ - name: _submitted_somatic_mutations_count
+ path: samples.aliquots.read_groups.submitted_somatic_mutations
+ fn: count
+ joining_props:
+ - index: file
+ join_on: _case_id
+ props:
+ - name: data_format
+ src: data_format
+ fn: set
+ - name: data_type
+ src: data_type
+ fn: set
+ - name: _file_id
+ src: file_id
+ fn: set
+ - name: file
+ doc_type: file
+ type: collector
+ root: None
+ category: data_file
+ props:
+ - name: object_id
+ - name: md5sum
+ - name: file_name
+ - name: file_size
+ - name: data_format
+ - name: data_type
+ - name: state
+ injecting_props:
+ case:
+ props:
+ - name: _case_id
+ src: id
+ fn: set
+ - name: project_id
+ target_nodes:
+ - name: slide_image
+ path: slides.samples.cases
diff --git a/examples/Secrets/etl_creds.json b/examples/Secrets/etl_creds.json
new file mode 100644
index 00000000..eb8428a8
--- /dev/null
+++ b/examples/Secrets/etl_creds.json
@@ -0,0 +1,7 @@
+{
+ "db_host": "postgres",
+ "db_username": "sheepdog_user",
+ "db_password": "sheepdog_pass",
+ "db_database": "metadata_db"
+ }
+
\ No newline at end of file
diff --git a/examples/Secrets/fence-config.yaml b/examples/Secrets/fence-config.yaml
new file mode 100644
index 00000000..a4e98077
--- /dev/null
+++ b/examples/Secrets/fence-config.yaml
@@ -0,0 +1,491 @@
+---
+############################### Fence Configuration ####################################
+# This file contains various configurations for the fence microservice.
+#
+# README:
+# - This is initially configured for minimal local development with reasonable defaults.
+# - Descriptions for each of the configurations (if any) will be *above* the variable as
+# comments.
+# - Some configuration variables will have examples commented out below them.
+# - This is broken up into 2 main sections for REQUIRED and OPTIONAL configurations.
+# - Optional configs will note what features or endpoints they support
+# - Underneath each main section the variables are logically grouped under named
+# sections.
+#
+# NOTE: Login is NOT ready out of the box. Fill out REQUIRED configurations first
+
+########################################################################################
+# REQUIRED CONFIGURATIONS #
+########################################################################################
+
+# //////////////////////////////////////////////////////////////////////////////////////
+# GENERAL
+# - Fill out all variables!
+# //////////////////////////////////////////////////////////////////////////////////////
+APP_NAME: 'Gen3 Data Commons'
+# Where fence microservice is deployed
+BASE_URL: 'https://aced-training.compbio.ohsu.edu/user'
+# postgres db to connect to
+# connection url format:
+# postgresql://[user[:password]@][netloc][:port][/dbname]
+DB: 'postgresql://fence_user:fence_pass@postgres:5432/fence_db'
+
+# A URL-safe base64-encoded 32-byte key for encrypting keys in db
+# in python you can use the following script to generate one:
+# import base64
+# import os
+# key = base64.urlsafe_b64encode(os.urandom(32))
+# print(key)
+ENCRYPTION_KEY: 'mHgNKggngJrLQUoTsYh4AFFhTlrO7yKTZ22ELqS347U='
+
+# //////////////////////////////////////////////////////////////////////////////////////
+# DEBUG & SECURITY SETTINGS
+# - Modify based on whether you're in a dev environment or in production
+# //////////////////////////////////////////////////////////////////////////////////////
+# flask's debug setting
+# WARNING: DO NOT ENABLE IN PRODUCTION
+DEBUG: true
+# if true, will automatically login a user with username "test"
+MOCK_AUTH: true
+# if true, will only fake a successful login response from Google in /login/google
+MOCK_GOOGLE_AUTH: false
+# if true, will ignore anything configured in STORAGE_CREDENTIALS
+MOCK_STORAGE: true
+# allow OIDC traffic on http for development. By default it requires https.
+#
+# WARNING: ONLY set to true when fence will be deployed in such a way that it will
+# ONLY receive traffic from internal clients and can safely use HTTP.
+AUTHLIB_INSECURE_TRANSPORT: false
+
+# set if you want browsers to only send cookies with requests over HTTPS
+SESSION_COOKIE_SECURE: true
+
+ENABLE_CSRF_PROTECTION: true
+
+# //////////////////////////////////////////////////////////////////////////////////////
+# OPEN ID CONNECT (OIDC) CLIENTS
+# - Fully configure at least one client so login works
+# //////////////////////////////////////////////////////////////////////////////////////
+OPENID_CONNECT:
+ # These Google values must be obtained from Google's Cloud Console
+ # Follow: https://developers.google.com/identity/protocols/OpenIDConnect
+ #
+ # You'll need to obtain a Client ID and Client Secret. Set the redirect URIs
+ # in Google to be '{{BASE_URL}}/login/google/login', but expand BASE_URL to
+ # whatever you set it to above.
+ google:
+ client_id: ''
+ client_secret: ''
+ # this is be the allowed redirect back to fence, should not need to change
+ redirect_url: '{{BASE_URL}}/login/google/login/'
+ # Support for multi-tenant fence (another fence is this fence's IDP)
+ # If this fence instance is a client of another fence, fill this cfg out.
+ # REMOVE if not needed
+ fence:
+ # this api_base_url should be the root url for the OTHER fence
+ # something like: https://example.com
+ api_base_url: ''
+ # this client_id and client_secret should be obtained by registering THIS fence as
+ # a new client of the OTHER fence
+ client_id: ''
+ client_secret: ''
+ client_kwargs:
+ # openid is required to use OIDC flow
+ scope: 'openid'
+ # callback after logging in through the other fence
+ redirect_uri: '{{BASE_URL}}/login/fence/login'
+ # The next 3 should not need to be changed if the provider is following
+ # Oauth2 endpoint naming conventions
+ authorize_url: '{{api_base_url}}/oauth2/authorize'
+ access_token_url: '{{api_base_url}}/oauth2/token'
+ refresh_token_url: '{{api_base_url}}/oauth2/token'
+ # Custom name to display for consent screens. If not provided, will use `fence`.
+ # If the other fence is using NIH Login, you should make name: `NIH Login`
+ name: ''
+
+# //////////////////////////////////////////////////////////////////////////////////////
+# LOGIN
+# - Modify based on which OIDC client(s) you configured above
+# - NOTE: You can have multiple IDPs for users to login with, but one has to be set
+# as the default
+# //////////////////////////////////////////////////////////////////////////////////////
+# Login url for identity provider (IDP):
+# Google? Use: '{{BASE_URL}}/login/google'
+# Multi-tenant fence (e.g. another fence instance)? Use: '{{BASE_URL}}/login/fence'
+# Sibboleth? Use: '{{BASE_URL}}/login/shib'
+DEFAULT_LOGIN_URL: '{{BASE_URL}}/login/google'
+
+# Which Identity Provider fence will/can use
+#
+# See ``fence/blueprints/login/__init__.py`` for which identity providers can
+# be loaded.
+#
+LOGIN_OPTIONS:
+ - name: 'Login from Google'
+ desc: 'description'
+ idp: google
+
+ # secondary: True
+ # - name: 'ORCID Login'
+ # idp: orcid
+ # - name: 'Microsoft Login'
+ # idp: microsoft
+ # # Cognito login: You may want to edit the name to reflect Cognito's IdP,
+ # # especially if Cognito is only using one IdP
+ # - name: 'Login from Cognito'
+ # desc: 'Amazon Cognito login'
+ # idp: cognito
+ # - name: 'Login from RAS'
+ # idp: ras
+ # - name: 'NIH Login'
+ # idp: fence
+ # fence_idp: shibboleth
+ # - name: 'ORCID Login through other Fence'
+ # idp: fence
+ # fence_idp: orcid
+ # - name: 'InCommon Login'
+ # idp: fence
+ # fence_idp: shibboleth
+ # # "shib_idps" can be '*' or a list of one or more entity IDs
+ # shib_idps:
+ # - urn:mace:incommon:nih.gov
+ # - urn:mace:incommon:uchicago.edu
+
+
+DEFAULT_LOGIN_IDP: google
+
+# //////////////////////////////////////////////////////////////////////////////////////
+# LIBRARY CONFIGURATION (authlib & flask)
+# - Already contains reasonable defaults
+# //////////////////////////////////////////////////////////////////////////////////////
+# authlib-specific configs for OIDC flow and JWTs
+# NOTE: the OAUTH2_JWT_KEY cfg gets set automatically by fence if keys are setup
+# correctly
+OAUTH2_JWT_ALG: 'RS256'
+OAUTH2_JWT_ENABLED: true
+OAUTH2_JWT_ISS: '{{BASE_URL}}'
+OAUTH2_PROVIDER_ERROR_URI: '/api/oauth2/errors'
+
+# used for flask, "path mounted under by the application / web server"
+# since we deploy as microservices, fence is typically under {{base}}/user
+# this is also why our BASE_URL default ends in /user
+APPLICATION_ROOT: '/user'
+
+
+# //////////////////////////////////////////////////////////////////////////////////////
+# Tokens, Lifetimes, & Expirations
+# - Already contains reasonable defaults
+# //////////////////////////////////////////////////////////////////////////////////////
+# The name of the browser cookie in which the access token will be stored.
+ACCESS_TOKEN_COOKIE_NAME: "access_token"
+
+# The name of the browser cookie in which the session token will be stored.
+# Note that the session token also stores information for the
+# ``flask.session`` in the ``context`` field of the token.
+SESSION_COOKIE_NAME: "fence"
+
+OAUTH2_TOKEN_EXPIRES_IN:
+ "authorization_code": 1200
+ "implicit": 1200
+
+# The number of seconds after an access token is issued until it expires.
+ACCESS_TOKEN_EXPIRES_IN: 1200
+
+# The number of seconds after a refresh token is issued until it expires.
+REFRESH_TOKEN_EXPIRES_IN: 2592000
+
+# The number of seconds after which a browser session is considered stale.
+SESSION_TIMEOUT: 1800
+
+# The maximum session lifetime in seconds.
+SESSION_LIFETIME: 28800
+
+# The number of seconds the user's Google service account key used for
+# url signing will last before being expired/rotated
+# 30 days: 2592000 seconds
+GOOGLE_SERVICE_ACCOUNT_KEY_FOR_URL_SIGNING_EXPIRES_IN: 2592000
+
+# The number of seconds after a User's Google Service account is added to bucket
+# access until it expires.
+# 7 days: 604800 seconds
+GOOGLE_USER_SERVICE_ACCOUNT_ACCESS_EXPIRES_IN: 604800
+
+# The number of seconds after a User's Google account is added to bucket
+# access until it expires.
+GOOGLE_ACCOUNT_ACCESS_EXPIRES_IN: 86400
+
+# The number of seconds after a pre-signed url is issued until it expires.
+MAX_PRESIGNED_URL_TTL: 3600
+
+# The number of seconds after an API KEY is issued until it expires.
