Cellpose-mcp is a Model Context Protocol (MCP) server that enables AI assistants like Claude, Cursor IDE, etc. to perform cell segmentation through natural language commands. This tool exposes comprehensive Cellpose functionality through 13+ MCP tools, including 2D/3D segmentation, batch processing, image restoration (denoising, deblurring, upsampling), and custom model training. The system integrates seamlessly with Napari, enabling complete workflows from segmentation to interactive visualization.
📌 Note: This project started as a fun project inspired by napari-mcp and adapted for Cellpose segmentation workflows. If you would like to contribute then please get in touch with me at ssahu2@ucmerced.edu.
Requirements: Python 3.10 or later (use a virtual environment or conda).
Install from PyPI:
pip install cellpose-mcpInstall and configure for Cursor in one go:
pip install cellpose-mcp && cellpose-mcp-install cursorThe installer uses the Python that runs the command (or a conda env named Cellpose_mcp if present). Restart Cursor after configuring.
Development install (from source):
git clone https://github.com/surajinacademia/cellpose_mcp.git
cd cellpose_mcp
pip install -e .After pip install cellpose-mcp, run the installer for your app. It writes to the correct MCP config file using your current Python.
| Application | Command | Notes |
|---|---|---|
| Cursor IDE | cellpose-mcp-install cursor |
Writes to ~/.cursor/mcp.json |
| Claude Desktop | cellpose-mcp-install claude-desktop |
Adds to Claude Desktop config |
| Antigravity | cellpose-mcp-install antigravity |
Configures Antigravity MCP |
| VS Code (Cline/Roo Cline) | cellpose-mcp-install vscode |
Configures Cline/Roo Cline extension |
| Claude Code | Manual only | See Manual Configuration below |
Options: --python-path /path/to/python to use a specific Python; --env-name NAME to use a conda env (default: Cellpose_mcp).
Manual Configuration for Claude Code
If you prefer manual setup (or use Claude Code), create a .mcp.json file in your project root. Use the full path to your Python executable if python is not the one that has cellpose-mcp installed (e.g. a venv or conda):
{
"mcpServers": {
"cellpose": {
"command": "python",
"args": ["-m", "cellpose_mcp"],
"env": {
"KMP_DUPLICATE_LIB_OK": "TRUE",
"OMP_NUM_THREADS": "1"
}
}
}
}For Cursor, use the same structure in ~/.cursor/mcp.json (global) or .cursor/mcp.json in your project.
After installation, restart your AI app and try asking:
"Can you list available Cellpose models?"
