Leverage Jupyter Notebooks with the power of your NVIDIA GPU and perform GPU calculations using Tensorflow and Pytorch in collaborative notebooks.
First of all, thanks to docker-stacks for creating and maintaining a robust Python, R and Julia toolstack for Data Analytics/Science applications. This project uses the NVIDIA CUDA image as the base image and installs their toolstack on top of it to enable GPU calculations in the Jupyter notebooks. The image of this repository is available on Dockerhub.
-
A computer with an NVIDIA GPU is required.
-
Install Docker version 1.10.0+ and Docker Compose version 1.28.0+.
-
Get access to your GPU via CUDA drivers within Docker containers. You can be sure that you can access your GPU within Docker, if the command
docker run --gpus all nvidia/cuda:11.0.3-cudnn8-runtime-ubuntu20.04 nvidia-smireturns a result similar to this one:Mon Apr 26 13:59:53 2021 +-----------------------------------------------------------------------------+ | NVIDIA-SMI 465.19.01 Driver Version: 465.19.01 CUDA Version: 11.3 | |-------------------------------+----------------------+----------------------+ | GPU Name Persistence-M| Bus-Id Disp.A | Volatile Uncorr. ECC | | Fan Temp Perf Pwr:Usage/Cap| Memory-Usage | GPU-Util Compute M. | | | | MIG M. | |===============================+======================+======================| | 0 NVIDIA GeForce ... On | 00000000:01:00.0 On | N/A | | 0% 48C P8 8W / 215W | 283MiB / 7974MiB | 11% Default | | | | N/A | +-------------------------------+----------------------+----------------------+ +-----------------------------------------------------------------------------+ | Processes: | | GPU GI CI PID Type Process name GPU Memory | | ID ID Usage | |=============================================================================| +-----------------------------------------------------------------------------+
If you don't get an output similar than this one, follow the installation steps in this medium article. The CUDA toolkit is not required on the host system, as it will be installed within the Docker containers using NVIDIA-docker. It is also important to keep your installed CUDA version in mind, when you pull images. You can't run images based on
nvidia/cuda:11.2if you have only CUDA version 10.1 installed. Check your host's CUDA-version withnvcc --versionand update to at least the same version you want to pull. -
Pull and run the image. This can last some hours, as a whole data-science environment will be downloaded:
cd your-working-directory docker run --gpus all -d -it -p 8848:8888 -v $(pwd)/data:/home/jovyan/work -e GRANT_SUDO=yes -e JUPYTER_ENABLE_LAB=yes --user root cschranz/gpu-jupyter:v1.4_cuda-11.0_ubuntu-20.04_python-only
This starts an instance with of GPU-Jupyter the tag
v1.4_cuda-11.0_ubuntu-20.04_python-only at [http://localhost:8848](http://localhost:8848) (port8484). The default password isgpu-jupyter(previouslyasdf) which should be changed as described [below](#set-password). Furthermore, data within the host'sdata` directory is shared with the container. The following images of GPU-Jupyter are available on Dockerhub:v1.4_cuda-11.0_ubuntu-20.04(full image)v1.4_cuda-11.0_ubuntu-20.04_python-only(only with a python interpreter and without Julia and R)v1.4_cuda-11.0_ubuntu-20.04_slim(only with a python interpreter and without additional packages)v1.4_cuda-11.0_ubuntu-18.04(full image)v1.4_cuda-11.0_ubuntu-18.04_python-only(only with a python interpreter and without Julia and R)v1.4_cuda-11.0_ubuntu-18.04_slim(only with a python interpreter and without additional packages)v1.4_cuda-10.1_ubuntu-18.04(full image)v1.4_cuda-10.1_ubuntu-18.04_python-only(only with a python interpreter and without Julia and R)v1.4_cuda-10.1_ubuntu-18.04_slim(only with a python interpreter and without additional packages)
The version, e.g.
v1.4, specifies a certain commit of the underlying docker-stacks. The Cuda version, e.g.cuda-11.0, has to match the host's driver version and must be supported by the gpu-libraries. These and older versions of GPU-Jupyter are listed on Dockerhub.