+MAX_API_KEY_TTL: 2592000
+
+# The number of seconds after an access token is issued until it expires.
+MAX_ACCESS_TOKEN_TTL: 3600
+
+
+########################################################################################
+# OPTIONAL CONFIGURATIONS #
+########################################################################################
+
+# //////////////////////////////////////////////////////////////////////////////////////
+# SUPPORT INFO
+# //////////////////////////////////////////////////////////////////////////////////////
+# If you want an email address to show up when an unhandled error occurs, provide one
+# here. Something like: support@example.com
+SUPPORT_EMAIL_FOR_ERRORS: null
+
+# //////////////////////////////////////////////////////////////////////////////////////
+# SHIBBOLETH
+# - Support using `shibboleth` in ENABLED_IDENTITY_PROVIDERS
+# - Contains defaults for using NIH's Login.
+# //////////////////////////////////////////////////////////////////////////////////////
+# assumes shibboleth is deployed under {{BASE_URL}}/shibboleth
+SHIBBOLETH_HEADER: 'persistent_id'
+SSO_URL: 'https://auth.nih.gov/affwebservices/public/saml2sso?SPID={{BASE_URL}}/shibboleth&RelayState='
+ITRUST_GLOBAL_LOGOUT: 'https://auth.nih.gov/siteminderagent/smlogout.asp?mode=nih&AppReturnUrl='
+
+# //////////////////////////////////////////////////////////////////////////////////////
+# dbGaP USER SYNCING SUPPORT
+# - Support syncing authorization information from dbGaP
+# //////////////////////////////////////////////////////////////////////////////////////
+# "dbGaP project serves as an access gateway for researchers seeking to gain
+# access to genotype and phenotype data"
+#
+# User syncing and access can also be done throught a User Access file. See
+# fence's README for more information
+dbGaP:
+ info:
+ host: ''
+ username: ''
+ password: ''
+ port: 22
+ proxy: ''
+ proxy_user: ''
+ protocol: 'sftp'
+ decrypt_key: ''
+ parse_consent_code: true
+
+# //////////////////////////////////////////////////////////////////////////////////////
+# STORAGE BACKENDS AND CREDENTIALS
+# - Optional: Used for `/admin` & `/credentials` endpoints for user management.
+# Also used during User Syncing process to automate managing Storage
+# access for users.
+# //////////////////////////////////////////////////////////////////////////////////////
+# Configuration for various storage systems for the backend
+# NOTE: Remove the [] and supply backends if needed. Example in comments below
+STORAGE_CREDENTIALS: []
+# Google Cloud Storage backend
+#
+# 'google':
+# backend: 'google'
+# # this should be the project id where the Google Groups for data access are managed
+# google_project_id: 'some-project-id-12378923'
+
+# Cleversafe data storage backend
+#
+# 'cleversafe-server-a':
+# backend: 'cleversafe'
+# aws_access_key_id: ''
+# aws_secret_access_key: ''
+# host: 'somemanager.osdc.io'
+# public_host: 'someobjstore.example.com'
+# port: 443
+# is_secure: true
+# username: 'someone'
+# password: 'somepass'
+# is_mocked: true
+
+# //////////////////////////////////////////////////////////////////////////////////////
+# AWS BUCKETS AND CREDENTIALS
+# - Support `/data` endpoints
+# //////////////////////////////////////////////////////////////////////////////////////
+
+AWS_CREDENTIALS:
+# NOTE: Remove the {} and supply creds if needed. Example in comments below
+ 'TEST-USER':
+ aws_access_key_id: 'test'
+ aws_secret_access_key: 'testtest'
+# 'CRED2':
+# aws_access_key_id: ''
+# aws_secret_access_key: ''
+
+
+# NOTE: the region is optional for s3_buckets, however it should be specified to avoid a
+# call to GetBucketLocation which you make lack the AWS ACLs for.
+S3_BUCKETS:
+ aced-default:
+ cred: 'TEST-USER'
+ endpoint_url: 'https://minio-default.compbio.ohsu.edu'
+ region: us-east-1
+ aced-ohsu:
+ cred: 'TEST-USER'
+ endpoint_url: 'https://minio-ohsu.compbio.ohsu.edu'
+ region: us-east-1
+ aced-ucl:
+ cred: 'TEST-USER'
+ endpoint_url: 'https://minio-ucl.compbio.ohsu.edu'
+ region: us-east-1
+ aced-manchester:
+ cred: 'TEST-USER'
+ endpoint_url: 'https://minio-manchester.compbio.ohsu.edu'
+ region: us-east-1
+ aced-stanford:
+ cred: 'TEST-USER'
+ endpoint_url: 'https://minio-stanford.compbio.ohsu.edu'
+ region: us-east-1
+
+# Configure which buckets fence is allowed to write to
+ALLOWED_DATA_UPLOAD_BUCKETS: ['aced-default', 'aced-ohsu', 'aced-ucl', 'aced-manchester', 'aced-stanford']
+
+# bucket2:
+# cred: 'CRED2'
+# bucket3:
+# cred: '*'
+# bucket4:
+# cred: 'CRED1'
+# role-arn: 'arn:aws:iam::role1'
+
+# `DATA_UPLOAD_BUCKET` specifies an S3 bucket to which data files are uploaded,
+# using the `/data/upload` endpoint. This must be one of the first keys under
+# `S3_BUCKETS` (since these are the buckets fence has credentials for).
+#DATA_UPLOAD_BUCKET: 'bucket1'
+DATA_UPLOAD_BUCKET: aced-default
+
+# //////////////////////////////////////////////////////////////////////////////////////
+# PROXY
+# - Optional: If the api is behind firewall that needs to set http proxy
+# //////////////////////////////////////////////////////////////////////////////////////
+# NOTE: leave as-is to not use proxy
+# this is only used by the Google Oauth2Client at the moment if provided
+HTTP_PROXY:
+ host: null
+ port: 3128
+
+# //////////////////////////////////////////////////////////////////////////////////////
+# MICROSERVICE PATHS
+# - Support `/data` endpoints & `/rbac` endpoints
+# //////////////////////////////////////////////////////////////////////////////////////
+# url where indexd microservice is running (for signed urls primarily)
+# NOTE: Leaving as null will force fence to default to {{BASE_URL}}/index
+# example value: 'https://example.com/index'
+INDEXD: http://indexd-service
+
+# this is the username which fence uses to make authenticated requests to indexd
+INDEXD_USERNAME: 'indexd_client'
+# this is the password which fence uses to make authenticated requests to indexd
+INDEXD_PASSWORD: 'indexd_client_pass'
+
+# url where role-based access control microservice is running
+ARBORIST: http://arborist-service
+
+# //////////////////////////////////////////////////////////////////////////////////////
+# CLOUD API LIBRARY (CIRRUS) CONFIGURATION
+# - Support Google Data Access Methods
+# //////////////////////////////////////////////////////////////////////////////////////
+# Setting this up allows fence to create buckets, manage Google groups, etc.
+# See directions here for setting up cirrus: https://github.com/uc-cdis/cirrus
+CIRRUS_CFG:
+ GOOGLE_API_KEY: ''
+ GOOGLE_PROJECT_ID: ''
+ GOOGLE_APPLICATION_CREDENTIALS: ''
+ GOOGLE_STORAGE_CREDS: ''
+ GOOGLE_ADMIN_EMAIL: ''
+ GOOGLE_IDENTITY_DOMAIN: ''
+ GOOGLE_CLOUD_IDENTITY_ADMIN_EMAIL: ''
+
+# Prefix to namespace Google Groups on a single Cloud Identity (see cirrus
+# setup for more info on Cloud Identity)
+#
+# NOTE: Make this short! Less than 8 characters if possible. Google has
+# length restrictions on group names.
+GOOGLE_GROUP_PREFIX: ''
+
+# //////////////////////////////////////////////////////////////////////////////////////
+# EMAIL
+# - Support for sending emails from fence. Used for user certificates
+# and `/google/service_accounts` endpoints
+# //////////////////////////////////////////////////////////////////////////////////////
+# Gun Mail Service (for sending emails from fence)
+#
+# NOTE: Example in comments below
+GUN_MAIL:
+ 'datacommons.io':
+ smtp_hostname: 'smtp.mailgun.org'
+ api_key: ''
+ default_login: 'postmaster@mailgun.example.com'
+ api_url: 'https://api.mailgun.net/v3/mailgun.example.com'
+ smtp_password: ''
+
+# For emails regarding users certificates
+EMAIL_SERVER: 'aced-training.compbio.ohsu.edu'
+SEND_FROM: 'example@gmail.com'
+SEND_TO: 'example@gmail.com'
+
+# //////////////////////////////////////////////////////////////////////////////////////
+# DATA ACCESS: GOOGLE LINKING & SERVICE ACCOUNT REGISTRATION
+# - Support `/google/service_accounts` endpoints
+# //////////////////////////////////////////////////////////////////////////////////////
+# A Google Project with controlled data access will be determined INVALID if
+# if it has a parent organization UNLESS that parent organization's ID is in this
+# whitelist.
+#
+# NOTE: Remove the [] and Google Organization IDs if needed. Example in comments below
+WHITE_LISTED_GOOGLE_PARENT_ORGS: []
+# - '12345678910'
+
+# A Google Project with Google Service Accounts determined INVALID will result in the
+# the entire project being invalid UNLESS that service accounts's email is in this
+# whitelist.
+#
+# NOTE: Remove the [] and service account emails if needed. Example in comments below
+WHITE_LISTED_SERVICE_ACCOUNT_EMAILS: []
+# - 'example@developer.gserviceaccount.com'
+# - 'example@test.iam.gserviceaccount.com'
+
+# when service accounts or google projects are determined invalid, an email is sent
+# to the project owners. These settings are for that email
+REMOVE_SERVICE_ACCOUNT_EMAIL_NOTIFICATION:
+ # this domain MUST exist in GUN_MAIL config
+ enable: false
+ domain: 'example.com'
+ from: 'do-not-reply@example.com'
+ subject: 'User service account removal notification'
+ # the {} gets replaced dynamically in the Python code to be the Project ID
+ content: >
+ Service accounts were removed from access control data because some users or
+ service accounts of GCP Project {} are not authorized to access the data sets
+ associated to the service accounts, or do not adhere to the security policies.
+ # this admin email will be included as a recipient to *any* email to anyone about
+ # service account removal.
+ #
+ # WARNING: This is NOT a bcc so the email is visible to the end-user
+ admin:
+ - 'admin@example.edu'
+
+# Service account email domains that represent a service account that Google owns.
+# These are usually created when a sepcific GCP service is enabled.
+# This is used for Service Account Validation for Data Access.
