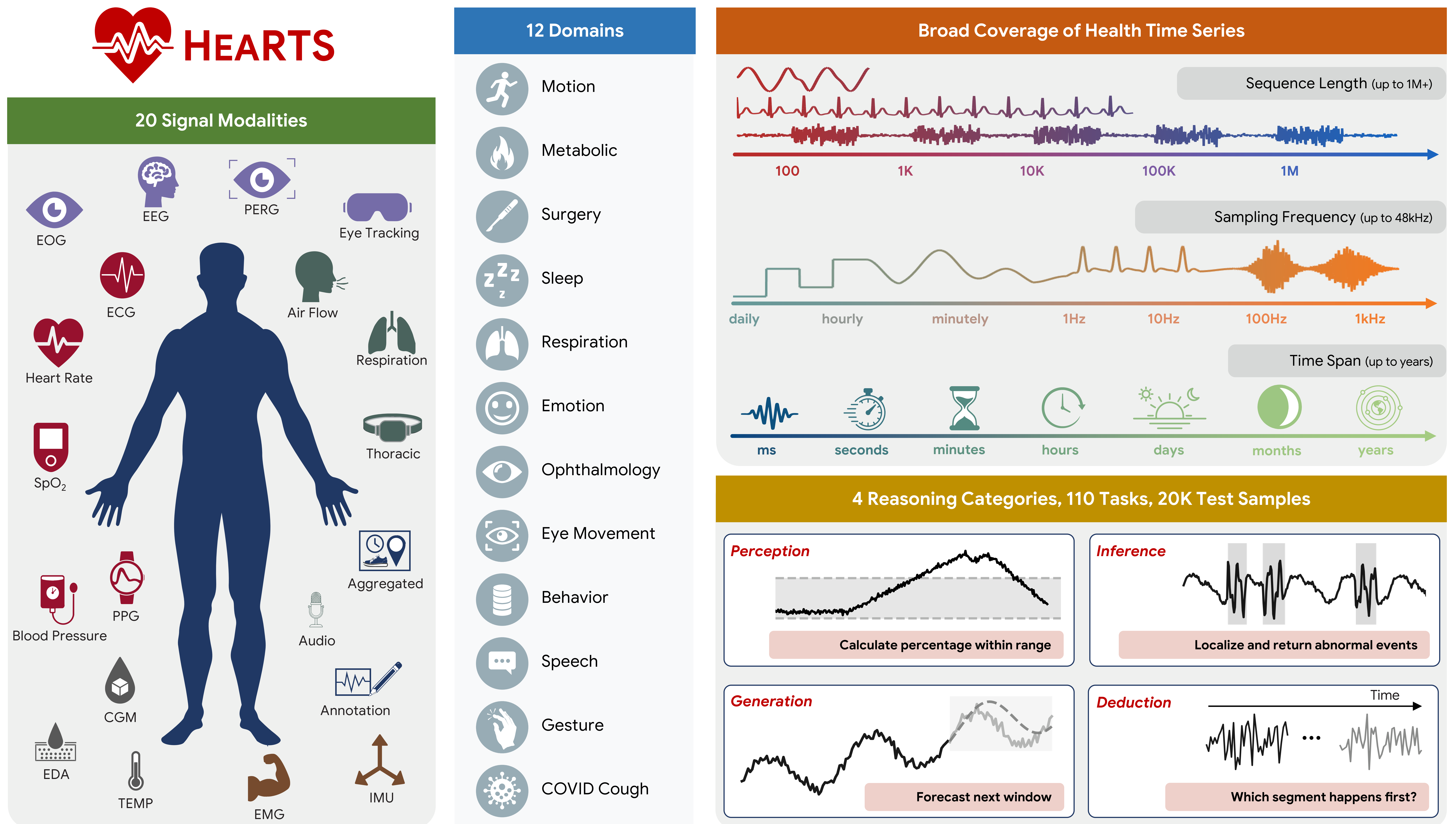
HEARTS (Health Reasoning over Time Series) is a benchmark and evaluation framework for testing how well LLM agents reason over real-world health time-series data.
📄 Paper | 🌐 Website | 🤗 Data | 🏃 Quickstart | ⚙️ Configuration | 🧪 Experiments | 🕵️ Agents | 📚 Citation
HEARTS goes beyond narrow forecasting and simple QA by evaluating four hierarchical capabilities:
- Perception: signal-level measurement and feature extraction
- Inference: event localization, physiological classification, and subject-level profiling
- Generation: forecasting, imputation, and cross-modal translation
- Deduction: temporal ordering and longitudinal trajectory analysis
| Item | Count |
|---|---|
| Datasets | 16 |
| Health domains | 12 |
| Signal modalities | 20 |
| Tasks | 110 |
| Test samples | 20,226 |
Domains include motion, metabolic health, surgery, sleep, respiration, emotion, ophthalmology, eye movement, behavior, speech, gesture, and COVID cough.
Health time-series reasoning is hard because data vary dramatically across:
- modality (ECG, EEG, PPG, EMG, audio, eye tracking, CGM, etc.)
- frequency (daily aggregates to 48 kHz)
- sequence length (tens of points to 1M+)
- time span (seconds to years)
HEARTS is designed to evaluate this full range in a single unified setting.
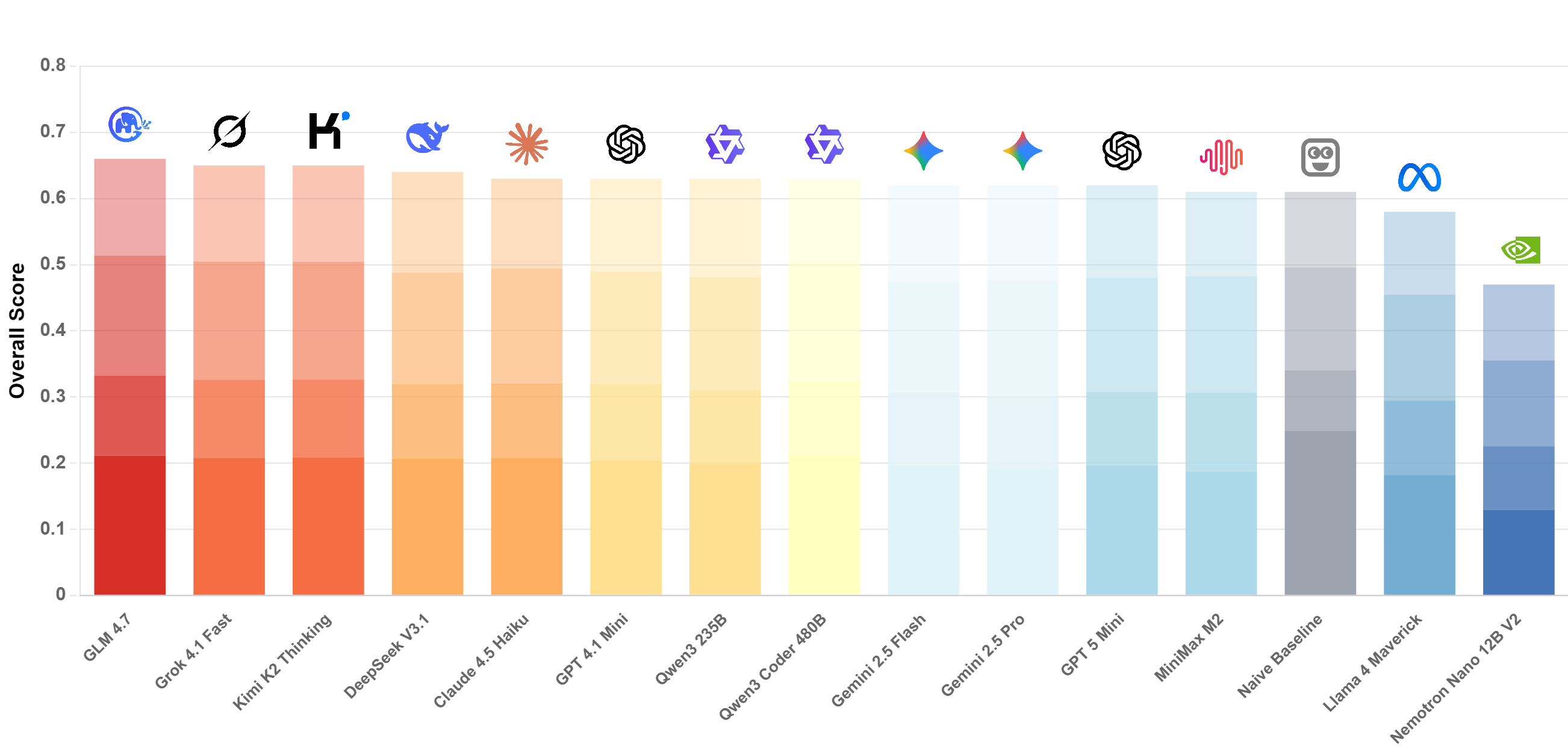
- LLMs underperform specialized time-series models on many health reasoning tasks. And performance on HEARTS is only weakly correlated with broad "general reasoning" indices.
- Models often rely on low-complexity heuristics (copying/interpolation/rule shortcuts) instead of deep temporal reasoning.
- Performance degrades with longer sequences and higher sampling frequencies, with a shared, model-agnostic difficulty ordering across domains and input modalities.
- Models in the same family show similar performance patterns, suggesting scaling alone is not sufficient.
- The input format of time-series (text/image/raw file) mainly shifts absolute performance, while relative task difficulty remains consistent across formats.
- Modular architecture: experiments (
exp/), agents (agents/), and shared utilities (utils/). - Diverse datasets: supports multiple healthcare datasets (e.g., SHHS, Capture24, VitalDB, Bridge2AI Voice).
- Task variety: Perception, Inference, Generation, and Deduction tasks.
- Agent flexibility: built-in support for different agent architectures (e.g., CodeAct).
- Model support: interfaces for major LLM providers (OpenAI, AWS Bedrock, Google Gemini, XAI).
- Reproducibility: "frozen" experiment execution on fixed test cases for consistent benchmarking.
This project uses uv for dependency management.
Requirements: Python >=3.11.
curl -LsSf https://astral.sh/uv/install.sh | shuv syncCreate a .env file (API keys are read from environment variables).
cp .env.example .envOn Windows PowerShell:
copy .env.example .envThen edit .env and set at least one provider key (examples in .env.example).
The primary entry point for running fixed experiments is run_exp_freeze.py. It executes a set of pre-defined ("fixed") test cases to ensure reproducibility.
Provide a directory containing fixed test cases (pickles) laid out as:
Fixed test cases can be downloaded from https://huggingface.co/datasets/yang-ai-lab/HEARTS (extract locally and point --fix-test-cases-dir to the extracted root).
fix_test_cases_dir/{dataset}/{task}/{index}.pkl
uv run run_exp_freeze.py --fix-test-cases-dir /path/to/test_casesTip: use --dry-run to validate your setup without calling a model.
You can configure an experiment using config.yaml and/or command-line arguments. The repository includes an example at config.yaml.example.
Example config.yaml:
dataset_name: cgmacros
task:
name: cgm_stat_calculation
params:
example_param: value
num_test: 50
model_name: gpt-4.1-mini
agent:
name: codeact
params:
verbose: true
fix_test_cases_dir: /path/to/fixed_cases # can also be set via CLI
result_dir: results/Common command-line overrides (these take precedence over config.yaml):
--task: task name--dataset-name: dataset name--num-test: number of test cases to run--model-name: model identifier (e.g.,gpt-4.1,gemini-3-pro-preview)--result-dir: directory to save results--logs-dir: directory to save agent logs--n-jobs: number of concurrent jobs (default: 2)--dry-run: run without calling the model
Experiment results and agent execution logs are saved automatically.
Results:
- JSON files written to
result_dir(default:results/) - filename format:
{dataset}_{task}_{agent}_{timestamp}.json - each file contains per-test-case entries (e.g.,
query_id,subject_id,GT,solution) plus a final entry with computedmetricsand the fullconfig
Logs (agent state):
- pickles written to
logs/{task}/{query_id}.pkl - include conversation history, code execution, and intermediate state; useful for debugging and analysis
.
├── agents/ # agent implementations
│ ├── base/ # abstract base class for agents
│ ├── codeact/ # CodeAct agent implementation
├── exp/ # experiment definitions (datasets & tasks)
│ ├── base/ # base experiment classes
│ ├── templates/ # task templates (classification, forecasting, etc.)
│ ├── {dataset_name}/ # dataset-specific experiment implementations
│ └── utils/ # experiment registry and utilities
├── utils/ # core utilities
│ ├── exp.py # experiment runner logic
│ ├── model_enums.py # supported model definitions
│ ├── metric.py # evaluation metrics
│ └── ...
├── run_exp_freeze.py # main execution script for frozen experiments
├── .env.example # environment variable template for API keys
├── config.yaml.example # example configuration file
└── pyproject.toml # project dependencies and metadata
Experiments are defined in exp/{dataset_name}/{task}.py. For details on adding new experiments, see exp/README.md.
Examples of included datasets:
bridge2ai_voice: voice analysis tasks (Parkinson's prediction, etc.)capture24: activity tracking and health monitoringcoswara: COVID-19 detection from audiovitaldb: vital signs monitoring and prediction
Agents are defined in agents/{agent_name}/. For details on implementing new agents, see agents/README.md.
Available agents:
- CodeAct: uses executable code to interact with data and solve tasks
- Follow
exp/README.mdto add new experiments. - Follow
agents/README.mdto add new agents. - Run
uv syncto keep dependencies up to date.
If you use HEARTS in your research, please cite:
@article{hearts2026,
title={HEARTS: Benchmarking LLM Reasoning on Health Time Series},
author={Sirui Li and Shuhan Xiao and Mihir Joshi and Ahmed Metwally and Daniel McDuff and Wei Wang and Yuzhe Yang},
journal={arXiv preprint arXiv:2603.06638},
year={2026}
}