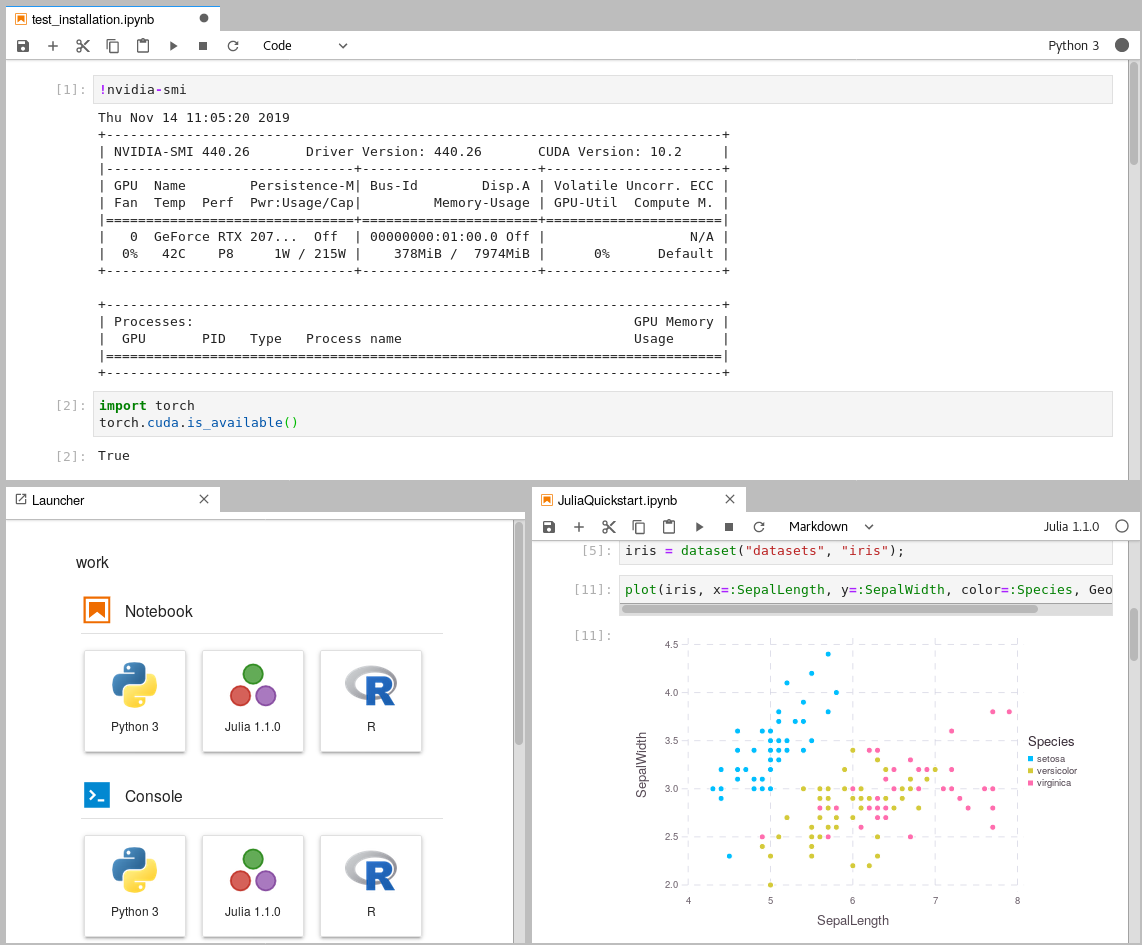
Within the Jupyterlab instance, you can check if you can access your GPU by opening a new terminal window and running
nvidia-smi. In terminal windows, you can also install new packages for your own projects.
Some example code can be found in the repository under extra/Getting_Started.
If you want to learn more about Jupyterlab, check out this tutorial.
This is the preferred option if one has a different GPU-architecture or one wants to
customize the pre-installed libraries. For the second point, the Dockerfiles in src/
are intended to be modified. In order the select a custom base image
alter src/Dockerfile.header, to install specific GPU-related libraries modify
src/Dockerfile.gpulibs and to add specific libraries append the in src/Dockerfile.usefulpackages.
After the modification, it is necessary to re-generate the Dockerfile in .build.
As soon as you have access to your GPU within Docker containers
(make sure the command docker run --gpus all nvidia/cuda:11.0.3-cudnn8-runtime-ubuntu20.04 nvidia-smi
shows your GPU statistics), you can generate the Dockerfile, build and run it.