+GOOGLE_MANAGED_SERVICE_ACCOUNT_DOMAINS:
+ - 'dataflow-service-producer-prod.iam.gserviceaccount.com'
+ - 'cloudbuild.gserviceaccount.com'
+ - 'cloud-ml.google.com.iam.gserviceaccount.com'
+ - 'container-engine-robot.iam.gserviceaccount.com'
+ - 'dataflow-service-producer-prod.iam.gserviceaccount.com'
+ - 'sourcerepo-service-accounts.iam.gserviceaccount.com'
+ - 'dataproc-accounts.iam.gserviceaccount.com'
+ - 'gae-api-prod.google.com.iam.gserviceaccount.com'
+ - 'genomics-api.google.com.iam.gserviceaccount.com'
+ - 'containerregistry.iam.gserviceaccount.com'
+ - 'container-analysis.iam.gserviceaccount.com'
+ - 'cloudservices.gserviceaccount.com'
+ - 'stackdriver-service.iam.gserviceaccount.com'
+ - 'appspot.gserviceaccount.com'
+ - 'partnercontent.gserviceaccount.com'
+ - 'trifacta-gcloud-prod.iam.gserviceaccount.com'
+ - 'gcf-admin-robot.iam.gserviceaccount.com'
+ - 'compute-system.iam.gserviceaccount.com'
+ - 'gcp-sa-websecurityscanner.iam.gserviceaccount.com'
+ - 'storage-transfer-service.iam.gserviceaccount.com'
+
+ENABLE_DB_MIGRATION: false
diff --git a/examples/Secrets/gitops-logo.png b/examples/Secrets/gitops-logo.png
new file mode 100644
index 00000000..33f5bb2f
Binary files /dev/null and b/examples/Secrets/gitops-logo.png differ
diff --git a/examples/Secrets/gitops.json b/examples/Secrets/gitops.json
new file mode 100644
index 00000000..e192df1f
--- /dev/null
+++ b/examples/Secrets/gitops.json
@@ -0,0 +1,605 @@
+{
+ "gaTrackingId": "UA-119127212-1",
+ "showArboristAuthzOnProfile": true,
+ "showFenceAuthzOnProfile": false,
+ "useArboristUI": true,
+
+
+ "featureFlags": {
+ "explorer": true,
+ "discovery": true,
+ "discoveryUseAggMDS": false
+ },
+
+
+ "explorerConfig":[
+
+ {
+
+ "tabTitle": "File",
+ "charts": {
+ "data_type": {
+ "chartType": "stackedBar",
+ "title": "File Type"
+ },
+ "data_format": {
+ "chartType": "stackedBar",
+ "title": "File Format"
+ },
+ "patient_gender": {
+ "chartType": "stackedBar",
+ "title": "Gender"
+ },
+ "patient_maritalStatus_coding_0_display": {
+ "chartType": "stackedBar",
+ "title": "Marital Status"
+ },
+ "patient_ombCategory": {
+ "chartType": "stackedBar",
+ "title": "OMB Category"
+ }
+ },
+ "filters": {
+ "tabs": [
+ {
+ "title": "File",
+ "fields": [
+ "project_id",
+ "file_category",
+ "data_type",
+ "data_format"
+ ]
+ },
+ {
+ "title": "Patient",
+ "fields": [
+ "project_id",
+ "patient_id",
+ "patient_gender",
+ "patient_disability_adjusted_life_years",
+ "patient_ombCategory",
+ "patient_ombCategory_detail",
+ "patient_us_core_birthsex",
+ "patient_quality_adjusted_life_years"
+ ]
+ }
+
+ ]
+ },
+ "table": {
+ "enabled": true,
+ "fields": [
+ "project_id",
+ "patient_id",
+ "patient_gender",
+ "patient_disability_adjusted_life_years",
+ "patient_ombCategory",
+ "patient_ombCategory_detail",
+ "patient_us_core_birthsex",
+ "patient_quality_adjusted_life_years",
+ "file_category",
+ "file_name",
+ "file_size",
+ "object_id"
+ ]
+ },
+ "dropdowns": {},
+ "guppyConfig": {
+ "dataType": "file",
+ "fieldMapping": [
+ { "field": "object_id", "name": "Download ID" },
+ { "field": "patient_gender", "name": "Gender" },
+ { "field": "patient_disability_adjusted_life_years", "name": "Disability Adjusted Life Years" },
+ { "field": "patient_quality_adjusted_life_years", "name": "Quality Adjusted Life Years" },
+ { "field": "patient_ombCategory", "name": "OMB Category" },
+ { "field": "patient_ombCategory_detail", "name": "OMB Category Detail" },
+ { "field": "patient_us_core_birthsex", "name": "Birth Sex" },
+ { "field": "file_category", "name": "Category" },
+ { "field": "file_name", "name": "Name" },
+ { "field": "data_type", "name": "Type" },
+ { "field": "data_format", "name": "Format" }
+ ],
+ "nodeCountTitle": "Files",
+ "manifestMapping": {
+ "resourceIndexType": "Patient",
+ "resourceIdField": "_patient_id",
+ "referenceIdFieldInResourceIndex": "object_id",
+ "referenceIdFieldInDataIndex": "object_id"
+ },
+ "accessibleFieldCheckList": ["project_id"],
+ "accessibleValidationField": "project_id",
+ "downloadAccessor": "object_id"
+ }
+ },
+
+
+ {
+ "tabTitle": "Observation",
+ "charts": {
+ "category": {
+ "chartType": "stackedBar",
+ "title": "Category"
+ },
+ "encounter_type": {
+ "chartType": "stackedBar",
+ "title": "Encounter Type"
+ },
+ "code_display": {
+ "chartType": "stackedBar",
+ "title": "Code"
+ }
+ },
+ "filters": {
+ "tabs": [
+ {
+ "title": "Observation",
+ "fields": [
+ "category",
+ "code_display"
+ ]
+ },
+ {
+ "title": "Encounter",
+ "fields": [
+ "patient_id",
+ "encounter_type",
+ "encounter_start"
+ ]
+ }
+ ]
+ },
+ "table": {
+ "enabled": true,
+ "fields": [
+ "patient_id",
+ "category",
+ "code_display",
+ "valueQuantity_value",
+ "valueQuantity_unit",
+ "valueCodeableConcept_coding_0_display",
+ "component_0_code_coding_0_display",
+ "component_0_valueQuantity_value",
+ "component_0_valueQuantity_unit",
+ "component_1_code_coding_0_display",
+ "component_1_valueQuantity_value",
+ "component_1_valueQuantity_unit"
+ ]
+ },
+ "dropdowns": {},
+ "guppyConfig": {
+ "dataType": "case",
+ "fieldMapping": [
+ { "field": "category", "name": "Category" },
+ { "field": "code_display", "name": "Code" },
+ { "field": "valueQuantity_value", "name": "Quantity Value" },
+ { "field": "valueQuantity_unit", "name": "Quantity Unit" },
+ { "field": "valueCodeableConcept_coding_0_display", "name": "Coding" },
+ { "field": "component_0_code_coding_0_display", "name": "Component" },
+ { "field": "component_0_valueQuantity_value", "name": "Component Value" },
+ { "field": "component_0_valueQuantity_unit", "name": "Component Unit" },
+ { "field": "component_1_code_coding_0_display", "name": "Component" },
+ { "field": "component_1_valueQuantity_value", "name": "Component Value" },
+ { "field": "component_1_valueQuantity_unit", "name": "Component Unit" }
+ ],
+ "nodeCountTitle": "Observations",
+ "manifestMapping": {
+ "resourceIndexType": "Patient",
+ "resourceIdField": "_patient_id",
+ "referenceIdFieldInResourceIndex": "object_id",
+ "referenceIdFieldInDataIndex": "object_id"
+ },
+ "accessibleFieldCheckList": ["project_id"],
+ "accessibleValidationField": "project_id",
+ "downloadAccessor": "object_id"
+ }
+ }
+
+ ],
+
+
+ "discoveryConfig": {
+ "features": {
+ "advSearchFilters": {
+ "enabled": true,
+ "field": "advSearchFilters",
+ "filters": [
+ {
+ "key": "Research Focus Area"
+ },
+ {
+ "key": "Administering IC(s)"
+ },
+ {
+ "key": "Simulated vs. Real Data"
+ },
+ {
+ "key": "Year Grant Awarded"
+ },
+ {
+ "key": "Study Description"
+ }
+ ]
+ },
+ "exportToWorkspace": {
+ "enabled": true,
+ "manifestFieldName": "__manifest",
+ "enableDownloadManifest": true,
+ "downloadManifestButtonText": "Download",
+ "documentationLinks": {
+ "gen3Client": "https://gen3.org/resources/user/gen3-client/",
+ "gen3Workspaces": "https://gen3.org/resources/user/analyze-data/"
+ }
+ },
+ "pageTitle": {
+ "enabled": false,
+ "text": "Study Discovery"
+ },
+ "search": {
+ "searchBar": {
+ "enabled": true
+ },
+ "tagSearchDropdown": {
+ "enabled": true,
+ "collapsibleButtonText": "Study Characteristics"
+ }
+ },
+ "authorization": {
+ "enabled": false
+ }
+ },
+ "aggregations": [
+ {
+ "name": "Studies",
+ "field": "frontend_uid",
+ "type": "count"
+ },
+ {
+ "name": "Total Subjects",
+ "field": "_research_subject_count",
+ "type": "sum"
+ }
+ ],
+ "tagSelector": {
+ "title": "Tags by category"
+ },
+ "studyColumns": [
+ {
+ "name": "Study Name",
+ "field": "name",
+ "errorIfNotAvailable": false,
+ "valueIfNotAvailable": "n/a",
+ "width": "157px"
+ },
+ {
+ "name": "Full Name",
+ "field": "full_name",
+ "contentType": "string",
+ "errorIfNotAvailable": false,
+ "valueIfNotAvailable": "n/a",
+ "width": "400px"
+ },
+ {
+ "name": "Number of Subjects",
+ "field": "_research_subject_count",
+ "errorIfNotAvailable": false,
+ "valueIfNotAvailable": "n/a",
+ "contentType": "number",
+ "width": "98px"
+ },
+ {
+ "name": "ID Number",
+ "field": "study_id",
+ "errorIfNotAvailable": false,
+ "valueIfNotAvailable": "n/a",
+ "width": "152px"
+ },
+ {
+ "name": "Data Commons",
+ "field": "commons",
+ "errorIfNotAvailable": false,
+ "valueIfNotAvailable": "n/a",
+ "hrefValueFromField": "commons_url"
+ }
+ ],
+ "studyPreviewField": {
+ "name": "Description",
+ "field": "study_description",
+ "contentType": "string",
+ "includeName": false,
+ "includeIfNotAvailable": true,
+ "valueIfNotAvailable": "No description has been provided for this study."
+ },
+ "studyPageFields": {
+ "showAllAvailableFields": false,
+ "header": {
+ "field": "name"
+ },
+ "fieldsToShow": [
+ {
+ "groupName": "Study Identifiers",
+ "groupWidth": "half",
+ "includeName": false,
+ "fields": [
+ {
+ "name": "Number of Subjects",
+ "field": "_subjects_count",
+ "contentType": "number"
+ },
+ {
+ "name": "Full Name",
+ "field": "full_name",
+ "contentType": "string"
+ },
+ {
+ "name": "Short Name",
+ "field": "short_name",
+ "contentType": "string",
+ "includeName": true,
+ "includeIfNotAvailable": true,
+ "valueIfNotAvailable": "N/A"
+ },
+ {
+ "name": "dbGaP Accession Number",
+ "field": "dbgap_accession",
+ "contentType": "string",
+ "includeName": true,
+ "includeIfNotAvailable": false
+ },
+ {
+ "name": "Project ID",
+ "field": "project_id",
+ "contentType": "string",
+ "includeIfNotAvailable": false
+ },
+ {
+ "name": "Data Commons",
+ "field": "commons",
+ "contentType": "string",
+ "includeIfNotAvailable": false
+ }
+ ]
+ },
+ {
+ "groupWidth": "full",
+ "fields": [
+ {
+ "name": "Description",
+ "field": "study_description",
+ "contentType": "paragraphs",
+ "includeName": true,
+ "includeIfNotAvailable": true,
+ "valueIfNotAvailable": "No description has been provided for this study."
+ }
+ ]
+ }
+ ]
+ },
+ "minimalFieldMapping": {
+ "tagsListFieldName": "tags",
+ "authzField": "authz",
+ "uid": "frontend_uid"
+ },
+ "tagCategories": [
+ {
+ "name": "Commons",
+ "displayName": "Data Commons",
+ "color": "rgba(129, 211, 248, 1)",
+ "display": true
+ },
+ {
+ "name": "Program",
+ "color": "rgba(129, 211, 248, 1)",
+ "display": false
+ },
+ {
+ "name": "Study Registration",
+ "color": "rgba(236, 128, 141, 1)",
+ "display": false
+ },
+ {
+ "name": "Data Type",
+ "color": "rgba(112, 182, 3, 1)",
+ "display": false
+ },
+ {
+ "name": "primary_site",
+ "color": "rgba(112, 182, 3, 1)",
+ "display": false
+ },
+ {
+ "name": "disease_type",
+ "color": "rgba(112, 182, 3, 1)",
+ "display": false
+ },
+ {
+ "name": "data_type",
+ "color": "rgba(112, 182, 3, 1)",
+ "display": false
+ },
+ {
+ "name": "clinical_study_type",
+ "color": "rgba(112, 182, 3, 1)",
+ "display": false
+ }
+ ]
+ },
+
+ "components": {
+ "appName": "The Open Access Data Commons Portal",
+ "index": {
+ "introduction": {
+ "heading": "Open Access Data Commons",
+ "text": "The Open Access Data Commons supports the management, analysis and sharing of data for the research community with the aim of accelerating discovery and development of diagnostics, treatment and prevention of diseases.",
+ "link": "/submission"
+ },
+ "buttons": [
+ {
+ "name": "Data Dictionary",
+ "icon": "data-field-define",
+ "body": "Browse the nodes and properties of the graph data model used in the Open Access Data Commons.",
+ "link": "/DD",
+ "label": "Explore Data Model"
+ },
+ {
+ "name": "Explore Data",
+ "icon": "data-explore",
+ "body": "Search and download subsets of data from the Open Access Data Commons using intuitive navigation tools.",
+ "link": "/explorer",
+ "label": "Explore data"
+ },
+ {
+ "name": "Query Data",
+ "icon": "data-access",
+ "body": "Search and download subsets of data from the Open Access Data Commons using GraphQL queries.",
+ "link": "/query",
+ "label": "Query data"
+ },
+ {
+ "name": "Discover Study Metadata",
+ "icon": "data-explore",
+ "body": "Discover study descriptions and other metadata stored on the Data Commons",
+ "link": "/discovery",
+ "label": "Discover Study Metadata"
+ },
+ {
+ "name": "Analyze Data",
+ "icon": "data-analyze",
+ "body": "Perform analysis on the Open Access Data Commons data using Jupyter Notebooks.",
+ "link": "/workspace",
+ "label": "Run analysis"
+ }
+ ]
+ },
+ "navigation": {
+ "items": [
+ {
+ "icon": "query",
+ "link": "/discovery",
+ "name": "Discovery",
+ "tooltip": "Use free-text search and tags to rapidly find relevant studies."
+ },
+ {
+ "icon": "dictionary",
+ "link": "/DD",
+ "color": "#a2a2a2",
+ "name": "Dictionary",
+ "tooltip": "The Data Dictionary serves to inform the data model and is updated as new data is ingested."
+ },
+ {
+ "icon": "exploration",
+ "link": "/explorer",
+ "color": "#a2a2a2",
+ "name": "Exploration",
+ "tooltip": "The Exploration Page enables discovery of the data at the subject level and features a cohort builder."
+ },
+ {
+ "icon": "query",
+ "link": "/query",
+ "color": "#a2a2a2",
+ "name": "Query",
+ "tooltip": "Query the databases using GraphiQL."
+ },
+ {
+ "name": "Notebook Browser",
+ "link": "/resource-browser",
+ "icon": "analysis",
+ "tooltip": "Learn how to use Jupyter Notebooks to explore and visualize data available by running a tutorial notebook, or use one of these examples as a launching point for your own custom analysis."
+ },
+ {
+ "icon": "workspace",
+ "link": "#hostname#workspace/",
+ "color": "#a2a2a2",
+ "name": "Workspace",
+ "tooltip": "Gen3 workspaces are secure data analysis environments in the cloud that can access data from one or more data resources."
+ },
+ {
+ "icon": "profile",
+ "link": "/identity",
+ "color": "#a2a2a2",
+ "name": "Profile",
+ "tooltip": "Create API keys for programmatic data access, and review your authorization privileges to datasets and services."
+ }
+ ]
+ },
+ "topBar": {
+ "items": [
+ {
+ "icon": "upload",
+ "link": "/submission",
+ "name": "Submit Data"
+ },
+ {
+ "link": "https://gen3.org/resources/user",
+ "name": "Documentation"
+ }
+ ]
+ },
+ "login": {
+ "title": "Open Access Data Commons",
+ "subTitle": "Explore, Analyze, and Share Data",
+ "text": "The Open Access Data Commons is a scalable cloud-based platform for computational discovery designed for the health community.",
+ "contact": "If you have any questions about access or the registration process, please contact ",
+ "email": "support@datacommons.io",
+ "image": "/../../../../custom/sponsors/gitops-sponsors/gene_bgy"
+ },
+ "footerLogos": [
+ {
+ "src": "/src/img/gen3.png",
+ "href": "https://ctds.uchicago.edu/gen3",
+ "alt": "Gen3 Data Commons"
+ },
+ {
+ "src": "/src/img/createdby.png",
+ "href": "https://ctds.uchicago.edu/",
+ "alt": "Center for Translational Data Science at the University of Chicago"
+ }
+ ]
+ },
+
+ "resourceBrowser": {
+ "title": "My Data Commons' Jupyter Notebooks",
+ "description": "Here's a collection of notebooks I'm proud of",
+ "public": true,
+ "resources": [
+ {
+ "title": "Custom Notebook",
+ "category": "Notebooks",
+ "description": "This is a custom notebook that generates a bunch of charts",
+ "link": "https://blah.net/notebooks/nb.html",
+ "imageUrl": "/src/img//resource-browser/notebook_gene.png"
+ },
+ {
+ "title": "Custom Notebook 2",
+ "link": "/notebooks/nb.html",
+ "imageUrl": "/src/img/gen3.png"
+ }
+ ]
+ },
+
+ "graphql": {
+ "boardCounts": [
+ {
+ "graphql": "_research_subject_count",
+ "name": "research_subject",
+ "plural": "research_subjects"
+ },
+ {
+ "graphql": "_specimen_count",
+ "name": "specimen",
+ "plural": "specimens"
+ },
+ {
+ "graphql": "_observation_count",
+ "name": "observation",
+ "plural": "observations"
+ }
+ ],
+ "chartCounts": [
+ {
+ "graphql": "_research_subject_count",
+ "name": "research_subjects"
+ }
+ ],
+ "projectDetails": "boardCounts"
+ }
+}
diff --git a/examples/Secrets/guppy_config.json b/examples/Secrets/guppy_config.json
new file mode 100644
index 00000000..34d4f8df
--- /dev/null
+++ b/examples/Secrets/guppy_config.json
@@ -0,0 +1 @@
+{ "indices": [ { "index": "etl", "type": "case" }, { "index": "file", "type": "file" } ], "config_index": ["etl_array-config", "file_array-config"], "auth_filter_field": "auth_resource_path" }
\ No newline at end of file
diff --git a/examples/Secrets/metadata/aggregate_config.json b/examples/Secrets/metadata/aggregate_config.json
new file mode 100644
index 00000000..e6ee200c
--- /dev/null
+++ b/examples/Secrets/metadata/aggregate_config.json
@@ -0,0 +1,57 @@
+{
+ "gen3_commons": {
+ "IBD Commons": {
+ "mds_url": "https://ibdgc.datacommons.io",
+ "commons_url" : "ibdgc.datacommons.io",
+ "study_data_field" : "my_metadata",
+ "guid_type" : "my_metadata",
+ "columns_to_fields": {
+ "_subjects_count" : "subjects_count",
+ "study_description" : "brief_summary",
+ "short_name": "dataset_title",
+ "full_name": "dataset_title"
+ }
+ },
+ "BioData Catalyst": {
+ "mds_url": "https://gen3.biodatacatalyst.nhlbi.nih.gov",
+ "commons_url" : "gen3.biodatacatalyst.nhlbi.nih.gov",
+ "columns_to_fields": {
+ "short_name": "name",
+ "_unique_id" : "study_id"
+ }
+ },
+ "MIDRC": {
+ "mds_url": "https://data.midrc.org",
+ "commons_url" : "data.midrc.org",
+ "study_data_field" : "discovery_metadata",
+ "columns_to_fields": {
+ "_subjects_count" : "cases_count",
+ "study_description" : "research_description",
+ "_unique_id": "study_id"
+ }
+ },
+ "NIAID ClinicalData": {
+ "mds_url": "https://accessclinicaldata.niaid.nih.gov",
+ "commons_url" : "accessclinicaldata.niaid.nih.gov",
+ "study_data_field" : "my_metadata",
+ "guid_type" : "my_metadata",
+ "columns_to_fields": {
+ "full_name": "title",
+ "study_id" : "nct_number",
+ "_unique_id": "nct_number",
+ "study_description" : "brief_summary"
+ }
+ },
+ "AnVIL": {
+ "mds_url": "https://internalstaging.theanvil.io",
+ "commons_url": "gen3.theanvil.io",
+ "columns_to_fields": {
+ "name": "name",
+ "full_name": "full_name",
+ "_subjects_count" : "_subjects_count",
+ "_unique_id" : "study_id",
+ "study_description" : "study_description"
+ }
+ }
+ }
+}
\ No newline at end of file
diff --git a/examples/Secrets/user.yaml b/examples/Secrets/user.yaml
new file mode 100644
index 00000000..7475c82f
--- /dev/null
+++ b/examples/Secrets/user.yaml
@@ -0,0 +1,251 @@
+authz:
+ # policies automatically given to anyone, even if they are not authenticated
+ anonymous_policies:
+ - open_data_reader
+ - aced_submitter
+