"Segment the cells in ./data/sample.tif using the cyto2 model"
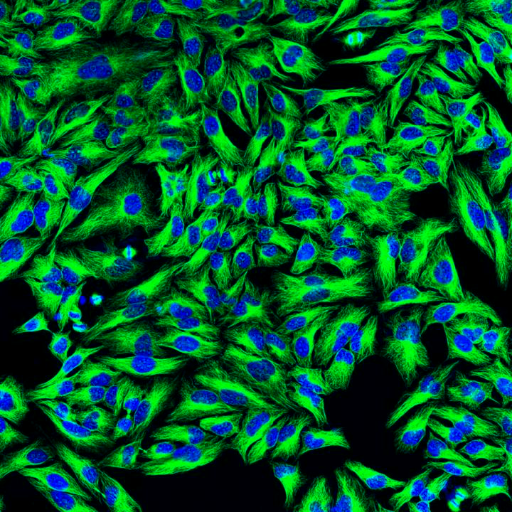
Original Image: Fluorescence microscopy with green-stained cytoplasm and blue-stained nuclei |
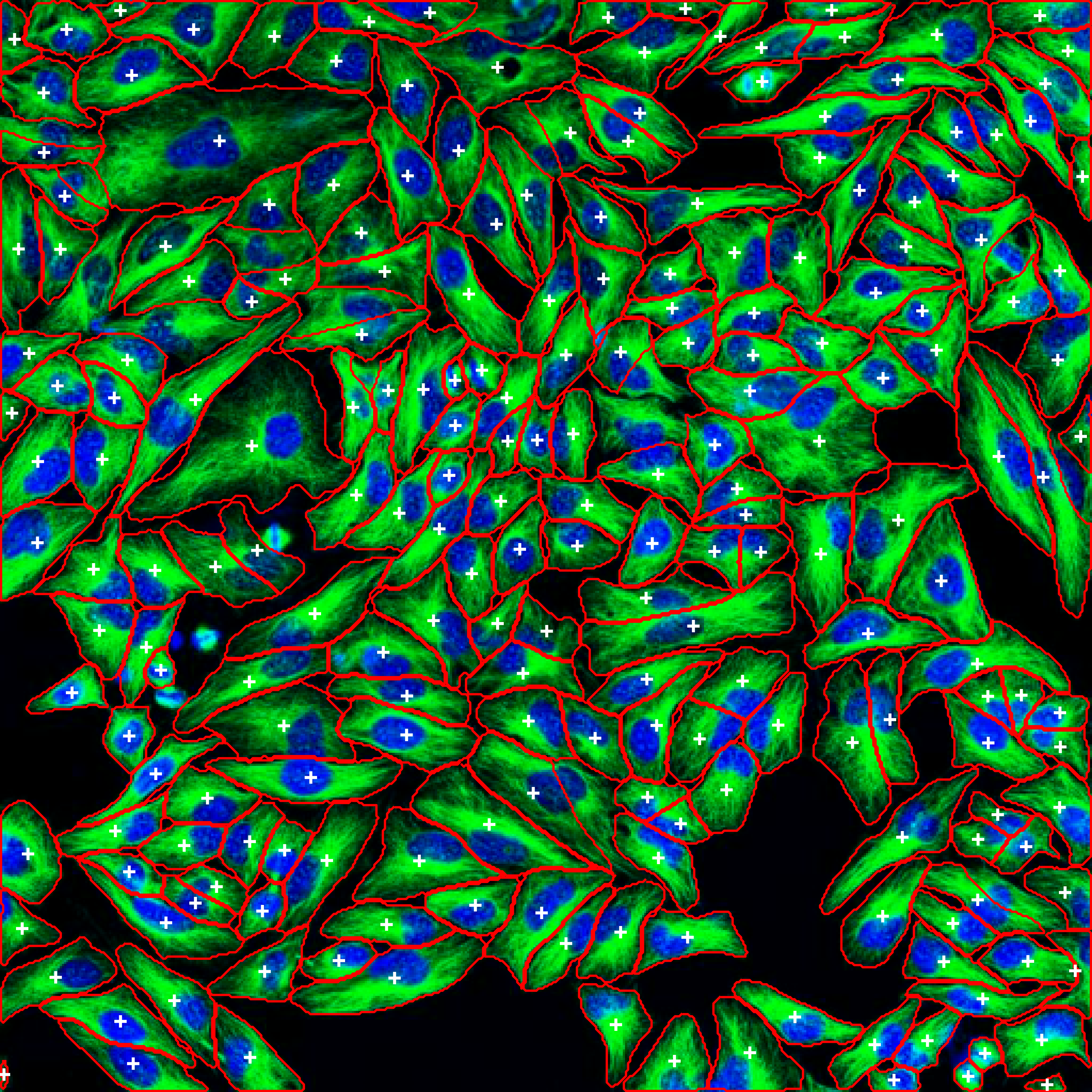
Segmented Result: Cells automatically detected with boundaries and labels |
"Segment the cells in ./data/sample.tif using the cyto2 model"
"List available Cellpose models"
"Estimate cell diameter from ./data/image.tif"
"Segment all TIFF files in ./data/images/ and save masks to ./output/"
"Train a custom segmentation model using images in ./train/images/ and masks in ./train/masks/"
"Restore and segment the noisy image in ./data/noisy.tif using oneclick_cyto3"
"Process all images in ./data/ with the cyto2 model and save results to ./output/"
The server exposes 13+ tools for complete Cellpose functionality:
segment_cells_2d- Segment cells in 2D imagessegment_cells_3d- Segment cells in 3D volumessegment_cells_batch- Batch process multiple images
denoise_image- Denoise microscopy imagesdeblur_image- Deblur microscopy imagesupsample_image- Upsample low-resolution imagesrestore_and_segment- Combined restoration + segmentation
train_segmentation_model- Train custom segmentation modeltrain_restoration_model- Train custom restoration model
list_available_models- List all pretrained modelsestimate_cell_diameter- Estimate cell diameter from imagesave_masks- Save masks in various formatsload_image_info- Get image metadata
- Quick Start Guide - Get running in 3 steps
- Available Tools - Complete tool list
- Release Notes - Detailed v0.1.0 release information
- Changelog - Version history and changes
- FastMCP Server: Handles MCP protocol communication
- Cellpose Integration: Manages model loading and segmentation operations
- Tool Layer: Exposes Cellpose functionality as MCP tools
- File I/O: Handles image reading, writing, and mask generation
Key features:
- Thread-safe: All operations are properly serialized
- Non-blocking: Async operations for better performance
- Napari Integration: Integration with Napari for visualization and analysis
Author: Suraj Sahu
Affiliation: Department of Physics, University of California Merced, CA, USA
Email: ssahu2@ucmerced.edu
BSD-3-Clause License - see LICENSE file for details.
- Napari MCP by royerlab
- Cellpose team for the excellent segmentation library
- FastMCP for the MCP framework
- Anthropic for Claude and MCP development
- Model Context Protocol - Open standard for AI-tool integration