The following commands will start GPU-Jupyter on localhost:8848
with the default password gpu-jupyter (previously asdf).
git clone https://github.com/iot-salzburg/gpu-jupyter.git
cd gpu-jupyter
git branch # Check for available supported CUDA versions
git checkout v1.4_cuda-11.0_ubuntu-20.04 # select the desired (CUDA)-version
# generate a Dockerfile with python and without Julia and R
./generate-Dockerfile.sh --python-only # generate the Dockerfile with only a python interpreter
docker build -t gpu-jupyter .build/ # will take a while
docker run --gpus all -d -it -p 8848:8888 -v $(pwd)/data:/home/jovyan/work -e GRANT_SUDO=yes -e JUPYTER_ENABLE_LAB=yes -e NB_UID="$(id -u)" -e NB_GID="$(id -g)" --user root --restart always --name gpu-jupyter_1 gpu-jupyterThis starts a container WITH GPU support, a shared local data volume data
and some other configurations like root permissions which are necessary to install packages within the container.
For more configurations, scroll down to Configuration of the Dockerfile-Generation.
Start GPU-Jupyter using docker-compose.yml:
docker-compose up --build -dThis step requires a docker-compose version of at least 1.28.0,
as the dockerfile requests GPU resources
(see changelog).
To update docker-compose, this discussion may be useful.
With these commands we can see if everything worked well:
docker ps
docker logs [service-name] # or
docker-compose logs -fIn order to stop the local deployment, run:
docker ps
docker rm -f [service-name] # or
docker-compose downThe script generate-Dockerfile.sh generates a Dockerfile within the .build/
directory.
This implies that this Dockerfile is overwritten by each generation.
The Dockerfile-generation script generate-Dockerfile.sh
has the following parameters (note that 2, 3 and 4 are exclusive):
-
-h|--help: Show a help message. -
-p|--pw|--password: Set the password for GPU-Jupyter by updating the salted hashed token insrc/jupyter_notebook_config.json. -
-c|--commit: specify a commit or"latest"for thedocker-stacks, the default commit is a working one. -
-s|--slim: Generate a slim Dockerfile. As some installations are not needed by everyone, there is the possibility to skip some installations to reduce the size of the image. Here thedocker-stackscipy-notebookis used instead ofdatascience-notebookthat comes with Julia and R. Moreover, none of the packages withinsrc/Dockerfile.usefulpackagesis installed. -
--python-only|--no-datascience-notebook: As the name suggests, thedocker-stackdatascience-notebookis not installed on top of thescipy-notebook, but the packages withinsrc/Dockerfile.usefulpackagesare. -
--no-useful-packages: On top of thedocker-stackdatascience-notebook(Julia and R), the essentialgpulibsare installed, but not the packages withinsrc/Dockerfile.usefulpackages.
Custom packages can be installed within a container, or by modifying the file
src/Dockerfile.usefulpackages.
As .build/Dockerfile is overwritten each time a Dockerfile is generated,
it is suggested to append custom installations either
within src/Dockerfile.usefulpackages or in generate-Dockerfile.sh.
If an essential package is missing in the default stack, please let us know!
The easiest way to set a password is by giving it as an parameter:
bash generate-Dockerfile.sh --password your_passwordThis updates the salted hashed token within src/jupyter_notebook_config.json.
Another way to specify your password is to directly change the token in src/jupyter_notebook_config.json.
Therefore, hash your password in the form (password)(salt) using a sha1 hash generator, e.g.,
the sha1 generator of sha1-online.com. The input with the
default password gpu-jupyter (previously asdf) is concatenated by an arbitrary salt
3b4b6378355 to gpu-jupyter3b4b6378355 and is hashed to 642693b20f0a33bcad27b94293d0ed7db3408322.
Never give away your own unhashed password!
Then update the config file as shown below, generate the Dockerfile and restart GPU-Jupyter.
{
"NotebookApp": {
"password": "sha1:3b4b6378355:642693b20f0a33bcad27b94293d0ed7db3408322"
}
}The host's CUDA-version must be equal or higher than that of the
container itself (in Dockerfile.header).
Check the host's version with nvcc --version and the version compatibilities
for CUDA-dependent packages as Pytorch
respectively Tensorflow previously.
Then modify, if supported, the CUDA-version in Dockerfile.header to, e.g.:
the line:
FROM nvidia/cuda:11.1-base-ubuntu20.04
and in the Dockerfile.pytorch the line:
cudatoolkit=11.1
Then re-generate, re-build and run the updated image, as closer described above: Note that a change in the first line of the Dockerfile will re-build the whole image.
/generate-Dockerfile.sh --python-only # generate the Dockerfile with only a python interpreter
docker build -t gpu-jupyter .build/ # will take a while
docker run --gpus all -d -it -p 8848:8888 -v $(pwd)/data:/home/jovyan/work -e GRANT_SUDO=yes -e JUPYTER_ENABLE_LAB=yes -e NB_UID="$(id -u)" -e NB_GID="$(id -g)" --user root --restart always --name gpu-jupyter_1 gpu-jupyterThe docker-stacks are used as a
submodule within .build/docker-stacks. Per default, the head of the commit is reset to a commit on which gpu-jupyter runs stable.