+ # policies automatically given to authenticated users (in addition to their other policies)
+ all_users_policies: []
+
+ groups:
+ # can CRUD programs and projects and upload data files
+ - name: data_submitters
+ policies:
+ - services.sheepdog-admin
+ - data_upload
+ - MyFirstProject_submitter
+ - aced_submitter
+ users:
+ - test
+
+ # can create/update/delete indexd records
+ - name: indexd_admins
+ policies:
+ - indexd_admin
+ users:
+ - test
+
+ resources:
+ - name: workspace
+ - name: data_file
+ - name: services
+ subresources:
+ - name: sheepdog
+ subresources:
+ - name: submission
+ subresources:
+ - name: program
+ - name: project
+ - name: open
+ - name: programs
+ subresources:
+ - name: MyFirstProgram
+ subresources:
+ - name: projects
+ subresources:
+ - name: MyFirstProject
+ - name: jnkns
+ subresources:
+ - name: projects
+ subresources:
+ - name: jenkins
+ - name: program1
+ subresources:
+ - name: projects
+ subresources:
+ - name: P1
+ - name: aced
+ subresources:
+ - name: projects
+ subresources:
+ - name: Alcoholism
+ - name: Alzheimers
+ - name: Breast_Cancer
+ - name: Colon_Cancer
+ - name: Diabetes
+ - name: Lung_Cancer
+ - name: Prostate_Cancer
+ policies:
+ - id: workspace
+ description: be able to use workspace
+ resource_paths:
+ - /workspace
+ role_ids:
+ - workspace_user
+ - id: data_upload
+ description: upload raw data files to S3
+ role_ids:
+ - file_uploader
+ resource_paths:
+ - /data_file
+ - id: services.sheepdog-admin
+ description: CRUD access to programs and projects
+ role_ids:
+ - sheepdog_admin
+ resource_paths:
+ - /services/sheepdog/submission/program
+ - /services/sheepdog/submission/project
+ - id: indexd_admin
+ description: full access to indexd API
+ role_ids:
+ - indexd_admin
+ resource_paths:
+ - /programs
+ - /data_file
+ - id: open_data_reader
+ role_ids:
+ - reader
+ - storage_reader
+ resource_paths:
+ - /open
+ - id: all_programs_reader
+ role_ids:
+ - reader
+ - storage_reader
+ resource_paths:
+ - /programs
+ - id: MyFirstProject_submitter
+ role_ids:
+ - reader
+ - creator
+ - updater
+ - deleter
+ - storage_reader
+ - storage_writer
+ resource_paths:
+ - /programs/MyFirstProgram/projects/MyFirstProject
+ - id: aced_submitter
+ role_ids:
+ - reader
+ - creator
+ - updater
+ - deleter
+ - storage_reader
+ - storage_writer
+ resource_paths:
+ - /programs/aced/projects/Alcoholism
+ - /programs/aced/projects/Alzheimers
+ - /programs/aced/projects/Breast_Cancer
+ - /programs/aced/projects/Colon_Cancer
+ - /programs/aced/projects/Diabetes
+ - /programs/aced/projects/Lung_Cancer
+ - /programs/aced/projects/Prostate_Cancer
+ - id: jnkns
+ role_ids:
+ - reader
+ - creator
+ - updater
+ - deleter
+ - storage_reader
+ - storage_writer
+ resource_paths:
+ - /programs/jnkns
+ - /programs/jnkns/projects/jenkins
+ - id: program1
+ role_ids:
+ - reader
+ - creator
+ - updater
+ - deleter
+ - storage_reader
+ - storage_writer
+ resource_paths:
+ - /programs/program1
+ - /programs/program1/projects/P1
+
+ roles:
+ - id: file_uploader
+ permissions:
+ - id: file_upload
+ action:
+ service: fence
+ method: file_upload
+ - id: workspace_user
+ permissions:
+ - id: workspace_access
+ action:
+ service: jupyterhub
+ method: access
+ - id: sheepdog_admin
+ description: CRUD access to programs and projects
+ permissions:
+ - id: sheepdog_admin_action
+ action:
+ service: sheepdog
+ method: '*'
+ - id: indexd_admin
+ description: full access to indexd API
+ permissions:
+ - id: indexd_admin
+ action:
+ service: indexd
+ method: '*'
+ - id: admin
+ permissions:
+ - id: admin
+ action:
+ service: '*'
+ method: '*'
+ - id: creator
+ permissions:
+ - id: creator
+ action:
+ service: '*'
+ method: create
+ - id: reader
+ permissions:
+ - id: reader
+ action:
+ service: '*'
+ method: read
+ - id: updater
+ permissions:
+ - id: updater
+ action:
+ service: '*'
+ method: update
+ - id: deleter
+ permissions:
+ - id: deleter
+ action:
+ service: '*'
+ method: delete
+ - id: storage_writer
+ permissions:
+ - id: storage_creator
+ action:
+ service: '*'
+ method: write-storage
+ - id: storage_reader
+ permissions:
+ - id: storage_reader
+ action:
+ service: '*'
+ method: read-storage
+
+clients:
+ wts:
+ policies:
+ - all_programs_reader
+ - open_data_reader
+
+users:
+ test:
+ tags:
+ name: User One
+# email: mustbe@differentemail.com
+ policies:
+ - workspace
+ - data_upload
+ - MyFirstProject_submitter
+ - aced_submitter
+ - jnkns
+ - program1
+
+ username2:
+ tags:
+ name: John Doe
+ email: johndoe@gmail.com
+
+cloud_providers: {}
+groups: {}
diff --git a/fhir/README.md b/fhir/README.md
new file mode 100644
index 00000000..e7486781
--- /dev/null
+++ b/fhir/README.md
@@ -0,0 +1,22 @@
+
+# create database in postgres
+
+```
+DROP DATABASE hapi ; CREATE DATABASE hapi owner fhir_user ;
+```
+
+# delete all data in all tables
+
+```
+\c hapi;
+DO $$ DECLARE
+ r RECORD;
+BEGIN
+ FOR r IN (SELECT tablename FROM pg_tables WHERE schemaname = current_schema()) LOOP
+ EXECUTE 'TRUNCATE TABLE ' || quote_ident(r.tablename) || ' CASCADE ';
+ END LOOP;
+END $$;
+
+```
+
+
diff --git a/fhir/data/application.yaml b/fhir/data/application.yaml
new file mode 100644
index 00000000..4f36af87
--- /dev/null
+++ b/fhir/data/application.yaml
@@ -0,0 +1,206 @@
+#Adds the option to go to eg. http://localhost:8080/actuator/health for seeing the running configuration
+#see https://docs.spring.io/spring-boot/docs/current/reference/html/actuator.html#actuator.endpoints
+management:
+ endpoints:
+ web:
+ exposure:
+ include: "health,prometheus"
+spring:
+ main:
+ allow-circular-references: true
+ #allow-bean-definition-overriding: true
+ flyway:
+ enabled: false
+ check-location: false
+ baselineOnMigrate: true
+ datasource:
+ url: 'jdbc:postgresql://postgres:5432/hapi'
+ username: 'fhir_user'
+ password: 'fhir_pass'
+ driverClassName: org.postgresql.Driver
+ max-active: 20
+ # 6dB&ifdH3dRH
+# url: 'jdbc:h2:file:./target/database/h2'
+# #url: jdbc:h2:mem:test_mem
+# username: sa
+# password: null
+# driverClassName: org.h2.Driver
+# max-active: 15
+
+ # database connection pool size
+ hikari:
+ maximum-pool-size: 25
+ jpa:
+ properties:
+ hibernate.format_sql: false
+ hibernate.show_sql: false
+ #Hibernate dialect is automatically detected except Postgres and H2.
+ #If using H2, then supply the value of ca.uhn.fhir.jpa.model.dialect.HapiFhirH2Dialect
+ #If using postgres, then supply the value of ca.uhn.fhir.jpa.model.dialect.HapiFhirPostgres94Dialect
+
+ hibernate.dialect: ca.uhn.fhir.jpa.model.dialect.HapiFhirPostgres94Dialect
+ # hibernate.hbm2ddl.auto: update
+ # hibernate.jdbc.batch_size: 20
+ # hibernate.cache.use_query_cache: false
+ # hibernate.cache.use_second_level_cache: false
+ # hibernate.cache.use_structured_entries: false
+ # hibernate.cache.use_minimal_puts: false
+ ### These settings will enable fulltext search with lucene or elastic
+ hibernate.search.enabled: true
+ ### lucene parameters
+# hibernate.search.backend.type: lucene
+# hibernate.search.backend.analysis.configurer: ca.uhn.fhir.jpa.search.HapiHSearchAnalysisConfigurers$HapiLuceneAnalysisConfigurer
+# hibernate.search.backend.directory.type: local-filesystem
+# hibernate.search.backend.directory.root: target/lucenefiles
+# hibernate.search.backend.lucene_version: lucene_current
+ ### elastic parameters ===> see also elasticsearch section below <===
+# hibernate.search.backend.type: elasticsearch
+# hibernate.search.backend.analysis.configurer: ca.uhn.fhir.jpa.search.HapiHSearchAnalysisConfigurers$HapiElasticAnalysisConfigurer
+hapi:
+ fhir:
+ ### This enables the swagger-ui at /fhir/swagger-ui/index.html as well as the /fhir/api-docs (see https://hapifhir.io/hapi-fhir/docs/server_plain/openapi.html)
+ openapi_enabled: true
+ ### This is the FHIR version. Choose between, DSTU2, DSTU3, R4 or R5
+ fhir_version: R4
+ ### enable to use the ApacheProxyAddressStrategy which uses X-Forwarded-* headers
+ ### to determine the FHIR server address
+ # use_apache_address_strategy: false
+ ### forces the use of the https:// protocol for the returned server address.
+ ### alternatively, it may be set using the X-Forwarded-Proto header.
+ # use_apache_address_strategy_https: false
+ ### enable to set the Server URL
+ # server_address: http://hapi.fhir.org/baseR4
+ # defer_indexing_for_codesystems_of_size: 101
+ # install_transitive_ig_dependencies: true
+ # implementationguides:
+ ### example from registry (packages.fhir.org)
+ # swiss:
+ # name: swiss.mednet.fhir
+ # version: 0.8.0
+ # example not from registry
+ # ips_1_0_0:
+ # url: https://build.fhir.org/ig/HL7/fhir-ips/package.tgz
+ # name: hl7.fhir.uv.ips
+ # version: 1.0.0
+ # supported_resource_types:
+ # - Patient
+ # - Observation
+ ##################################################
+ # Allowed Bundle Types for persistence (defaults are: COLLECTION,DOCUMENT,MESSAGE)
+ ##################################################
+ # allowed_bundle_types: COLLECTION,DOCUMENT,MESSAGE,TRANSACTION,TRANSACTIONRESPONSE,BATCH,BATCHRESPONSE,HISTORY,SEARCHSET
+ # allow_cascading_deletes: true
+ # allow_contains_searches: true
+ # allow_external_references: true
+ # allow_multiple_delete: true
+ # allow_override_default_search_params: true
+ # auto_create_placeholder_reference_targets: false
+ # cql_enabled: true
+ # default_encoding: JSON
+ # default_pretty_print: true
+ # default_page_size: 20
+ # delete_expunge_enabled: true
+ # enable_repository_validating_interceptor: true
+ # enable_index_missing_fields: false
+ # enable_index_of_type: true
+ # enable_index_contained_resource: false
+ ### !!Extended Lucene/Elasticsearch Indexing is still a experimental feature, expect some features (e.g. _total=accurate) to not work as expected!!
+ ### more information here: https://hapifhir.io/hapi-fhir/docs/server_jpa/elastic.html
+ advanced_lucene_indexing: false
+ bulk_export_enabled: true
+ bulk_import_enabled: true
+ # enforce_referential_integrity_on_delete: false
+ # This is an experimental feature, and does not fully support _total and other FHIR features.
+ # enforce_referential_integrity_on_delete: false
+ # enforce_referential_integrity_on_write: false
+ # etag_support_enabled: true
+ # expunge_enabled: true
+ client_id_strategy: ANY
+ # fhirpath_interceptor_enabled: false
+ # filter_search_enabled: true
+ # graphql_enabled: true
+ # narrative_enabled: true
+ # mdm_enabled: true
+ # local_base_urls:
+ # - https://hapi.fhir.org/baseR4
+ mdm_enabled: false
+ # partitioning:
+ # allow_references_across_partitions: false
+ # partitioning_include_in_search_hashes: false
+ cors:
+ allow_Credentials: true
+ # These are allowed_origin patterns, see: https://docs.spring.io/spring-framework/docs/current/javadoc-api/org/springframework/web/cors/CorsConfiguration.html#setAllowedOriginPatterns-java.util.List-
+ allowed_origin:
+ - '*'
+
+ # Search coordinator thread pool sizes
+ search-coord-core-pool-size: 20
+ search-coord-max-pool-size: 100
+ search-coord-queue-capacity: 200
+
+ # Threadpool size for BATCH'ed GETs in a bundle.
+ # bundle_batch_pool_size: 10
+ # bundle_batch_pool_max_size: 50
+
+ # logger:
+ # error_format: 'ERROR - ${requestVerb} ${requestUrl}'
+ # format: >-
+ # Path[${servletPath}] Source[${requestHeader.x-forwarded-for}]
+ # Operation[${operationType} ${operationName} ${idOrResourceName}]
+ # UA[${requestHeader.user-agent}] Params[${requestParameters}]
+ # ResponseEncoding[${responseEncodingNoDefault}]
+ # log_exceptions: true
+ # name: fhirtest.access
+ # max_binary_size: 104857600
+ # max_page_size: 200
+ # retain_cached_searches_mins: 60
+ # reuse_cached_search_results_millis: 60000
+ tester:
+ home:
+ name: Local Tester
+ server_address: 'http://localhost:8080/fhir'
+ refuse_to_fetch_third_party_urls: false
+ fhir_version: R4
+ global:
+ name: Global Tester
+ server_address: "http://hapi.fhir.org/baseR4"
+ refuse_to_fetch_third_party_urls: false
+ fhir_version: R4
+ # validation:
+ # requests_enabled: true
+ # responses_enabled: true
+ # binary_storage_enabled: true
+ inline_resource_storage_below_size: 10000
+# bulk_export_enabled: true
+# subscription:
+# resthook_enabled: true
+# websocket_enabled: false
+# email:
+# from: some@test.com
+# host: google.com
+# port:
+# username:
+# password:
+# auth:
+# startTlsEnable:
+# startTlsRequired:
+# quitWait:
+# lastn_enabled: true
+# store_resource_in_lucene_index_enabled: true
+### This is configuration for normalized quantity serach level default is 0
+### 0: NORMALIZED_QUANTITY_SEARCH_NOT_SUPPORTED - default
+### 1: NORMALIZED_QUANTITY_STORAGE_SUPPORTED
+### 2: NORMALIZED_QUANTITY_SEARCH_SUPPORTED
+# normalized_quantity_search_level: 2
+#elasticsearch:
+# debug:
+# pretty_print_json_log: false
+# refresh_after_write: false
+# enabled: false
+# password: SomePassword
+# required_index_status: YELLOW
+# rest_url: 'localhost:9200'
+# protocol: 'http'
+# schema_management_strategy: CREATE
+# username: SomeUsername
+#
diff --git a/guppy_setup.sh b/guppy_setup.sh
index 559668d2..b16c8b2b 100644
--- a/guppy_setup.sh
+++ b/guppy_setup.sh
@@ -1,16 +1,11 @@
#!/bin/bash
# Script to create and re-create es indices and setup guppy
-sleep 2
-docker exec esproxy-service curl -X DELETE http://localhost:9200/etl_0
-sleep 2
-docker exec esproxy-service curl -X DELETE http://localhost:9200/file_0
-sleep 2
-docker exec esproxy-service curl -X DELETE http://localhost:9200/file-array-config_0
-sleep 2
-docker exec esproxy-service curl -X DELETE http://localhost:9200/etl-array-config_0
-sleep 2
-docker exec tube-service bash -c "python run_config.py && python run_etl.py"
+docker exec esproxy-service curl -X DELETE http://localhost:9200/gen3.aced.*
+docker compose stop guppy-service
+docker compose rm -f guppy-service
-docker container stop guppy-service
-docker container start guppy-service
+# start two processes one for observation and one for file, run them in parallel at a higher 'niceness' level
+echo -e observation"\n"file | nice -n 10 xargs -L 1 -P 2 nice -n 10 ./etl/tube_lite --credentials_path $1 --elastic http://localhost:9200 --batch_size 20 --entity
+
+docker compose up -d guppy-service
diff --git a/minio-webhook/Dockerfile b/minio-webhook/Dockerfile
new file mode 100644
index 00000000..b8bc59fd
--- /dev/null
+++ b/minio-webhook/Dockerfile
@@ -0,0 +1,16 @@
+# set base image (host OS)
+FROM python:3.10
+
+# set the working directory in the container
+WORKDIR /minio-webhook
+
+# copy the dependencies file to the working directory
+COPY requirements.txt .
+
+# install dependencies
+RUN pip install -r requirements.txt
+
+
+RUN wget https://dl.min.io/client/mc/release/linux-amd64/mc
+RUN chmod +x mc
+RUN mv mc /usr/local/bin
diff --git a/minio-webhook/config.py b/minio-webhook/config.py
new file mode 100644
index 00000000..6e1c8b0c
--- /dev/null
+++ b/minio-webhook/config.py
@@ -0,0 +1,29 @@
+from pydantic import BaseModel
+
+class LogConfig(BaseModel):
+ """Logging configuration to be set for the server"""
+
+ LOGGER_NAME: str = "minio-webhook"
+ LOG_FORMAT: str = "%(levelprefix)s | %(asctime)s | %(message)s"
+ LOG_LEVEL: str = "DEBUG"
+
+ # Logging config
+ version = 1
+ disable_existing_loggers = False
+ formatters = {
+ "default": {
+ "()": "uvicorn.logging.DefaultFormatter",
+ "fmt": LOG_FORMAT,
+ "datefmt": "%Y-%m-%d %H:%M:%S",
+ },
+ }
+ handlers = {
+ "default": {
+ "formatter": "default",
+ "class": "logging.StreamHandler",
+ "stream": "ext://sys.stderr",
+ },
+ }
+ loggers = {
+ "minio-webhook": {"handlers": ["default"], "level": LOG_LEVEL},
+ }
diff --git a/minio-webhook/main.py b/minio-webhook/main.py
new file mode 100644
index 00000000..e5713aa6
--- /dev/null
+++ b/minio-webhook/main.py
@@ -0,0 +1,43 @@
+from logging.config import dictConfig
+import logging
+from config import LogConfig
+
+from fastapi import FastAPI
+
+# https://docs.aws.amazon.com/AmazonS3/latest/userguide/notification-content-structure.html
+# https://pydantic-docs.helpmanual.io/datamodel_code_generator/
+from model import Model
+
+app = FastAPI()
+
+
+@app.get("/")
+async def root():
+ return {"message": "Health OK."}
+
+
+@app.head("/file-uploaded")
+async def uploaded_head():
+ """Receive HEAD from minio server when minio starts."""
+ pass
+
+
+@app.post("/file-uploaded")
+async def uploaded(body: Model):
+ """Receive POST from minio server when file uploaded."""
+ if body.EventName != 's3:ObjectCreated:Put':
+ logger.warning(f"Ignoring unexpected event type {body}")
+ return
+ # logger.info(f"Processing event type {body.EventName} {body}")
+ for record in body.Records:
+ logger.info(
+ f"Update gen3 sheepdog and indexd with: {record.s3.bucket.name} {record.s3.object.key} {record.s3.object.size} {record.s3.object.eTag} {record.s3.object.userMetadata}")
+ return
+
+# https://aced-training.compbio.ohsu.edu/index/index?acl=null&uploader=test&start=00000000-0000-0000-0000-000000000000&limit=1024
+
+
+# setup logging
+dictConfig(LogConfig().dict())
+logger = logging.getLogger("minio-webhook")
+
diff --git a/minio-webhook/model.py b/minio-webhook/model.py
new file mode 100644
index 00000000..16d15537
--- /dev/null
+++ b/minio-webhook/model.py
@@ -0,0 +1,87 @@
+# generated by datamodel-codegen:
+# filename: event.json
+# timestamp: 2022-09-10T13:03:37+00:00
+
+from __future__ import annotations
+
+from typing import List
+
+from pydantic import BaseModel, Field, Extra
+from typing import Optional
+
+
+class UserIdentity(BaseModel):
+ principalId: str
+
+
+class RequestParameters(BaseModel):
+ principalId: str
+ region: str
+ sourceIPAddress: str
+
+
+class ResponseElements(BaseModel):
+ content_length: str = Field(..., alias='content-length')
+ x_amz_request_id: str = Field(..., alias='x-amz-request-id')
+ x_minio_deployment_id: str = Field(..., alias='x-minio-deployment-id')
+ x_minio_origin_endpoint: str = Field(..., alias='x-minio-origin-endpoint')
+
+
+class OwnerIdentity(BaseModel):
+ principalId: str
+
+
+class Bucket(BaseModel):
+ name: str
+ ownerIdentity: OwnerIdentity
+ arn: str
+
+
+class UserMetadata(BaseModel, extra=Extra.allow):
+ content_type: str = Field(..., alias='content-type')
+ # our extensions, set by etl/file::upload
+ datanode_type: Optional[str] = Field(..., alias='X-Amz-Meta-Datanode_type')
+ datanode_submitter_id: Optional[str] = Field(..., alias='X-Amz-Meta-Datanode_submitter_id')
+ datanode_object_id: Optional[str] = Field(..., alias='X-Amz-Meta-Datanode_object_id')
+ md5: Optional[str] = Field(..., alias='X-Amz-Meta-Md5')
+
+
+class Object(BaseModel):
+ key: str
+ size: int
+ eTag: str
+ contentType: str
+ userMetadata: UserMetadata
+ sequencer: str
+
+
+class S3(BaseModel):
+ s3SchemaVersion: str
+ configurationId: str
+ bucket: Bucket
+ object: Object
+
+
+class Source(BaseModel):
+ host: str
+ port: str
+ userAgent: str
+
+
+class Record(BaseModel):
+ eventVersion: str
+ eventSource: str
+ awsRegion: str
+ eventTime: str
+ eventName: str
+ userIdentity: UserIdentity
+ requestParameters: RequestParameters
+ responseElements: ResponseElements
+ s3: S3
+ source: Source
+
+
+class Model(BaseModel):
+ EventName: str
+ Key: str
+ Records: List[Record]
diff --git a/minio-webhook/requirements.txt b/minio-webhook/requirements.txt
new file mode 100644
index 00000000..611e7d21
--- /dev/null
+++ b/minio-webhook/requirements.txt
@@ -0,0 +1,7 @@
+pyjwt
+Click
+gen3
+jsonpath-ng
+certifi
+
+fastapi[all]
\ No newline at end of file
diff --git a/minio.conf b/minio.conf
new file mode 100644
index 00000000..9b2255e4
--- /dev/null
+++ b/minio.conf
@@ -0,0 +1,441 @@
+#
+# MINIO configuration
+#
+
+ # For load balancing, cluster support: coordinate with docker-compose.override.yml
+ upstream minio-default {
+ server minio-default:9000;
+ # add more servers for clustering
+ }
+ upstream minio-default-console {
+ ip_hash;
+ server minio-default:9001;
+ # add more servers for clustering
+ }
+ upstream minio-ohsu {
+ server minio-ohsu:9000;
+ }
+ upstream minio-ohsu-console {
+ ip_hash;
+ server minio-ohsu:9001;
+ }
+ upstream minio-ucl {
+ server minio-ucl:9000;
+ }
+ upstream minio-ucl-console {
+ ip_hash;
+ server minio-ucl:9001;
+ }
+ upstream minio-manchester {
+ server minio-manchester:9000;
+ }
+ upstream minio-manchester-console {
+ ip_hash;
+ server minio-manchester:9001;
+ }
+ upstream minio-stanford {
+ server minio-stanford:9000;
+ }
+ upstream minio-stanford-console {
+ ip_hash;
+ server minio-stanford:9001;
+ }
+
+ #
+ # MINIO S3 storage host(s)
+ #
+ server {
+ server_name minio-default.compbio.ohsu.edu;
+
+ listen 80;
+ resolver 127.0.0.11;
+ listen 443 ssl;
+
+ ssl_certificate /etc/nginx/ssl/nginx.crt;
+ ssl_certificate_key /etc/nginx/ssl/nginx.key;
+
+
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+
+ proxy_connect_timeout 300;
+ # Default is HTTP/1, keepalive is only enabled in HTTP/1.1
+ proxy_http_version 1.1;
+ proxy_set_header Connection "";
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-default;
+ }
+ }
+ server {
+ server_name minio-ohsu.compbio.ohsu.edu;
+
+ listen 80;
+ resolver 127.0.0.11;
+ listen 443 ssl;
+
+ ssl_certificate /etc/nginx/ssl/nginx.crt;
+ ssl_certificate_key /etc/nginx/ssl/nginx.key;
+
+
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+
+ proxy_connect_timeout 300;
+ # Default is HTTP/1, keepalive is only enabled in HTTP/1.1
+ proxy_http_version 1.1;
+ proxy_set_header Connection "";
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-ohsu;
+ }
+ }
+ server {
+ server_name minio-ucl.compbio.ohsu.edu;
+
+ listen 80;
+ resolver 127.0.0.11;
+ listen 443 ssl;
+
+ ssl_certificate /etc/nginx/ssl/nginx.crt;
+ ssl_certificate_key /etc/nginx/ssl/nginx.key;
+
+
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+
+ proxy_connect_timeout 300;
+ # Default is HTTP/1, keepalive is only enabled in HTTP/1.1
+ proxy_http_version 1.1;
+ proxy_set_header Connection "";
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-ucl;
+ }
+ }
+ server {
+ server_name minio-manchester.compbio.ohsu.edu;
+
+ listen 80;
+ resolver 127.0.0.11;
+ listen 443 ssl;
+
+ ssl_certificate /etc/nginx/ssl/nginx.crt;
+ ssl_certificate_key /etc/nginx/ssl/nginx.key;
+
+
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+
+ proxy_connect_timeout 300;
+ # Default is HTTP/1, keepalive is only enabled in HTTP/1.1
+ proxy_http_version 1.1;
+ proxy_set_header Connection "";
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-manchester;
+ }
+ }
+ server {
+ server_name minio-stanford.compbio.ohsu.edu;
+
+ listen 80;
+ resolver 127.0.0.11;
+ listen 443 ssl;
+
+ ssl_certificate /etc/nginx/ssl/nginx.crt;
+ ssl_certificate_key /etc/nginx/ssl/nginx.key;
+
+
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+
+ proxy_connect_timeout 300;
+ # Default is HTTP/1, keepalive is only enabled in HTTP/1.1
+ proxy_http_version 1.1;
+ proxy_set_header Connection "";
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-stanford;
+ }
+ }
+
+ #
+ # MINIO S3 console host(s)
+ #
+ server {
+ server_name minio-default-console.compbio.ohsu.edu;
+
+ listen 80;
+ resolver 127.0.0.11;
+
+ listen 443 ssl;
+
+ ssl_certificate /etc/nginx/ssl/nginx.crt;
+ ssl_certificate_key /etc/nginx/ssl/nginx.key;
+
+
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+ proxy_set_header X-NginX-Proxy true;
+
+ # This is necessary to pass the correct IP to be hashed
+ real_ip_header X-Real-IP;
+
+ proxy_connect_timeout 300;
+
+ # To support websocket
+ proxy_http_version 1.1;
+ proxy_set_header Upgrade $http_upgrade;
+ proxy_set_header Connection "upgrade";
+
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-default-console;
+ }
+ }
+ server {
+ server_name minio-ohsu-console.compbio.ohsu.edu;
+
+ listen 80;
+ resolver 127.0.0.11;
+
+ listen 443 ssl;
+
+ ssl_certificate /etc/nginx/ssl/nginx.crt;
+ ssl_certificate_key /etc/nginx/ssl/nginx.key;
+
+
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+ proxy_set_header X-NginX-Proxy true;
+
+ # This is necessary to pass the correct IP to be hashed
+ real_ip_header X-Real-IP;
+
+ proxy_connect_timeout 300;
+
+ # To support websocket
+ proxy_http_version 1.1;
+ proxy_set_header Upgrade $http_upgrade;
+ proxy_set_header Connection "upgrade";
+
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-ohsu-console;
+ }
+ }
+ server {
+ server_name minio-ucl-console.compbio.ohsu.edu;
+
+ listen 80;
+ resolver 127.0.0.11;
+
+ listen 443 ssl;
+
+ ssl_certificate /etc/nginx/ssl/nginx.crt;
+ ssl_certificate_key /etc/nginx/ssl/nginx.key;
+
+
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+ proxy_set_header X-NginX-Proxy true;
+
+ # This is necessary to pass the correct IP to be hashed
+ real_ip_header X-Real-IP;
+
+ proxy_connect_timeout 300;
+
+ # To support websocket
+ proxy_http_version 1.1;
+ proxy_set_header Upgrade $http_upgrade;
+ proxy_set_header Connection "upgrade";
+
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-ucl-console;
+ }
+ }
+ server {
+ server_name minio-manchester-console.compbio.ohsu.edu;
+
+ listen 80;
+ resolver 127.0.0.11;
+
+ listen 443 ssl;
+
+ ssl_certificate /etc/nginx/ssl/nginx.crt;
+ ssl_certificate_key /etc/nginx/ssl/nginx.key;
+
+
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+ proxy_set_header X-NginX-Proxy true;
+
+ # This is necessary to pass the correct IP to be hashed
+ real_ip_header X-Real-IP;
+
+ proxy_connect_timeout 300;
+
+ # To support websocket
+ proxy_http_version 1.1;
+ proxy_set_header Upgrade $http_upgrade;
+ proxy_set_header Connection "upgrade";
+
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-manchester-console;
+ }
+ }
+ server {
+ server_name minio-stanford-console.compbio.ohsu.edu;
+
+ listen 80;
+ resolver 127.0.0.11;
+
+ listen 443 ssl;
+
+ ssl_certificate /etc/nginx/ssl/nginx.crt;
+ ssl_certificate_key /etc/nginx/ssl/nginx.key;
+
+
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+ proxy_set_header X-NginX-Proxy true;
+
+ # This is necessary to pass the correct IP to be hashed
+ real_ip_header X-Real-IP;
+
+ proxy_connect_timeout 300;
+
+ # To support websocket
+ proxy_http_version 1.1;
+ proxy_set_header Upgrade $http_upgrade;
+ proxy_set_header Connection "upgrade";
+
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-stanford-console;
+ }
+ }
+
diff --git a/minio.conf.staging b/minio.conf.staging
new file mode 100644
index 00000000..a423e82c
--- /dev/null
+++ b/minio.conf.staging
@@ -0,0 +1,457 @@
+#
+# MINIO configuration
+#
+
+ # For load balancing, cluster support: coordinate with docker-compose.override.yml
+ upstream minio-default {
+ server minio-default:9000;
+ # add more servers for clustering
+ }
+ upstream minio-default-console {
+ ip_hash;
+ server minio-default:9001;
+ # add more servers for clustering
+ }
+ upstream minio-ohsu {
+ server minio-ohsu:9000;
+ }
+ upstream minio-ohsu-console {
+ ip_hash;
+ server minio-ohsu:9001;
+ }
+ upstream minio-ucl {
+ server minio-ucl:9000;
+ }
+ upstream minio-ucl-console {
+ ip_hash;
+ server minio-ucl:9001;
+ }
+ upstream minio-manchester {
+ server minio-manchester:9000;
+ }
+ upstream minio-manchester-console {
+ ip_hash;
+ server minio-manchester:9001;
+ }
+ upstream minio-stanford {
+ server minio-stanford:9000;
+ }
+ upstream minio-stanford-console {
+ ip_hash;
+ server minio-stanford:9001;
+ }
+
+ #
+ # MINIO S3 storage host(s)
+ #
+ server {
+ server_name minio-default-staging.aced-idp.org;
+ listen 80;
+ resolver 127.0.0.11;
+ listen 443 ssl;
+ ssl_certificate /etc/letsencrypt/live/staging.aced-idp.org/fullchain.pem;
+ ssl_certificate_key /etc/letsencrypt/live/staging.aced-idp.org/privkey.pem;
+
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+
+ proxy_connect_timeout 300;
+ # Default is HTTP/1, keepalive is only enabled in HTTP/1.1
+ proxy_http_version 1.1;
+ proxy_set_header Connection "";
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-default;
+ }
+ location /.well-known/acme-challenge/ {
+ root /var/www/certbot;
+ }
+ }
+ server {
+ server_name minio-ohsu-staging.aced-idp.org;
+
+ listen 80;
+ resolver 127.0.0.11;
+ listen 443 ssl;
+
+ ssl_certificate /etc/letsencrypt/live/staging.aced-idp.org/fullchain.pem;
+ ssl_certificate_key /etc/letsencrypt/live/staging.aced-idp.org/privkey.pem;
+
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+
+ proxy_connect_timeout 300;
+ # Default is HTTP/1, keepalive is only enabled in HTTP/1.1
+ proxy_http_version 1.1;
+ proxy_set_header Connection "";
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-ohsu;
+ }
+ location /.well-known/acme-challenge/ {
+ root /var/www/certbot;
+ }
+ }
+ server {
+ server_name minio-ucl-staging.aced-idp.org;
+ listen 80;
+ resolver 127.0.0.11;
+ listen 443 ssl;
+
+ ssl_certificate /etc/letsencrypt/live/staging.aced-idp.org/fullchain.pem;
+ ssl_certificate_key /etc/letsencrypt/live/staging.aced-idp.org/privkey.pem;
+
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+
+ proxy_connect_timeout 300;
+ # Default is HTTP/1, keepalive is only enabled in HTTP/1.1
+ proxy_http_version 1.1;
+ proxy_set_header Connection "";
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-ucl;
+ }
+ location /.well-known/acme-challenge/ {
+ root /var/www/certbot;
+ }
+ }
+ server {
+ server_name minio-manchester-staging.aced-idp.org;
+
+ listen 80;
+ resolver 127.0.0.11;
+ listen 443 ssl;
+
+ ssl_certificate /etc/letsencrypt/live/staging.aced-idp.org/fullchain.pem;
+ ssl_certificate_key /etc/letsencrypt/live/staging.aced-idp.org/privkey.pem;
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+
+ proxy_connect_timeout 300;
+ # Default is HTTP/1, keepalive is only enabled in HTTP/1.1
+ proxy_http_version 1.1;
+ proxy_set_header Connection "";
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-manchester;
+ }
+ location /.well-known/acme-challenge/ {
+ root /var/www/certbot;
+ }
+ }
+ server {
+ server_name minio-stanford-staging.aced-idp.org;
+
+ listen 80;
+ resolver 127.0.0.11;
+ listen 443 ssl;
+
+ ssl_certificate /etc/letsencrypt/live/staging.aced-idp.org/fullchain.pem;
+ ssl_certificate_key /etc/letsencrypt/live/staging.aced-idp.org/privkey.pem;
+
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+
+ proxy_connect_timeout 300;
+ # Default is HTTP/1, keepalive is only enabled in HTTP/1.1
+ proxy_http_version 1.1;
+ proxy_set_header Connection "";
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-stanford;
+ }
+ location /.well-known/acme-challenge/ {
+ root /var/www/certbot;
+ }
+ }
+
+ #
+ # MINIO S3 console host(s)
+ #
+ server {
+ server_name minio-default-console-staging.aced-idp.org;
+
+ listen 80;
+ resolver 127.0.0.11;
+
+ listen 443 ssl;
+
+ ssl_certificate /etc/letsencrypt/live/staging.aced-idp.org/fullchain.pem;
+ ssl_certificate_key /etc/letsencrypt/live/staging.aced-idp.org/privkey.pem;
+
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+ proxy_set_header X-NginX-Proxy true;
+
+ # This is necessary to pass the correct IP to be hashed
+ real_ip_header X-Real-IP;
+
+ proxy_connect_timeout 300;
+
+ # To support websocket
+ proxy_http_version 1.1;
+ proxy_set_header Upgrade $http_upgrade;
+ proxy_set_header Connection "upgrade";
+
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-default-console;
+ }
+ location /.well-known/acme-challenge/ {
+ root /var/www/certbot;
+ }
+ }
+ server {
+ server_name minio-ohsu-console-staging.aced-idp.org;
+
+ listen 80;
+ resolver 127.0.0.11;
+
+ listen 443 ssl;
+
+ ssl_certificate /etc/letsencrypt/live/staging.aced-idp.org/fullchain.pem;
+ ssl_certificate_key /etc/letsencrypt/live/staging.aced-idp.org/privkey.pem;
+
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+ proxy_set_header X-NginX-Proxy true;
+
+ # This is necessary to pass the correct IP to be hashed
+ real_ip_header X-Real-IP;
+
+ proxy_connect_timeout 300;
+
+ # To support websocket
+ proxy_http_version 1.1;
+ proxy_set_header Upgrade $http_upgrade;
+ proxy_set_header Connection "upgrade";
+
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-ohsu-console;
+ }
+ location /.well-known/acme-challenge/ {
+ root /var/www/certbot;
+ }
+ }
+ server {
+ server_name minio-ucl-console-staging.aced-idp.org;
+
+ listen 80;
+ resolver 127.0.0.11;
+
+ listen 443 ssl;
+
+ ssl_certificate /etc/letsencrypt/live/staging.aced-idp.org/fullchain.pem;
+ ssl_certificate_key /etc/letsencrypt/live/staging.aced-idp.org/privkey.pem;
+
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+ proxy_set_header X-NginX-Proxy true;
+
+ # This is necessary to pass the correct IP to be hashed
+ real_ip_header X-Real-IP;
+
+ proxy_connect_timeout 300;
+
+ # To support websocket
+ proxy_http_version 1.1;
+ proxy_set_header Upgrade $http_upgrade;
+ proxy_set_header Connection "upgrade";
+
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-ucl-console;
+ }
+ location /.well-known/acme-challenge/ {
+ root /var/www/certbot;
+ }
+ }
+ server {
+ server_name minio-manchester-console-staging.aced-idp.org;
+
+ listen 80;
+ resolver 127.0.0.11;
+
+ listen 443 ssl;
+
+ ssl_certificate /etc/letsencrypt/live/staging.aced-idp.org/fullchain.pem;
+ ssl_certificate_key /etc/letsencrypt/live/staging.aced-idp.org/privkey.pem;
+
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+ proxy_set_header X-NginX-Proxy true;
+
+ # This is necessary to pass the correct IP to be hashed
+ real_ip_header X-Real-IP;
+
+ proxy_connect_timeout 300;
+
+ # To support websocket
+ proxy_http_version 1.1;
+ proxy_set_header Upgrade $http_upgrade;
+ proxy_set_header Connection "upgrade";
+
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-manchester-console;
+ }
+ location /.well-known/acme-challenge/ {
+ root /var/www/certbot;
+ }
+ }
+ server {
+ server_name minio-stanford-console-staging.aced-idp.org;
+
+ listen 80;
+ resolver 127.0.0.11;
+
+ listen 443 ssl;
+
+ ssl_certificate /etc/letsencrypt/live/staging.aced-idp.org/fullchain.pem;
+ ssl_certificate_key /etc/letsencrypt/live/staging.aced-idp.org/privkey.pem;
+
+ # To allow special characters in headers
+ ignore_invalid_headers off;
+ # Allow any size file to be uploaded.
+ # Set to a value such as 1000m; to restrict file size to a specific value
+ client_max_body_size 0;
+ # To disable buffering
+ proxy_buffering off;
+ proxy_request_buffering off;
+
+ location / {
+ proxy_set_header Host $http_host;
+ proxy_set_header X-Real-IP $remote_addr;
+ proxy_set_header X-Forwarded-For $proxy_add_x_forwarded_for;
+ proxy_set_header X-Forwarded-Proto $scheme;
+ proxy_set_header X-NginX-Proxy true;
+
+ # This is necessary to pass the correct IP to be hashed
+ real_ip_header X-Real-IP;
+
+ proxy_connect_timeout 300;
+
+ # To support websocket
+ proxy_http_version 1.1;
+ proxy_set_header Upgrade $http_upgrade;
+ proxy_set_header Connection "upgrade";
+
+ chunked_transfer_encoding off;
+
+ proxy_pass http://minio-stanford-console;
+ }
+ location /.well-known/acme-challenge/ {
+ root /var/www/certbot;
+ }
+ }
+
diff --git a/nginx.conf b/nginx.conf
index 81813923..1531baa4 100644
--- a/nginx.conf
+++ b/nginx.conf
@@ -14,6 +14,7 @@ events {
}
http {
+ server_names_hash_bucket_size 128;
include /etc/nginx/mime.types;
default_type application/octet-stream;
@@ -28,15 +29,31 @@ http {
keepalive_timeout 65;
+ #
+ # GEN3 configuration
+ #
+ # Redirect all HTTP requests to HTTPS.
+ # Reference:
+ # https://linuxize.com/post/redirect-http-to-https-in-nginx/
server {
listen 80;
- server_name revproxy-service;
- resolver 127.0.0.11;
+ server_name _;
+ resolver 127.0.0.11;
+ # Using 307 to temporarily reidrect to HTTPS.
+ return 307 https://$host$request_uri;
+ }
+ server {
listen 443 ssl;
+ server_name revproxy-service;
+
+ # Original certificates
+ # ssl_certificate /etc/nginx/ssl/nginx.crt;
+ # ssl_certificate_key /etc/nginx/ssl/nginx.key;
- ssl_certificate /etc/nginx/ssl/nginx.crt;
- ssl_certificate_key /etc/nginx/ssl/nginx.key;
+ # New Certbot certificates
+ ssl_certificate /etc/letsencrypt/live/staging.aced-idp.org/fullchain.pem;
+ ssl_certificate_key /etc/letsencrypt/live/staging.aced-idp.org/privkey.pem;
set $access_token "";
set $csrf_check "ok-tokenauth";
@@ -89,6 +106,10 @@ http {
proxy_pass http://portal-service/;
}
+ location /.well-known/ {
+ root /etc/nginx;
+ }
+
location /user/ {
proxy_pass http://fence-service/;
}
@@ -121,6 +142,10 @@ http {
proxy_pass http://indexd-service/;
}
+ location /ga4gh/ {
+ proxy_pass http://indexd-service/ga4gh/;
+ }
+
location = /_status {
default_type application/json;
return 200 "{ \"message\": \"Feelin good!\" }\n";
@@ -137,9 +162,9 @@ http {
proxy_pass http://peregrine-service/v0/submission/getschema;
}
- location /guppy/ {
- proxy_pass http://guppy-service/;
- }
+# location /guppy/ {
+# proxy_pass http://guppy-service/;
+# }
location /api/v0/submission/graphql {
if ($cookie_csrftoken = "") {
@@ -276,5 +301,13 @@ http {
location /lw-workspace/ {
return 302 /lw-workspace/proxy;
}
+
+ location /.well-known/acme-challenge/ {
+ root /var/www/certbot;
+ }
}
+
+
+ include /etc/nginx/minio.conf;
+
}
diff --git a/scripts/discovery_metadata.sh b/scripts/discovery_metadata.sh
new file mode 100644
index 00000000..53f8dc07
--- /dev/null
+++ b/scripts/discovery_metadata.sh
@@ -0,0 +1,10 @@
+# adds a sample record to meta
+
+# in metadata container
+
+foo=$(cat <