To update the generated Dockerfile to a specific commit, run:
./generate-Dockerfile.sh --commit c1c32938438151c7e2a22b5aa338caba2ec01da2To update the generated Dockerfile to the latest commit, run:
./generate-Dockerfile.sh --commit latestA new build can last some time and may consume a lot of data traffic. Note, that the latest version may result in a version conflict! More info to submodules can be found in this tutorial.
A Jupyter instance often requires data from other services. If that data-source is containerized in Docker and sharing a port for communication shouldn't be allowed, e.g., for security reasons, then connecting the data-source with GPU-Jupyter within a Docker Swarm is a great option!
This step requires a running Docker Swarm on a cluster or at least on this node. In order to register custom images in a local Docker Swarm cluster, a registry instance must be deployed in advance. Note that the we are using the port 5001, as many services use the default port 5000.
sudo docker service create --name registry --publish published=5001,target=5000 registry:2
curl 127.0.0.1:5001/v2/This should output {}. \
Afterwards, check if the registry service is available using docker service ls.
Additionally, GPU-Jupyter is connected to the data-source via the same docker-network. Therefore, This network must be set to attachable in the source's docker-compose.yml:
services:
data-source-service:
...
networks:
- default
- datastack
...
networks:
datastack:
driver: overlay
attachable: true In this example,
- the docker stack was deployed in Docker swarm with the name elk (
docker stack deploy ... elk), - the docker network has the name datastack within the
docker-compose.ymlfile, - this network is configured to be attachable in the
docker-compose.ymlfile - and the docker network has the name elk_datastack, see the following output:
sudo docker network ls # ... # [UID] elk_datastack overlay swarm # ...
The docker network name elk_datastack is used in the next step as a parameter.
Finally, GPU-Jupyter can be deployed in the Docker Swarm with the shared network, using:
./generate-Dockerfile.sh
./add-to-swarm.sh -p [port] -n [docker-network] -r [registry-port]
# e.g. ./add-to-swarm.sh -p 8848 -n elk_datastack -r 5001where:
- -p: port specifies the port on which the service will be available.
- -n: docker-network is the name of the attachable network from the previous step, e.g., here it is elk_datastack.
- -r: registry port is the port that is published by the registry service, default is
5000.
Now, gpu-jupyter will be accessible here on localhost:8848
with the default password gpu-jupyter (previously asdf) and shares the network with the other data-source, i.e.,
all ports of the data-source will be accessible within GPU-Jupyter,
even if they aren't routed it the source's docker-compose file.
Check if everything works well using:
sudo docker service ps gpu_gpu-jupyter
docker service ps gpu_gpu-jupyterIn order to remove the service from the swarm, use:
./remove-from-swarm.sh-
Can't execute the bash scripts.
$ bash generate-Dockerfile.sh generate-Dockerfile.sh: line 2: cd: $'/path/to/gpu-jupyter\r': No such file or directory generate-Dockerfile.sh: line 3: $'\r': command not found generate-Dockerfile.sh: line 9: $'\r': command not found generate-Dockerfile.sh: line 11: syntax error near unexpected token `$'in\r'' generate-Dockerfile.sh: line 11: `while [[ "$#" -gt 0 ]]; do case $1 in
The reason for this issue is that the line-breaks between Unix and Windows based systems are different and in the current version different.
Solution: Change the line-endings of the bash scripts. This can be done e.g. in the Notepad++ under Edit - EOL Conversion, or using
dos2unixsudo apt install dos2unix dos2unix generate-Dockerfile.sh
-
No GPU available - error The docker-compose start breaks with:
ERROR: for fc8d8dfbebe9_gpu-jupyter_gpu-jupyter_1 Cannot start service gpu-jupyter: OCI runtime create failed: container_linux.go:370: starting container process caused: process_linux.go:459: container init ca used: Running hook #0:: error running hook: exit status 1, stdout: , stderr: nvidia-container-cli: initialization error: driver error: failed to process request: unknown
Solution:
Check if the GPU is available on the host node vianvidia-smiand run with the describeddocker-commands. If the error still occurs, so try there could be an issue that docker can't visualize the GPU.
This project has the intention to create a robust image for CUDA-based GPU-applications, which is built on top of the docker-stacks. You are free to help to improve this project, by: